Abstract
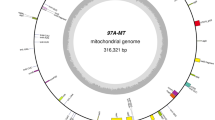
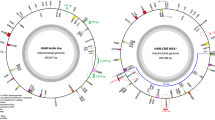
To study genetic relatedness of two male sterility-inducing cytotypes, the phylogenetic relationship among three cytotypes of onions (Allium cepa L.) was assessed by analyzing polymorphisms of the mitochondrial DNA organization and chloroplast sequences. The atp6 gene and a small open reading frame, orf22, did not differ between the normal and CMS-T cytotypes, but two SNPs and one 4-bp insertion were identified in CMS-S cytotype. Partial sequences of the chloroplast ycf2 gene were integrated in the upstream sequence of the cob gene via short repeat sequence-mediated recombination. However, this chloroplast DNA-integrated organization was detected only in CMS-S. Interestingly, disruption of a group II intron of cox2 was identified for the first time in this study. Like other trans-splicing group II introns in mitochondrial genomes, fragmentation of the intron occurred in domain IV. Two variants of each exon1 and exon2 flanking sequences were identified. The predominant types of four variants were identical in both the normal and the CMS-T cytotypes. These predominant types existed as sublimons in CMS-S cytotypes. Altogether, no differences were identified between normal and CMS-T, but significant differences in gene organization and nucleotide sequences were identified in CMS-S, suggesting recent origin of CMS-T male-sterility from the normal cytotype.






Similar content being viewed by others
References
Abdelnoor RV, Yule R, Elo A, Christensen AC, Meyer-Gauen G, Mackenzie SA (2003) Substoichiometric shifting in the plant mitochondrial genome is influenced by a gene homologous to MutS. Proc Natl Acad Sci 100:5968–5973
Abdelnoor RV, Christensen AC, Mohammed S, Munoz-Castillo B, Moriyama H, Mackenzie SA (2006) Mitochondrial genome dynamics in plants and animals: convergent gene fusions of a MutS homologue. J Mol Evol 63:165–173
Albert B, Godelle B, Gouyon PH (1998) Evolution of the plant mitochondrial genome: dynamics of duplication and deletion of sequences. J Mol Evol 46:155–158
Allen JO, Fauron CM, Mink P, Roark L, Oddiraju S, Lin GN, Meyer L, Sun H, Kim K, Wang C, Du F, Xu D, Gibson M, Cifrese J, Clifton SW, Newton KJ (2007) Comparisons among two fertile and three male-sterile mitochondrial genomes of maize. Genetics 177:1173–1192
Arrieta-Montiel M, Lyznik A, Woloszynska M, Janska H, Tohme J, Mackenzie S (2001) Tracing evolutionary and developmental implications of mitochondrial stoichiometric shifting in the common bean. Genetics 158:851–864
Backert S, Neilsen BL, Börner T (1997) The mystery of the rings: structure and replication of mitochondrial genomes from higher plants. Trend Plant Sci 2:477–483
Bellaoui M, Martin-Canadell A, Pelletier G, Budar F (1998) Low-copy-number molecules are produced by recombination, actively maintained and can be amplified in the mitochondrial genome of Brassicaceae: relationship to reversion of the male sterile phenotype in some cybrids. Mol Gen Genet 257:177–185
Berninger E (1965) Contribution à l’étude de la sterilité mâle de l’oignon (Allium cepa L.). Ann Amélior Plant 15:183–199
Bonen L (2008) Cis- and trans-splicing of group II introns in plant mitochondria. Mitochondrion 8:26–34
Budar F, Touzet P, De Paepe R (2003) The nucleo-mitochondrial conflict in cytoplasmic male sterilities revised. Genetica 117:3–16
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Engelke T, Terefe D, Tatlioglu T (2003) A PCR-based marker system monitoring CMS-(S), CMS-(T) and (N)-cytoplasm in the onion (Allium cepa L.). Theor Appl Genet 107:162–167
Gao LM, Möller M, Zhang XM, Hollingsworth ML, Liu J, Mill RR, Gibby M, Li DZ (2007) High variation and strong phylogeographic pattern among cpDNA haplotypes in Taxus wallichiana (Taxaceae) in China and North Vietnam. Mol Ecol 16:4684–4698
Glanz S, Kück U (2009) Trans-splicing of organelle introns—a detour to continuos RNAs. Bioessays 31:921–934
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Window 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Handa H (2003) The complete nucleotide sequence and RNA editing content of the mitochondrial genome of rapeseed (Brassica napus L.): comparative analysis of the mitochondrial genomes of rapeseed and Arabidopsis thaliana. Nucleic Acids Res 31:5907–5916
Hanson MR, Bentolila S (2004) Interactions of mitochondrial and nuclear genes that affect male gametophyte development. Plant Cell 16:S154–S169
Havey MJ (1995) Identification of cytoplasms using the polymerase chain reaction to aid in the extraction of maintainer lines from open-pollinated populations of onion. Theor Appl Genet 90:263–268
Havey MJ (2000) Diversity among male-sterility-inducing and male-fertile cytoplasms of onion. Theor Appl Genet 101:778–782
Janska H, Sarria R, Woloszynska M, Arrieta-Montiel M, Mackenzie SA (1998) Stoichiometric shifts in the common bean mitochondrial genome leading to male sterility and spontaneous reversion to fertility. Plant Cell 10:1163–1180
Jones HA, Clarke A (1943) Inheritance of male sterility in the onion and the production of hybrid seed. Proc Am Soc Hortic Sci 43:189–194
Jones HA, Emsweller SL (1936) A male-sterile onion. Proc Am Soc Hortic Sci 34:582–585
Kanazawa A, Tsutsumi N, Hirai A (1994) Reversible changes in the composition of the population of mtDNAs during dedifferentiation and regeneration in tobacco. Genetics 138:865–870
Kim S, Lim H, Park S, Cho K, Sung S, Oh D, Kim K (2007) Identification of a novel mitochondrial genome type and development of molecular makers for cytoplasm classification in radish (Raphanus sativus L.). Theor Appl Genet 115:1137–1145
Kim S, Lee E, Cho DY, Han T, Bang H, Patil BS, Ahn YK, Yoon M (2009a) Identification of a novel chimeric gene, orf725. and its use in development of a molecular marker for distinguishing among three cytoplasm types in onion (Allium cepa L.). Theor Appl Genet 118:433–441
Kim S, Lee Y, Lim H, Ahn Y, Sung S (2009b) Identification of highly variable chloroplast sequences and development of cpDNA-based molecular markers that distinguish four cytoplasm types in radish (Raphanus sativus L.). Theor Appl Genet 119:189–198
Kmiec B, Woloszynska M, Janska H (2006) Heteroplasmy as a common state of mitochondrial genetic information in plants and animals. Curr Genet 50:149–159
Knoop V (2004) The mitochondrial DNA of land plants: peculiarities in phylogenetic perspective. Curr Genet 46:123–139
Kubo T, Newton KJ (2008) Angiosperm mitochondrial genomes and mutations. Mitochondrion 8:5–14
Kubo T, Nishizawa S, Sugawara A, Itchoda N, Estiati A, Mikami T (2000) The complete nucleotide sequence of the mitochondrial genome of sugar beet (Beta vulgaris L.) reveals a novel gene for tRNAcys(GCA). Nucleic Acids Res 28:2571–2576
Kudla J, Albertazzi FJ, Blazevic D, Hermann M, Bock R (2002) Loss of the mitochondrial cox2 intron 1 in a family of monocotyledonous plants and utilization of mitochondrial intron sequences for the construction of a nuclear intron. Mol Genet Genomics 267:223–230
Lee Y, Park S, Lim C, Kim H, Lim H, Ahn Y, Sung S, Yoon M, Kim S (2008) Discovery of a novel cytoplasmic male-sterility and its restorer lines in radish (Raphanus sativus L.). Theor Appl Genet 117:905–913
Mackenzie SA, Chase CD (1990) Fertility restoration is associated with loss of a portion of the mitochondrial genome in cytoplasmic male-sterile common bean. Plant Cell 2:905–912
Meng L, Yang R, Abbott RJ, Miehe G, Hu T, Liu J (2007) Mitochondrial and chloroplast phylogeography of Picea crassifolia Kom. (Pinaceae) in the Qinghai-Tibetan plateau and adjacent highlands. Mol Ecol 16:4128–4137
Notsu Y, Masood S, Nishikawa T, Kubo N, Akiduki G, Nakazono M, Kirai A, Kadowaki K (2002) The complete sequence of the rice (Oryza sativa L.) mitochondrial genome: frequent DNA sequence acquisition and loss during the evolution of flowering plants. Mol Genet Genomics 268:434–445
Ohsako T, Wang G, Miyashita NT (1996) Polymerase chain reaction-single strand conformational polymorphism analysis of intra- and interspecific variations in organellar DNA regions of Aegilops mutica and related species. Genes Genet Syst 71:281–292
Oldenburg DJ, Bendich AJ (2001) Mitochondrial DNA from the Liverwort Marchantia polymorpha: circularly permuted linear molecules, head-to-tail concatemers, and a 5′ protein. J Mol Biol 310:549–562
Palmer JD (1988) Intraspecific variation and multicircularity in Brassica mitochondrial DNAs. Genetics 118:341–351
Palmer JD, Herbon LA (1987) Unicircular structure of the Brassica hirta mitochondrial genome. Curr Genet 11:565–570
Pyle AM, Fedorova O, Waldsich C (2007) Folding of group II introns: a model system for large, multidomain RNAs? Trend Biochem Sci 32:138–145
Qiu Y, Palmer JD (2004) Many independent origins of trans splicing of a plant mitochondrial group II intron. J Mol Evol 59:80–89
Reboud X, Zeyl C (1993) Organelle inheritance in plants. Heredity 72:132–140
Sakai T, Imamura J (1993) Evidence for a mitochondrial sub-genome containing radish Atp1 in a Brassica napus cybrid. Plant Sci 90:95–103
Sandhu AS, Abdelnoor RV, Mackenzie SA (2007) Transgenic induction of mitochondrial rearrangements for cytoplasmic male sterility in crop plants. Proc Natl Acad Sci 104:1766–1770
Sato Y (1998) PCR amplification of CMS-specific mitochondrial nucleotide sequences to identify cytoplasmic genotypes of onion (Allium cepa L.). Theor Appl Genet 96:367–370
Satoh M, Kubo T, Nishizawa S, Estiati A, Itchoda N, Mikami T (2004) The cytoplasmic male-sterile type and normal type mitochondrial genomes of sugarbeet share the same complement of genes of known function but differ in the content of expressed ORFs. Mol Gen Genomics 272:247–256
Schweisguth B (1973) Étude d’un nouveau type de stérilité male chez l’oignon, Allium cepa L. Ann Amélior Plant 23:221–233
Shedge V, Arrieta-Montiel M, Christensen AC, Mackenzie SA (2007) Plant mitochondrial recombination surveillance requires unusual RecA and MutS homologs. Plant Cell 19:1251–1264
Small I, Suffolk R, Leaver CJ (1989) Evolution of plant mitochondrial genomes via substoichiometric intermediates. Cell 58:69–76
Sugiyama Y, Watase Y, Nagase M, Makita N, Yagura S, Hirai A, Sugiura M (2005) The complete nucleotide sequence and multipartite organization of the tobacco mitochondrial genome: comparative analysis of mitochondrial genomes in higher plants. Mol Gen Genomics 272:603–615
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Toor N, Keating KS, Taylor SD, Pyle AM (2008) Crystal structure of a self-spliced group II intron. Science 320:77–82
Unseld M, Marienfeld JR, Brandt P, Brennicke A (1997) The mitochondrial genome of Arabidopsis thaliana contains 57 genes in 366, 924 nucleotides. Nature genet 15:57–61
Ward BL, Anderson RS, Bendich AJ (1981) The mitochondrial genome is large and variable in a family of plants (Cucurbitaceae). Cell 25:793–803
Woloszynska M, Trojanowski D (2009) Counting mtDNA molecules in Phaseolus vulgaaris: sublimons are constantly produced by recombination via short repeats and undergo rigorous selection during substoichiometric shifting. Plant Mol Biol 70:511–521
Zaegel V, Guermann B, Le Ret M, Andrés C, Meyer D, Erhardt M, Canaday J, Gualberto JM, Imbault P (2006) The plant-specific ssDNA binding protein OSB1 is involved in the stoichiometric transmission of mitochondrial DNA in Arabidopsis. Plant Cell 18:3548–3563
Acknowledgments
This study was supported by the Specific Joint Agricultural Research-Promoting Projects (Project No. 200803A01081219), RDA, Republic of Korea, and by the Biotechnology Research Institute, Chonnam National University, Republic of Korea. We thank Seon Chong and Mi-Young Kim for their dedicated technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Bock.
Rights and permissions
About this article
Cite this article
Kim, S., Yoon, MK. Comparison of mitochondrial and chloroplast genome segments from three onion (Allium cepa L.) cytoplasm types and identification of a trans-splicing intron of cox2 . Curr Genet 56, 177–188 (2010). https://doi.org/10.1007/s00294-010-0290-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-010-0290-6