Abstract
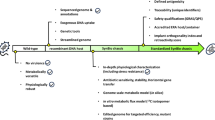
A genome-shuffled Stenotrophomonas maltophilia strain showing the enhanced ability of RDX degradation was constructed, and its characteristics were compared with those of the wild-type one. The shuffled strain was able to completely degrade 25, 50, and 75 µM RDX (hexahydro-1,3,5-trinitro-1,3,5-triazine) within 10, 30, and 50 days, respectively. However, it took 30 and 70 days for the wild-type strain to degrade 25 and 50 µM RDX, respectively, and at day 70, the strain degraded only 67% of 75 µM RDX. The shuffled strain reached its maximum growth at 50–60 days and exhibited approximately 1.5-fold increased cell numbers. SEM revealed more severe damage on the surface of the wild-type cells compared to the genome-shuffled cells. The mRNA levels of dnaK and groEL encoding the heat shock proteins were increased by 2.5-fold and fourfold, and DnaK and GroEL proteins were more highly produced in the shuffled cells. In addition, the mRNA levels of pnrB encoding a TNT nitroreductase, and algA involved in exopolymer biosynthesis, were slightly higher in the shuffled strain, but not as high as those of dnaK and groEL. These results indicate that the genome shuffling rendered the shuffled cells more resistant to RDX stress. A proteomic comparison revealed changes in the production levels of certain proteins including nitrate and cell protection, particularly those involved in metabolism. These proteomic analyses provide clues for understanding the improved RDX degradation by the genome-shuffled S. maltophilia strain.






Similar content being viewed by others
References
Best EP, Geter KN, Tatem HE, Lane BK (2006) Effects, transfer, and fate of RDX from aged soil in plants and worms. Chemosphere 62:616–625
Binks PR, Nicklin S, Bruce NC (1995) Degradation of hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) by Stenotrophomonas maltophilia PB1. Appl Environ Microbiol 61:1318–1322
Biot-Pelletier D, Martin VJ (2014) Evolutionary engineering by genome shuffling. Appl Microbiol Biotechnol 98:3877–3887
Bollag DM, Rozycki MD, Edelstein SJ (1996) Protein methods, 2nd edn. Wiley, New York
Chalopagorn P, Charoenpanich J, Choowongkomon K (2014) Genome shuffling enhances lipase production of thermophilic Geobacillus sp. Appl Biochem Biotechnol 174:1444–1454
Chang HW, Kahng HY, Kim SI, Chun JW, Oh KH (2004) Characterization of Pseudomonas sp. HK-6 cells responding to explosive RDX (hexahydro-1,3,5-trinitro-1,3,5-triazine). Appl Microbiol Biotechnol 65:323–329
Cho SH, Cho YS, Oh KH (2009) Biological treatment of TNT containing wastewater, pink water by Stenotrophomonas maltophilia OK-5, and RT-PCR quantification of the nitroreductase (pnrB) gene. Kor J Biotechnol Eng 24:556–562
Coretzee JN, Sirgel FA, Lecatsas G (1979) Genetic recombination in fused spheroplast of Providence alcalifaciens. J Gen Microbiol 114:313–322
Copley S (2009) Evolution of efficient pathways for degradation of anthropogenic chemicals. Nat Chem Biol 5:559–566
Fournier D, Halasz A, Spain J, Spanggord RJ, Bottaro JC, Hawari J (2004) Biodegradation of the hexahydro-1,3,5-trinitro-1,3,5-triazine ring cleavage product 4-nitro-2,4-diazabutanal by Phanerochaete chrysosporium. Appl Environ Microbiol 70:1123–1128
Gacesa P (1998) Bacterial alginate biosynthesis-recent progress and future prospects. Microbiology 144:1133–1143
Haas R, Schreiber I, von LöwE Stork G (1990) Conception for the investigation of contaminated munitions plants, 2: investigation of former RDX plants and filling stations. Anal Bioanal Chem 338:41–45
Heukeshoven J, Dernick R (1985) Simplified method for silver staining of proteins in polyacrylamide gels and the mechanism of silver staining. Electrophoresis 6:103–112
Ho EM, Chang HW, Kim SI, Kahng HY, Oh KH (2004) Analysis of TNT (2,4,6-trinitrotoluene)-inducible cellular responses and stress shock proteome in Stenotrophomonas sp. OK-5. Curr Microbiol 49:346–352
Hou LH, Meng M, Guo L, He JY (2015) A comparison of whole cell directed evolution approaches in breeding of industrial strain of Saccharomyces cerevisiae. Biotechnol Lett 37:1393–1398
Khan MI, Lee J, Park J (2012) Microbial degradation and toxicity of hexahydro-1,3,5-trinitro-1,3,5-triazine. J Microbiol Biotechnol 22:1321–1333
Kwon MJ, Wei N, Millerick K, Popovic J, Finneran K (2014) Clostridium geopurificans strain MJ1 sp. nov., a strictly anaerobic bacterium that grows via fermentation and reduces the cyclic nitramine explosive hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX). Curr Microbiol 68:743–750
Lee BU, Park SC, Cho YS, Kahng HY, Oh KH (2008) Expression and characterization of the TNT nitroreductase of Pseudomonas sp. HK-6 in Escherichia coli. Curr Microbiol 56:386–390
Lee BU, Park SC, Cho YS, Oh KH (2008) Exopolymer biosynthesis and proteomic changes of Pseudomonas sp. HK-6 under stress of TNT (2,4,6-trinitrotoluene). Curr Microbiol 57:477–483
Lee BU, Cho YS, Park SC, Oh KH (2009) Enhanced degradation of TNT by genome-shuffled Stenotrophomonas maltophilia OK-5. Curr Microbiol 59:346–351
Nejidat A, Kafka L, Tekoah Y, Ronen Z (2008) Effect of organic and inorganic nitrogenous compounds on RDX degradation and cytochrome P450 expression in Rhodococcus strain YH1. Biodegradation 19:313–320
Ng LK, Sherburne R, Taylor DE, Stiles ME (1985) Morphological forms and viability of Campylobacter species studied by electron microscopy. J Bacteriol 164:338–343
Pertins DN, Pappin DJ, Creasy DM, Cottrell JS (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20:3551–3567
Ramos JL, Duque E, Huertas MJ, Haidour A (1995) Isolation and expression of catabolic potential of a Pseudomonas putida strain able to grow in presence of high concentrations of aromatic hydrocarbons. J Bacteriol 177:3911–3916
Rosenblatt DH, Burrows EP, Mitchell WR, Parme DL (1991) Organic explosive and related compounds. In: Hutzinger O (ed) Handbook of environmental chemistry. Springer, Berlin, pp 195–234
Rylott EL, Budarina MV, Barker A, Lorenz A, Strand SE (2011) Engineering plants for the phytoremediation of RDX in the presence of the co-contaminating explosive TNT. New Phytol 192:405–413
Sikkema J, de Bont JA, Poolman B (1995) Mechanisms of membrane toxicity of hydrocarbons. Microbiol Rev 59:201–222
Stewart V (1982) Requirement of Fnr and NarL functions for nitrate reductase expression in Escherichia coli K-12. J Bacteriol 151:1320–1325
Yinon J (1990) Toxicity and metabolism of explosives. CRC Press, Boca Raton, Florida
Zhang YX, Perry K, Vinci VA, Powell K, Stemmer WP (2002) Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415:644–646
Zhou YP, Ren XD, Wang L, Chen XS, Mao ZG (2015) Enhancement of ε-poly-lysine production in ε-poly-lysine-tolerant Streptomyces sp. by genome shuffling. Bioprocess Biosyst Eng 38:1705–1713
Acknowledgement
This study was supported by the Basic Science Research Program through the National Research Foundation of Korea, funded by the Ministry of Education, Science, and Technology (2011-0026690), and by the Soonchunhyang University Research Fund. We thank the ROK Agency for Defense Development for providing the RDX used in this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lee, BU., Choi, MS., Kim, DM. et al. Genome Shuffling of Stenotrophomonas maltophilia OK-5 for Improving the Degradation of Explosive RDX (Hexahydro-1,3,5-trinitro-1,3,5-triazine). Curr Microbiol 74, 268–276 (2017). https://doi.org/10.1007/s00284-016-1179-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-016-1179-5