Abstract
Key message
Analysis of terpenoids content, transcriptome from Chamaemelum nobile showed that the content of the terpenoids in the roots was the highest and key genes involved in the terpenoids synthesis pathway were identified.
Abstract
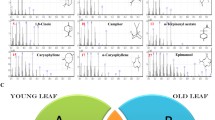
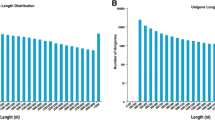
Chamaemelum nobile is a widely used herbaceous medicinal plant rich in volatile oils, mainly composed of terpenoids. It is widely used in food, cosmetics, medicine, and other fields. In this study, we analyzed the transcriptome and the content and chemical composition of the terpenoids in different organs of C. nobile. Gas chromatography–mass spectrometry analysis showed that the total content of the terpenoids among C. nobile organs was highest in the roots, followed by the flowers. Illumina HiSeq 2500 high-throughput sequencing technology was used to sequence the transcripts of roots, stems, leaves, and flowers of C. nobile. We obtained 139,757 unigenes using the Trinity software assembly. A total of 887 unigenes were annotated to secondary metabolism. In total, 55,711 differentially expressed genes were screened among different organs of C. nobile. We identified 16 candidate genes that may be involved in the terpenoid biosynthesis from C. nobile and analyzed their expression patterns using real-time PCR. Results showed that the expression pattern of these genes was tissue-specific and had significant differential expression levels in different organs of C. nobile. Among these genes, 13 were expressed in roots with the highest levels. Furthermore, the transcript levels of these 13 genes were positively correlated with the content of α-pinene, β-phellandrene, 1,8-cineole, camphor, α-terpineol, carvacrol, (E,E)-farnesol and chamazulene, suggesting that these 13 genes may be involved in the regulation of the synthesis of the volatile terpenoids. These results laid the foundation for the subsequent improvement of C. nobile quality through genetic engineering.










Similar content being viewed by others
References
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:1–12
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S (2004) UniProt: the universal protein knowledgebase. Nucleic Acids Res 32:115–119
Ashburner M (2000) Gene Ontology: tool for the unification of biology. Nat Genet 25:25–29
Banyai W, Kirdmanee C, Mii M, Supaibulwatana K (2010) Overexpression of farnesyl pyrophosphate synthase (FPS) gene affected artemisinin content and growth of Artemisia annua L. Plant Cell Tiss Org 103:255–265
Barabasi AL, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nat Rev Genet 5:101–113
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188
Bohlmann J, Keeling CI (2008) Terpenoid biomaterials. Plant J 54:656–669
Buhaescu I, Izzedine H (2007) Mevalonate pathway: A review of clinical and therapeutical implications. Clin Biochem 40:575–584
Carlson MR, Zhang B, Fang Z, Mischel PS, Horvath S, Nelson SF (2006) Gene connectivity, function, and sequence conservation: predictions from modular yeast co-expression networks. BMC Genom 7:40
Chen H, Xin C, Jing T, Yong Y, Liu Z, Hao X, Wang L, Wang S, Liang J, Zhang L, Yin F, Cheng X (2016) Development of gene-based SSR markers in rice bean (Vigna umbellata L.) based on transcriptome data. PLoS One 11:e0151040
Cheng S, Wang X, Xu F, Chen Q, Tao T, Lei J, Zhang W, Liao Y, Chang J, Li X (2016) Cloning, expression profiling and functional analysis of CnHMGS, a gene encoding 3-hydroxy-3-methylglutaryl coenzyme a synthase from Chamaemelum nobile. Molecules 21:316
Deng YY, Li JQ, Wu SF, Zhu YP, Chen YW, He FC (2006) Integrated nr database in protein annotation system and its localization. Comput En 32:71–74
Dudareva N, Cseke L, Blanc VM, Pichersky E (1996) Evolution of floral scent in Clarkia: novel patterns of S-linalool synthase gene expression in the C. breweri flower. Plant Cell 8:1137–1148
Dudareva N, Martin D, Kish CM, Kolosova N, Gorenstein N, Faldt J, Miller B, Bohlmann J (2003) (E)-beta-ocimene and myrcene synthase genes of floral scent biosynthesis in snapdragon: function and expression of three terpene synthase genes of a new terpene synthase subfamily. Plant Cell 15:1227–1241
Farhoudi R (2013) Chemical constituents and antioxidant properties of Matricaria recutita and Chamaemelum nobile essential oil growing wild in the south west of Iran. J Essent Oil Bear Plant 16:531–537
Gao B, Zhang D, Li X, Yang H, Zhang Y, Wood AJ (2015) De novo transcriptome characterization and gene expression profiling of the desiccation tolerant moss Bryum argenteum following rehydration. BMC Genom 16:1–14
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Heuston S, Begley M, Davey MS, Eberl M, Casey PG, Hill C, Gahan CGM (2012) HmgR, a key enzyme in the mevalonate pathway for isoprenoid biosynthesis, is essential for growth of Listeria monocytogenes EGDe. Microbiology 158:1684–1693
Huber W, Carey VJ, Long L, Falcon S, Gentleman R (2007) Gentleman R: Graphs in molecular biology. BMC Bioinformatics 8:S8
Irmisch S, Krause ST, Kunert G, Gershenzon J, Degenhardt J, Köllner TG (2012) The organ-specific expression of terpene synthase genes contributes to the terpene hydrocarbon composition of chamomile essential oils. BMC Plant Biol 12:1–13
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:277–280
Kent WJ (2002) BLAT-the BLAST-like alignment tool. Genome Res 12:656–664
Landmann C, Fink B, Festner M, Dregus M, Engel KH, Schwab W (2007) Cloning and functional characterization of three terpene synthases from lavender (Lavandula angustifolia). Arch Biochem Biophys 465:417–429
Liu Z, Li X, Simoneau AR, Jafari M, Zi X (2006) Isoprenoid biosynthesis in plants: pathways, genes, regulation and metabolic engineering. J Biol Sci 351:371–374
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2–∆∆C T method. Methods 25:402–408
Ma CM, Winsor L, Daneshtalab M (2007) Quantification of spiroether isomers and Herniarin of different parts of Matricaria matricarioides and flowers of Chamaemelum nobile. Phytochem Anal 18:42–49
Meng XX, Yan JP, Liu XM, Liao YL, Chang J, Xu F (2016) Cloning and expression analysis of HMGR gene from Chamaemelum nobile. Acta Agric Boreali Sin 31:68–75
Misra I, Wang CZ, Miziorko HM (2003) The influence of conserved aromatic residues in 3-hydroxy-3-methylglutaryl-CoA synthase. J Biol Chem 278:26443–26449
Murat C, Riccioni C, Belfiori B, Cichocki N, Labbé J, Morin E, Tisserant E, Paolocci F, Rubini A, Martin F (2011) Distribution and localization of microsatellites in the Perigord black truffle genome and identification of new molecular markers. Fungal Genet Biol 48:592–601
Nagegowda DA, Gutensohn M, Wilkerson CG, Dudareva N (2008) Two nearly identical terpene synthases catalyze the formation of nerolidol and linalool in snapdragon flowers. Plant J 55:224–239
Newall CA, Anderson LA, Phillipson JD (1996) Herbal medicines. A guide for health-care professionals. Pharm Press 296
Nikiforova VJ, Willmitzer L (2007) Network visualization and network analysis. Plant Syst Biol 97:245–275
Rama Reddy NR, Mehta RH, Soni PH, Makasana J, Gajbhiye NA, Ponnuchamy M, Kumar J (2015) Next generation sequencing and transcriptome analysis predicts biosynthetic pathway of sennosides from Senna (Cassia angustifolia Vahl.), a non-model plant with potent laxative properties. PLoS One 10:e0129422
Schilmiller AL, Schauvinhold I, Larson M, Xu R, Charbonneau AL, Schmidt A, Wilkerson C, Last RL, Pichersky E (2009) Monoterpenes in the glandular trichomes of tomato are synthesized from a neryl diphosphate precursor rather than geranyl diphosphate. Pro Natl Acad Sci USA 106:10865–10870
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28:33–36
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, Van Baren MJ, Salzberg LS, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515
Xu F, Ning Y, Zhang W, Liao Y, Li L, Cheng H, Cheng S (2014) An R2R3-MYB transcription factor as a negative regulator of the flavonoid biosynthesis pathway in Ginkgo biloba. Funct Integr Genomic 14:177–189
Yan JP, Meng XX, Li Z, Zhang WW, Jie C, Xu F (2017) Molecular cloning and expression analysis of germacrene A synthase gene in Chamaemelum nobile. Chinese Tradit Herb Drugs 48:1851–1859
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L, Wang J (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:293–297
Yu F, Utsumi R (2009) Deversity, regulation, and genetic manipulation of plant mono-and sesquiterpenoid biosynthesis. Cell Mol Life Sci 66:3043–3052
Zenoni S, Delledonne M (2010) Characterization of transcriptional complexity during berry development in Vitis vinifera using RNA-SEq. Plant Physiol 152:1787–1795
Zhao L, Chang W, Xiao Y, Liu H, Liu P (2013) Methylerythritol phosphate pathway of isoprenoid biosynthesis. Annu Rev Biochem 82:497–530
Acknowledgements
This work was supported by National Natural Science Foundation of China (Grant no. 31400603).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Salim Al-Babili.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, X., Wang, X., Chen, Z. et al. De novo assembly and comparative transcriptome analysis: novel insights into terpenoid biosynthesis in Chamaemelum nobile L.. Plant Cell Rep 38, 101–116 (2019). https://doi.org/10.1007/s00299-018-2352-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-018-2352-z