Abstract
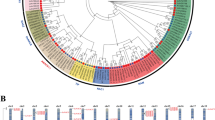
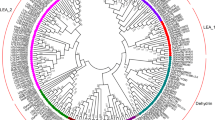
As the crucial members of the cold-regulated (COR) gene family, KIN genes are involved in diverse abiotic stress responses in plants. In the present study, KIN genes from the widespread plant Capsella bursa-pastoris were identified and analyzed to better understand the powerful adaptation of this species. Two KIN genes were cloned and sequenced by 3′ RACE. As some COR genes are homologous to LEA genes, three KIN-homologous LEA genes were also identified. We deduced the amino acid sequences of the five proteins to estimate their phylogenetic relationships, and grouped them into three subfamilies (CI, CII, and CIII). Variable 3′ UTRs were found in CI, CII, and CIII genes. Using qPCR, we evaluated the transcriptional levels of the five genes in different organs and embryonic stages. Two CI genes were exclusively expressed in early embryos and flowers. The CII and CIII genes showed obvious up-regulation in young leaves after heat stress, cold stress, and ABA treatment. Two of the CI genes, however, rarely responded to those stresses in young leaves. In contrast, all five genes showed differential responses in flowers when C. bursa-pastoris plants were sprayed with ABA. Furthermore, the expression of these genes in C. bursa-pastoris was compared to that of the corresponding Arabidopsis genes, and similar gene expression profiles were found in both species. Our findings suggest that these five genes play different roles in development and the responses to abiotic stresses in C. bursa-pastoris.
Key message We characterized two KIN and three KIN-homologous LEA genes, and analyzed their variable 3’UTR and organ-specific, embryo-developmental, stress-induced gene expression in Capsella bursa-pastoris.



Similar content being viewed by others
References
Battaglia M, Olvera-Carrillo Y, Garciarrubio A, Campos F, Covarrubias AA (2008) The enigmatic LEA proteins and other hydrophilins. Plant Physiol 148:6–24
Bhattramakki D, Dolan M, Hanafey M, Wineland R, Vaske D, Register JC, Tingey SV, Rafalski A (2002) Insertion-deletion polymorphisms in 3′ regions of maize genes occur frequently and can be used as highly informative genetic markers. Plant Mol Biol 48:539–547
Bies-Ethève N, Gaubier-Comella P, Debures A, Lasserre E, Jobet E, Raynal M, Cooke R, Delseny M (2008) Inventory, evolution and expression profiling diversity of the LEA (late embryogenesis abundant) protein gene family in Arabidopsis thaliana. Plant Mol Biol 67:107–124
Chen L, Zhong H, Ren F, Guo QQ, Hu XP, Li XB (2011) A novel cold-regulated gene, COR25, of Brassica napus is involved in plant response and tolerance to cold stress. Plant Cell Rep 30:463–471
Eveland AL, McCarty DR, Koch KE (2008) Transcript profiling by 3′-untranslated region sequencing resolves expression of gene families. Plant Physiol 146:32–44
Ganeshan S, Vitamvas P, Fowler DB, Chibbar RN (2008) Quantitative expression analysis of selected COR genes reveals their differential expression in leaf and crown tissues of wheat (Triticum aestivum L.) during an extended low temperature acclimation regimen. J Exp Bot 59:2393–2402
Heidarvand L, Amiri RM (2010) What happens in plant molecular responses to cold stress? Acta Physiol Plant 32:419–431
Huang D, Wu W, Abrams SR, Cutler AJ (2008) The relationship of drought-related gene expression in Arabidopsis thaliana to hormonal and environmental factors. J Exp Bot 59:2991–3007
Hundertmark M, Hincha DK (2008) LEA (late embryogenesis abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genomics 9:118
Hurka H, Neuffer B (1997) Evolutionary processes in the genus Capsella (Brassicaceae). Plant Syst Evol 206:295–316
Hurka H, Bleeker W, Neuffer B (2003) Evolutionary process associated with biological invasions in the Brassicaceae. Biol Invasions 5:281–292
Kosova K, Vitamvas P, Prasil IT (2007) The role of dehydrins in plant response to cold. Biol Plant 51:601–617
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Kurkela S, Franck M (1990) Cloning and characterization of a cold- and ABA-inducible Arabidopsis gene. Plant Mol Biol 15:137–144
Le BH, Cheng C, Bui AQ, Wameister JA, Henry KF, Pelletier J, Kwong L, Belmonte M, Kirkbride R, Horvath S, Drews GN, Fischer RL, Okamuro JK, Harada JJ, Goldberg RB (2010) Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. Proc Natl Acad Sci USA 170:8063–8070
Mazumder B, Seshadri V, Fox PL (2003) Translational control by the 3′ UTR: the ends specify the means. Trends Biochem Sci 28:91–98
Nishizawa A, Yabuta Y, Yoshida E, Maruta T, Yoshimura K, Shigeoka S (2006) Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J 48:535–547
Nutt P, Ziermann J, Hintz M, Neuffer B, Theisen G (2006) Capsella as a model system to study the evolutionary relevance of floral homeotic mutants. Plant Syst Evol 259:217–235
Orr W, Lu B, White TC, Robert LS, Singh J (1992) Complementary DNA sequence of a low temperature-induced Brassica napus gene with homology to the Arabidopsis thaliana kin1 gene. Plant Physiol 98:1532–1534
Ortega JL, Moguel-Esponda S, Potenza C, Conklin CF, Quintana A, Sengupta-Gopalan C (2006) The 3′ untranslated region of a soybean cytosolic glutamine synthetase (GS1) affects transcript stability and protein accumulation in transgenic alfalfa. Plant J 45:832–846
Park BS, Park YD, Kim HU, Jin YM, Kim HI (2002) BAN103, a pollen-preferential gene, from Chinese cabbage and its promoter activity. Mol Cells 14:150–157
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Scholkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Shimizu T, Kanamori Y, Furuki T, Kikawada T, Okuda T, Takahashi T, Mihara H, Sakurai M (2010) Desiccation-induced structuralization and glass formation of group 3 late embryogenesis abundant protein model peptides. Biochemistry 49:1093–1104
Simon B, Sengupta-Gopalan C (2010) The 3′ untranslated region of the two cytosolic glutamine synthetase (GS1) genes in alfalfa (Medicago sativa) regulates transcript stability in response to glutamine. Planta 232:1151–1162
Slotte T, Ceplitis A, Neuffer B, Hurka H, Lascoux M (2006) Intrageneric phylogeny of Capsella (Brassicaceae) and the origin of the tetraploid C. bursa-pastoris based on chloroplast and nuclear DNA sequences. Am J Bot 93:1714–1724
Slotte T, Holm K, McIntyre LM, Lagercrantz U, Lascoux M (2007) Differential expression of genes important for adaptation in Capsella bursa-pastoris (Brassicaceae). Plant Physiol 145:160–173
Tao P, Wang J (2012) Identification and characterization of transcripts differentially expressed during embryogenesis in Capsella bursa-pastoris. Biol Plant 56:415–421
Tao P, Liu L, Wang J (2012) Characterization of eight cytosolic sHSP genes and their expression in Capsella bursa-pastoris. Biol Plant. doi:10.1007/s10535-012-0239-3
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX_windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Vergnolle C, Vaultier M, Taconnat L, Renou J, Kader J, Zachowski A, Ruelland E (2005) The cold-induced early activation of phospholipase C and D pathways determines the response of two distinct clusters of genes in Arabidopsis cell suspensions. Plant Physiol 139:1217–1233
Vroh BI, McMullen MD, Sanchez-Villeda H, Schroeder S, Gardiner J, Polacco M, Soderlund C, Wing R, Fang Z, Coe EH Jr (2006) Single nucleotide polymorphisms and insertion-deletions for genetic markers and anchoring the maize fingerprint contig physical map. Crop Sci 46:12–21
Wachter A, Tunc-Ozdemir M, Grove BC, Green PJ, Shintani DK, Breaker RR (2007) Riboswitch control of gene expression in plants by splicing and alternative 3′ end processing of mRNAs. Plant Cell 19:3437–3450
Wang H, Cutler AJ (1995) Promoters from kinl and cor6.6, two Arabidopsis thaliana low-temperature- and ABA-inducible genes, direct strong p-glucuronidase expression in guard cells, pollen and young developing seeds. Plant Mol Biol 28:619–634
Wang H, Datla R, Georges F, Loewen M, Cutler AJ (1995) Promoters from kin1 and cor6.6, two homologous Arabidopsis thaliana genes: transcriptional regulation and gene expression induced by low temperature, ABA, osmoticum and dehydration. Plant Mol Biol 28:605–617
Xiong L, Lee H, Ishitani M, Zhu J (2002) Regulation of osmotic stress-responsive gene expression by the LOS6/ABA1 locus in Arabidopsis. J Biol Chem 277:8588–8596
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31070204, 30570112), National Found for Fostering Talents of Basic Sciences (J1103513) and the Chinese 111 Project.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Judelson.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tao, P., Peng, L., Huang, X. et al. Comparative analysis of the variable 3′ UTR and gene expression of the KIN and KIN-homologous LEA genes in Capsella bursa-pastoris . Plant Cell Rep 31, 1769–1777 (2012). https://doi.org/10.1007/s00299-012-1290-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-012-1290-4