Abstract
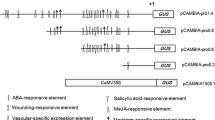
A 1,993 bp region upstream of the gene encoding the betaine aldehyde dehydrogenase (BADH) was isolated from Suaeda liaotungensis K., and the analysis of the promoter sequence has revealed the existence of several putative cis-elements by the PLACE database. In this study, according to the characteristic of the BADH promoter, five chimeric constructs varied in the length of promoter fragments from −1,993, −1,466, −1,084, −573 and −300 to +62 bp relative to the transcriptional start site were placed to the upstream of the β-glucuronidase (GUS) coding region and transferred to Nicotiana tabacum L.cv.89 by Agrobacterium tumefaciens-mediated leaf-disc transformation. The functional properties of each promoter fragment were examined by GUS histochemical staining and fluorescence quantitative analyses in the transgenic tobacco leaves treated with different NaCl concentrations for 48 h. The results show that healthy transgenic plants had decreased GUS activity in leaves, whereas a higher GUS activity was observed when the transgenic plants were challenged with sodium chloride (NaCl). Induction levels were proportional to the concentration of NaCl treatment, allowing fine-tuning of protein expression. GUS enzyme activity was enhanced 6.3-fold in transgenic tobacco leaves containing −300 bp promoter fragment in the presence of 400 mmol/l NaCl compared to the noninductive leaves. This suggests that the smallest promoter fragment (−300 to +62 bp) possesses all the essential cis-acting elements and is sufficient for NaCl induction.





Similar content being viewed by others
Abbreviations
- BADH:
-
Betaine aldehyde dehydrogenase
- CaMV:
-
Cauliflower mosaic virus
- GUS:
-
β-Glucuronidase
- MUG:
-
4-Methylumbelliferyl-β-d-glucuronide
- 4-MU:
-
4-Methylumbelliferone
- X-gluc:
-
5-Bromo-4-chloro-3-indolyl-β-d-glucuronide
References
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15(1):63–78. doi:10.1105/tpc.006
Agarwal M, Hao YJ, Kapoor A, Dong CH, Fujii H, Zheng XW, Zhu JK (2006) A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J Biol Chem 281(49):37636–37645. doi:10.1074/jbc.M605895200
Bastola DR, Pethe VV, Winicov I (1998) Alfin1, a novel zinc-finger protein in alfalfa roots that binds to promoter elements in the salt-inducible MsPRP2 gene. Plant Mol Biol 38(6):1123–1135. doi:10.1023/A:1006081926699
Boyer JS (1982) Plant productivity and environment. Science 218(4571):443–448. doi:10.1126/science.218.4571.443
Bradfork MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254. doi:10.1016/0003-2697(76)90527-3
Brouquisse R, Weigel P, Rhodes D, Yocum CF, Hanson AD (1989) Evidence for a ferredoxin-dependent choline monooxygenase from spinach chloroplast stroma. Plant Physiol 90(1):322–329. doi:0032-0889/89/90/0322/08/$01.00/0
Chinnusamy V, Ohta M, Kanrar S, Lee B, Hong XH, Agarwal M, Zhu K (2003) ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis. Genes Dev 17(8):1043–1054. doi:10.1101/gad.1077503
Chinnusamy V, Schumaker K, Zhu JK (2004) Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants. J Exp Bot 55(395):225–236. doi:10.1093/jxb/erh005
Hanson AD, Rathinasabapathi B, Chamberlin B, Gage DA (1991) Comparative physiological evidence that beta-alanine betaine and choline-O-sulfate act as compatible osmolytes in halophytic Limonium species. Plant Physiol 97(3):1199–1205
Hibino T, Meng YL, Kawamitsu Y, Uehara N, Matsuda N, Tanaka Y, Ishikawa H, Baba S, Takabe T, Wada K, Ishii T, Takabe T (2001) Molecular cloning and functional characterization of two kinds of betaine-aldehyde dehydrogenase in betaine-accumulating mangrove Avicennia marina (Forsk.) Vierh. Plant Mol Biol 45(3):353–363. doi:10.1023/A:1006497113323
Ishitani M, Nakamura T, Han SY, Takabe T (1995) Expression of the betaine aldehyde dehydrogenase gene in barley in response to osmotic stress and abscisic acid. Plant Mol Biol 27(2):307–315. doi:10.1007/BF00020185
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6(13):3901–3907
Jia GX, Zhu ZQ, Chang FQ, Li YX (2002) Transformation of tomato with the BADH gene from Atriplex improves salt tolerance. Plant Cell Rep 21(2):141–146. doi:10.1007/s00299-002-0489-1
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1999) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17(3):287–291. doi:10.1038/7036
Legaria J, Rajsbaum R, Muñoz-Clares RA, Villegas-Sepúlveda N, Simpson J, Iturriaga G (1998) Molecular characterization of two genes encoding betaine aldehyde dehydrogenase from amaranth. Expression in leaves under short-term exposure to osmotic stress of abscisic acid. Gene 218(1–2):69–76. doi:10.1016/S0378-1119(98)00381-3
Li QL, Gao XR, Yu XH, Wang XZ, An LJ (2003) Molecular cloning and characterization of betaine aldehyde dehydrogenase gene from Suaeda liaotungensis and its use in improved tolerance to salinity in transgenic tobacco. Biotechnol Lett 25(17):1431–1436. doi:10.1023/A:1025003628446
Li QL, Zhang Y, Yin H, Li D (2006) Isolation of BADH gene promoter from Suaeda liaotungensis and its sequence analysis. Sheng Wu Gong Cheng Xue Bao 22(1):77–81
Malnoy M, Venisse JS, Reynoird JP, Chevreau E (2003) Activation of three pathogen-inducible promoters of tobacco in transgenic pear (Pyrus communis L.) after abiotic and biotic elicitation. Planta 216(5):802–814. doi:10.1007/s00425-002-0932-0
McCue KF, Hanson AD (1992) Salt-inducible betaine aldehyde dehydrogenase from sugar beet: cDNA cloning and expression. Plant Mol Biol 18(1):1–11. doi:10.1007/BF00018451
Meng CX, Wei W, Su Z, Qin S (2006) Characterization of carotenoid hydroxylase gene promoter in Haematococcus pluvialis. Indian J Biochem Biophys 43(5):284–288
Nakamura T, Yokota S, Muramoto Y, Tsutsui K, Oguri Y, Fukui K, Takabe T (1997) Expression of a betaine aldehyde dehydrogenase gene in rice, a glycinebetaine nonaccumulator, and possible location of its protein in peroxisomes. Plant J 11(5):1115–1120
Narusaka Y, Nakashima K, Shinwari ZK, Sakuma Y, Furihata T, Abe H, Narusaka M, Shinozaki K, Yanguchi-Shinozaki K (2003) Interaction between two cis-element, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J 34:137–148
Papageorgiou GC, Murata N (1995) The unusually strong stabilizing effects of glycine betaine on the structure and function in the oxygen-evolving photosystem II complex. Photosynth Res 44(3):243–252. doi:10.1007/BF00048597
Park HC, Kim ML, Kang YH, Jeon JM, Yoo JH, Kim MC, Park CY, Jeong JC, Moon BC, Lee JH, Yoon HW, Lee SH, Chung WS, Lim CO, Lee JH, Hong JC, Cho MJ (2004) Pathogen- and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol 135(4):2150–2161
Pastuglia M, Roby D, Dumas C, Cockagi JM (1997) Rapid induction by wounding and bacterial infection of an S gene family receptor-like kinase gene in Brassica oleracea. Plant Cell 9:49–60
Rathinasabapathi B, Fouad WM, Sigua CA (2001) β-Alanine betaine synthesis in the Plumbaginaceae. Purification and characterization of a trifunctional, S-adenosyl-l-methionine-dependent N-methyltransferase from Limonium latifolium leaves. Plant Physiol 126(3):1241–1249
Rhodes D, Hanson AD (1993) Quaternary ammonium and tertiary sulfonium compounds in higher plants. Annu Rev Plant Physiol 44:357–384. doi:10.1146/annurev.pp.44.060193.002041
Robinson SP, Jones GP (1986) Accumulation of glycinebetaine in chloroplasts provides osmotic adjustment during salt stress. Aust J Plant Physiol 13:659–668
Uno Y, Furihata T, Yoshida HAR, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97(21):11632–11637
Weigel P, Weretilnyk EA, Hanson AD (1986) Betaine aldehyde oxidation by spinach chloroplasts. Plant Physiol 82(3):753–759
Weretilnyk EA, Hanson AD (1990) Molecular cloning of a plant betaine-aldehyde dehydrogenase, an enzyme implicated in adaptation to salinity and drought. Proc Natl Acad Sci USA 87(7):2745–2749
Wood AJ, Saneoka H, Rhodes D, Joly RJ, Goldsbrough PB (1996) Betaine aldehyde dehydrogenase in Sorghum. Plant Physiol 110(4):1301–1308
Xiao G, Zhang GY, Liu FH, Wang J, Chen SY (1995) The study of BADH gene in Atriplex hortensis. Chin Sci Bull 40:741–745
Yin XJ, Zhao YX, Luo D, Zhang H (2002a) Expression of the betaine aldehyde dehydrogenase (AcBADH) gene and isolation of its promoter from the halophyte Atriplex centralasiatica Iljin. J Plant Physiol Mol Biol 28(6):479–484
Yin XJ, Zhao YX, Luo D, Zhang H (2002b) Isolating the promoter of a stress-induced gene encoding betaine aldehyde dehydrogenase from the halophyte Atriplex centralasiatica Iljin. Biochim Biophys Acta 1577(3):452–456. doi:10.1016/S0167-4781(02)00495-5
Acknowledgments
This work was supported by the National Natural Science Foundation of China (no. 30370806), the Science and Technology Foundation of Liaoning Province (no. 20031060) and the Educational Department of Liaoning Province (no. 2004F091).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Register.
Rights and permissions
About this article
Cite this article
Zhang, Y., Yin, H., Li, D. et al. Functional analysis of BADH gene promoter from Suaeda liaotungensis K.. Plant Cell Rep 27, 585–592 (2008). https://doi.org/10.1007/s00299-007-0459-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-007-0459-8