Abstract
WRKY transcription factors participate in diverse physiological and developmental processes in plants. They have highly conserved WRKYGQK amino acid sequences in their N-termini, followed by the novel zinc-finger-like motifs, Cys2His2 or Cys2HisCys. To date, numerous WRKY genes have been identified and characterized in a number of herbaceous species. Survey and characterization of WRKY genes in a ligneous species would facilitate a better understanding of the evolutionary processes and functions of this gene family. In this study, 104 poplar WRKY genes (PtWRKY) were identified in the latest poplar genome sequence. According to their structural features, the predicted members were divided into the previously defined groups I–III, as described in rice. In addition, chromosomal localization of the genes demonstrated that there might be WRKY gene hot spots in 2.3 Mb regions on chromosome 14. Furthermore, approximately 83% (86 out of 104) WRKY genes participated in gene duplication events, including 69% (29 out of 42) gene pairs which exhibited segmental duplication. Using semi-quantitative RT-PCR, the expression patterns of subgroup III genes were investigated under different stresses [cold, drought, salinity and salicylic acid (SA)]. The data revealed that these genes presented different expression levels in response to various stress conditions. Expression analysis exhibited PtWRKY76 gene induced markedly in 0.1 mM SA or 25% PEG-6000 treatment. The results presented here provide a fundamental clue for cloning specific function genes in further studies and applications.
Key message This study identified 104 poplar WRKY genes and demonstrated WRKY gene hot spots on chromosome 14. Furthermore, semi-quantitative RT-PCR showed variable stress responses in subgroup III.








Similar content being viewed by others
References
Babu MM, Iyer LM, Balaji S, Aravind L (2006) The natural history of the WRKY-GCM1 zinc fingers and the relationship between transcription factors and transposons. Nucleic Acids Res 34:6505–6520
Brand LH, Kirchler T, Hummel S, Chaban C, Wanke D (2010) DPI-ELISA: a fast and versatile method to specify the binding of plant transcription factors to DNA in vitro. Plant Methods 6:25. doi:10.1186/1746-4811-6-25
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10. doi:10.1186/1471-2229-4-10
Chen CH, Chen ZX (2000) Isolation and characterization of two pathogen- and salicylic acid-induced genes encoding WRKY DNA-binding proteins from tobacco. Plant Mol Biol 42:387–396
Chothia C, Gough J, Vogel C, Teichmann SA (2003) Evolution of the protein repertoire. Science 300:1701–1703
Ciolkowski I, Wanke D, Birkenbihl RP, Somssich IE (2008) Studies on DNA-binding selectivity of WRKY transcription factors lend structural clues into WRKY-domain function. Plant Mol Biol 68:81–92
Cormack RS, Eulgem T, Rushton PJ, Kochner P, Hahlbrock K, Somssich IE (2002) Leucine zipper-containing WRKY proteins widen the spectrum of immediate early elicitor-induced WRKY transcription factors in parsley. Biochim Biophys Acta 1576:92–100
Dellagi A, Helibronn J, Avrova AO, Montesano M, Palva ET, Stewart HE, Toth IK, Cooke DE, Lyon GD, Birch PR (2000) A potato gene encoding a WRKY-like transcription factor is induced in interactins with Erwinia carotovora subsp. atroseptica and Phytophthora infestans and is coregulated with class I endochitinase expression. Mol Plant Microbe Interact 13:1092–1101
Du L, Chen Z (2000) Identification of genes encoding receptor-like protein kinases as possible targets of pathogen- and salicylic acid-induced WRKY DNA-binding proteins in Arabidopsis. Plant J 24:837–847
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5:199–206
Giacomelli JI, Ribichich KF, Dezar CA, Chan RL (2010) Expression analyses indicate the involvement of sunflower WRKY transcription factors in stress responses, and phylogenetic reconstructions reveal the existence of a novel clade in the Asteraceae. Plant Sci 178:398–410
Gu Z, Cavalcanti A, Chen FC, Bouman P, Li WH (2002) Extent of gene duplication in the genomes of Drosophila, nematode, and yeast. Mol Biol Evol 19:256–262
Guillaumie S, Mzid R, Mechin V, Leon C, Hichri I, Destrac-Irvine A, Trossat-Magnin C, Delrot S, Lauvergeat V (2010) The grapevine transcription factor WRKY2 influences the lignin pathway and xylem development in tobacco. Plant Mol Biol 72:215–234
Guo ZJ, Kan YC, Chen XJ, Li DB, Wang DW (2004) Characterization of a rice WRKY gene whose expression is induced upon pathogen attack and mechanical wounding. Acta Botanica Sinica 46:955–964
Hara K, Yagi M, Kusano T, Sano H (2000) Rapid systemic accumulation of transcripts encoding a tobacco WRKY transcription factor upon wounding. Mol Gen Genet 263:30–37
Hinderhofer K, Zentgraf U (2001) Identification of a transcription factor specifically expressed at the onset of leaf senescence. Planta 213:469–473
Holub EB (2001) The arms race is ancient history in Arabidopsis, the wildflower. Nat Rev Genet 2:516–527
Hwang DJ, Jang JY, Choi CH (2010) The WRKY superfamily of rice transcription factors. Plant Pathol J 26:110–114
Ishiguro S, Nakamura K (1994) Characterization of a cDNA encoding a novel DNA-binding protein, SPF1, that recognizes SP8 sequences in the 5′ upstream regions of genes coding for sporamin and beta-amylase from sweet potato. Mol Gen Genet 244:563–571
Izaguirre MM, Scopel AL, Baldwin IT, Ballare CL (2003) Convergent responses to stress. Solar ultraviolet-B radiation and Manduca sexta herbivory elicit overlapping transcriptional responses in field-grown plants of Nicotiana longiflora. Plant Physiol 132:1755–1767
Johnson CS, Kolevski B, Smyth DR (2002) TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 14:1359–1375
Kumar R, Tyagi AK, Sharma AK (2011) Genome-wide analysis of auxin response factor (ARF) gene family from tomato and analysis of their role in flower and fruit development. Mol Genet Genom 285:245–260
Landschulz WH, Johnson PF, McKnight SL (1988) The leucine zipper: a hypothetical structure common to a new class of DNA binding proteins. Science 240:1759–1764
Levee V, Major I, Levasseur C, Tremblay L, MacKay J, Seguin A (2009) Expression profiling and functional analysis of Populus WRKY23 reveals a regulatory role in defense. New Phytol 184:48–70
Li J, Brader G, Palva ET (2004) The WRKY70 transcription factor: a node of convergence for jasmonate-mediated and salicylate-mediated signals in plant defense. Plant Cell 16:319–331
Liu JJ, Ekramoddoullah AKM (2009) Identification and characterization of the WRKY transcription factor family in Pinus monticola. Genome 52:77–88
Liu Y, Schiff M, Dinesh-Kumar SP (2004) Involvement of MEK1 MAPKK, NTF6 MAPK, WRKY/MYB transcription factors, COI1 and CTR1 in N-mediated resistance to tobacco mosaic virus. Plant J 38:800–809
Liu Y, Jiang HY, Chen WJ, Qian YX, Ma Q, Cheng BJ, Zhu SW (2011) Genome-wide analysis of the auxin response factor (ARF) gene family in maize (Zea mays). Plant Growth Regul 63:225–234
Mangelsen E, Kilian J, Berendzen KW, Kolukisaoglu UH, Harter K, Jansson C, Wanke D (2008) Phylogenetic and comparative gene expression analysis of barley (Hordeum vulgare) WRKY transcription factor family reveals putatively retained functions between monocots and dicots. BMC Genom 9:194
Martin C, Paz-Ares J (1997) MYB transcription factors in plants. Trends Genet 13:67–73
McInerney EM, Rose DW, Flynn SE, Westin S, Mullen TM, Krones A, Inostroza J, Torchia J, Nolte RT, Assa-Munt N, Milburn MV, Glass CK, Rosenfeld MG (1998) Determinants of coactivator LXXLL motif specificity in nuclear receptor transcriptional activation. Genes Dev 12:3357–3368
Ohno S, Wolf U, Atkin NB (1968) Evolution from fish to mammals by gene duplication. Hereditas 59:169–187
Park CY, Lee JH, Yoo JH, Moon BC, Choi MS, Kang YH, Lee SM, Kim HS, Kang KY, Chung WS, Lim CO, Cho MJ (2005) WRKY group IId transcription factors interact with calmodulin. FEBS Lett 579:1545–1550
Pnueli L, Hallak-Herr E, Rozenberg M, Cohen M, Goloubinoff P, Kaplan A, Mittler R (2002) Molecular and biochemical mechanisms associated with dormancy and drought tolerance in the desert legume Retama raetam. Plant J 31:319–330
Ralph S, Oddy C, Cooper D, Yueh H et al (2006) Genomics of hybrid poplar (Populus trichocarpa × deltoides) interacting with forest tent caterpillars (Malacosoma disstria): normalized and full-length cDNA libraries, expressed sequence tags, and a cDNA microarray for the study of insect-induced defences in poplar. Mol Ecol 15:1275–1297
Ramamoorthy R, Jiang SY, Kumar N, Venkatesh PN, Ramachandran S (2008) A comprehensive transcriptional profiling of the WRKY gene family in rice under various abiotic and phytohormone treatments. Plant Cell Physiol 49:865–879
Robatzek S, Somssich IE (2002) Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev 16:1139–1149
Rushton PJ, Torres JT, Parniske M, Wernert P, Hahlbrock K, Somssich IE (1996) Interaction of elicitor-induced DNA-binding proteins with elicitor response elements in the promoters of parsley PR1 genes. EMBO J 15:5690–5700
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15:247–258
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Sun C, Palmqvist S, Olsson H, Boren M, Ahlandsberg S, Jansson C (2003) A novel WRKY transcription factor, SUSIBA2, participates in sugar signaling in barley by binding to the sugar-responsive elements of the iso1 promoter. Plant Cell 15:2076–2092
Taylor G (2002) Populus: arabidopsis for forestry. Do we need a model tree? Ann Bot 90:681–689
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tiwari SB, Hagen G, Guilfoyle TJ (2004) Aux/IAA proteins contain a potent transcriptional repression domain. Plant Cell 16:533–543
Vandenabeele S, Van Der Kelen K, Dat J, Gadjev I, Boonefaes T, Morsa S, Rottiers P, Slooten L, Van Montagu M, Zabeau M, Inze D, Van Breusegem F (2003) A comprehensive analysis of hydrogen peroxide-induced gene expression in tobacco. Proc Natl Acad Sci USA 100:16113–16118
Vision TJ, Brown DG, Tanksley SD (2000) The origins of genomic duplications in Arabidopsis. Science 290:2114–2117
Wenck AR, Conger BV, Trigiano RN, Sams CE (1988) Inhibition of somatic embryogenesis in orchardgrass by endogenous cytokinins. Plant Physiol 88:990–992
Wu KL, Guo ZJ, Wang HH, Li J (2005) The WRKY family of transcription factors in rice and Arabidopsis and their origins. DNA Res 12:9–26
Xie Z, Zhang ZL, Zou X, Huang J, Ruas P, Thompson D, Shen QJ (2005) Annotations and functional analyses of the rice WRKY gene superfamily reveal positive and negative regulators of abscisic acid signaling in aleurone cells. Plant Physiol 137:176–189
Xiong Y, Liu T, Tian C, Sun S, Li J, Chen M (2005) Transcription factors in rice: a genome-wide comparative analysis between monocots and eudicots. Plant Mol Biol 59:191–203
Xu YH, Wang JW, Wang S, Wang JY, Chen XY (2004) Characterization of GaWRKY1, a cotton transcription factor that regulates the sesquiterpene synthase gene (+)-delta-cadinene synthase-A. Plant Physiol 135:507–515
Yang S, Zhang X, Yue JX, Tian D, Chen JQ (2008) Recent duplications dominate NBS-encoding gene expansion in two woody species. Mol Genet Genom 280:187–198
Yu D, Chen C, Chen Z (2001) Evidence for an important role of WRKY DNA binding proteins in the regulation of NPR1 gene expression. Plant Cell 13:1527–1540
Yu DQ, Qiu YP, Jing SJ, Fu J, Li L (2004) Cloning and analysis of expression profile of 13 WRKY genes in rice. Chin Sci Bull 49:2159–2168
Zhang Y, Wang L (2005) The WRKY transcription factor superfamily: its origin in eukaryotes and expansion in plants. BMC Evol Biol 5:1. doi:10.1186/1471-2148-5-1
Zheng HQ, Lin SZ, Zhang Q, Lei Y, Hou L, Zhang ZY (2010) Functional identification and regulation of the PtDrl02 gene promoter from triploid white l. Plant Cell Rep 29:449–460
Acknowledgments
This work was supported by the Anhui Major Science and Technology Projects (No.08010302073) and National Science Technology Support Program (No.2009BADA6B06-1). We thank the members of the Laboratory of Modern Biotechnology and especially thank Daqiang Wu, Guo Wei and Yang Zhao for their skillful technical assistance in this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Q. Zhao.
Electronic supplementary material
Below is the link to the electronic supplementary material.
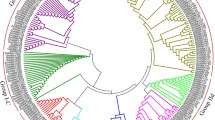
Fig. S1Phylogram based on Arabidopsis and poplar WRKY domains. The alignment of amino acid sequences was produced using the MEGA v4.0 program with the neighbor-joining (NJ) method. AtWRKY domain sequences were obtained from Wu et al. (2005).
Fig. S2Phylogenetic tree of triploid white poplar (( P. tomentosa x P. bolleana ) x P. tomentosa ), selected Arabidopsis and rice WRKY domains. The unrooted tree was constructed using MEGA 4.0 with the Neighbor-Joining (NJ) method. Clades of the WRKY domain are labeled according to the classifications of OsWRKY and AtWRKY domains by Wu et al. (2005). WRKY protein sequences of triploid white poplar were obtained from NCBI. Bootstrap values (≥500) based on 1,000 replications are exhibited beside the nodes.
Rights and permissions
About this article
Cite this article
He, H., Dong, Q., Shao, Y. et al. Genome-wide survey and characterization of the WRKY gene family in Populus trichocarpa . Plant Cell Rep 31, 1199–1217 (2012). https://doi.org/10.1007/s00299-012-1241-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-012-1241-0