Abstract
Purpose
We aimed to evaluate the performance of deep learning-based generalization of ultra-low-count amyloid PET/MRI enhancement when applied to studies acquired with different scanning hardware and protocols.
Methods
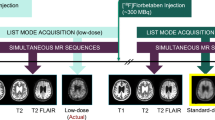
Eighty simultaneous [18F]florbetaben PET/MRI studies were acquired, split equally between two sites (site 1: Signa PET/MRI, GE Healthcare, 39 participants, 67 ± 8 years, 23 females; site 2: mMR, Siemens Healthineers, 64 ± 11 years, 23 females) with different MRI protocols. Twenty minutes of list-mode PET data (90–110 min post-injection) were reconstructed as ground-truth. Ultra-low-count data obtained from undersampling by a factor of 100 (site 1) or the first minute of PET acquisition (site 2) were reconstructed for ultra-low-dose/ultra-short-time (1% dose and 5% time, respectively) PET images. A deep convolution neural network was pre-trained with site 1 data and either (A) directly applied or (B) trained further on site 2 data using transfer learning. Networks were also trained from scratch based on (C) site 2 data or (D) all data. Certified physicians determined amyloid uptake (+/−) status for accuracy and scored the image quality. The peak signal-to-noise ratio, structural similarity, and root-mean-squared error were calculated between images and their ground-truth counterparts. Mean regional standardized uptake value ratios (SUVR, reference region: cerebellar cortex) from 37 successful site 2 FreeSurfer segmentations were analyzed.
Results
All network-synthesized images had reduced noise than their ultra-low-count reconstructions. Quantitatively, image metrics improved the most using method B, where SUVRs had the least variability from the ground-truth and the highest effect size to differentiate between positive and negative images. Method A images had lower accuracy and image quality than other methods; images synthesized from methods B–D scored similarly or better than the ground-truth images.
Conclusions
Deep learning can successfully produce diagnostic amyloid PET images from short frame reconstructions. Data bias should be considered when applying pre-trained deep ultra-low-count amyloid PET/MRI networks for generalization.






Similar content being viewed by others
References
Jack CR Jr, Bennett DA, Blennow K, Carrillo MC, Dunn B, Haeberlein SB, et al. NIA-AA research framework: toward a biological definition of Alzheimer’s disease. Alzheimers Dement. 2018;14:535–62. https://doi.org/10.1016/j.jalz.2018.02.018.
Barthel H, Schroeter ML, Hoffmann KT, Sabri O. PET/MR in dementia and other neurodegenerative diseases. Semin Nucl Med. 2015;45:224–33. https://doi.org/10.1053/j.semnuclmed.2014.12.003.
Catana C, Drzezga A, Heiss WD, Rosen BR. PET/MRI for neurologic applications. J Nucl Med. 2012;53:1916–25. https://doi.org/10.2967/jnumed.112.105346.
Drzezga A, Barthel H, Minoshima S, Sabri O. Potential clinical applications of PET/MR imaging in neurodegenerative diseases. J Nucl Med. 2014;55:47S–55S. https://doi.org/10.2967/jnumed.113.129254.
Rowe CC, Villemagne VL. Brain amyloid imaging. J Nucl Med. 2011;52:1733–40. https://doi.org/10.2967/jnumed.110.076315.
Sperling RA, Mormino EC, Schultz AP, Betensky RA, Papp KV, Amariglio RE, et al. The impact of amyloid-beta and tau on prospective cognitive decline in older individuals. Ann Neurol. 2019;85:181–93. https://doi.org/10.1002/ana.25395.
Barthel H, Gertz HJ, Dresel S, Peters O, Bartenstein P, Buerger K, et al. Cerebral amyloid-beta PET with florbetaben (18F) in patients with Alzheimer’s disease and healthy controls: a multicentre phase 2 diagnostic study. Lancet Neurol. 2011;10:424–35. https://doi.org/10.1016/S1474-4422(11)70077-1.
Villemagne VL. Selective Tau Imaging: Der Stand der Dinge. J Nucl Med. 2018;59:175–6. https://doi.org/10.2967/jnumed.117.198325.
Dickerson BC, Bakkour A, Salat DH, Feczko E, Pacheco J, Greve DN, et al. The cortical signature of Alzheimer’s disease: regionally specific cortical thinning relates to symptom severity in very mild to mild AD dementia and is detectable in asymptomatic amyloid-positive individuals. Cereb Cortex. 2009;19:497–510. https://doi.org/10.1093/cercor/bhn113.
Herholz K, Evans R, Anton-Rodriguez J, Hinz R, Matthews JC. The effect of 18F-florbetapir dose reduction on region-based classification of cortical amyloid deposition. Eur J Nucl Med Mol Imaging. 2014;41:2144–9. https://doi.org/10.1007/s00259-014-2842-3.
Tiepolt S, Barthel H, Butzke D, Hesse S, Patt M, Gertz HJ, et al. Influence of scan duration on the accuracy of beta-amyloid PET with florbetaben in patients with Alzheimer's disease and healthy volunteers. Eur J Nucl Med Mol Imaging. 2013;40:238–44. https://doi.org/10.1007/s00259-012-2268-8.
Schiller F, Frings L, Thurow J, Meyer PT, Mix M. Limits for reduction of acquisition time and administered activity in (18)F-FDG PET studies of Alzheimer dementia and frontotemporal dementia. J Nucl Med. 2019;60:1764–70. https://doi.org/10.2967/jnumed.119.227132.
Bland J, Mehranian A, Belzunce MA, Ellis S, McGinnity CJ, Hammers A, et al. MR-guided kernel EM reconstruction for reduced dose PET imaging. IEEE Trans Radiat Plasma Med Sci. 2018;2:235–43. https://doi.org/10.1109/TRPMS.2017.2771490.
Xiang L, Qiao Y, Nie D, An L, Wang Q, Shen D. Deep auto-context convolutional neural networks for standard-Dose PET image estimation from low-dose PET/MRI. Neurocomputing. 2017;267:406–16. https://doi.org/10.1016/j.neucom.2017.06.048.
Chen H Low-dose CT with a residual encoder-decoder convolutional neural network (RED-CNN). arXiv: arXiv. 2017.
Ronneberger O, Fischer P, Brox T U-Net: convolutional networks for biomedical image segmentation. arXiv. 2015.
Chen KT, Gong E, de Carvalho Macruz FB, Xu J, Boumis A, Khalighi M, et al. Ultra-low-dose (18)F-florbetaben amyloid PET imaging using deep learning with multi-contrast MRI inputs. Radiology. 2019;290:649–56. https://doi.org/10.1148/radiol.2018180940.
Chang K, Balachandar N, Lam C, Yi D, Brown J, Beers A, et al. Distributed deep learning networks among institutions for medical imaging. J Am Med Inform Assoc. 2018;25:945–54. https://doi.org/10.1093/jamia/ocy017.
Sheller MJ, Reina GA, Edwards B, Martin J, Bakas S. Multi-institutional deep learning modeling without sharing patient data: a feasibility study on brain tumor segmentation. Brainlesion. 2019;11383:92–104. https://doi.org/10.1007/978-3-030-11723-8_9.
McClure P, Zheng CY, Kaczmarzyk JR, Lee JA, Ghosh SS, Nielson D, et al. Distributed weight consolidation: a brain segmentation case study. In Proceedings of the 32nd International Conference on Neural Information Processing Systems (NIPS’18). Curran Associates Inc.: Red Hook, NY. 2018: 4097–4107.
Yune S, Lee H, Pomerantz SR, Romero JM, Kamalian S, Gonzalez RG, et al. Real-world performance of deep-learning-based automated detection system for intracranial hemorrhage. 2018 SIIM Conference on Machine Intelligence in Medical Imaging: San Francisco; 2018.
Pan SJ, Yang QA. A survey on transfer learning. IEEE Trans Knowl Data Eng. 2010;22:1345–59. https://doi.org/10.1109/Tkde.2009.191.
Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K, Kirby J, et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans Med Imaging. 2015;34:1993–2024. https://doi.org/10.1109/TMI.2014.2377694.
Guo J, Gong E, Fan AP, Goubran M, Khalighi MM, Zaharchuk G. Predicting (15)O-Water PET cerebral blood flow maps from multi-contrast MRI using a deep convolutional neural network with evaluation of training cohort bias. J Cereb Blood Flow Metab. 2019:271678X19888123. https://doi.org/10.1177/0271678X19888123.
Gatidis S, Wurslin C, Seith F, Schafer JF, la Fougere C, Nikolaou K, et al. Towards tracer dose reduction in PET studies: simulation of dose reduction by retrospective randomized undersampling of list-mode data. Hell J Nucl Med. 2016;19:15–8. https://doi.org/10.1967/s002449910333.
Iagaru A, Mittra E, Minamimoto R, Jamali M, Levin C, Quon A, et al. Simultaneous whole-body time-of-flight 18F-FDG PET/MRI: a pilot study comparing SUVmax with PET/CT and assessment of MR image quality. Clin Nucl Med. 2015;40:1–8. https://doi.org/10.1097/RLU.0000000000000611.
Ladefoged CN, Benoit D, Law I, Holm S, Kjaer A, Hojgaard L, et al. Region specific optimization of continuous linear attenuation coefficients based on UTE (RESOLUTE): application to PET/MR brain imaging. Phys Med Biol. 2015;60:8047–65. https://doi.org/10.1088/0031-9155/60/20/8047.
Chen KT, Schürer M, Ouyang J, Gong E, Tiepolt S, Sabri O, et al. How to generalize a deep learning model to new data lacking appropriate MR inputs? An Exploration using Ultra-low-dose Amyloid PET/MRI. Montreal: Annual Meeting ISMRM; 2019.
Fischl B, Sereno MI, Dale AM. Cortical surface-based analysis. II: Inflation, flattening, and a surface-based coordinate system. Neuroimage. 1999;9:195–207. https://doi.org/10.1006/nimg.1998.0396.
Dale AM, Fischl B, Sereno MI. Cortical surface-based analysis. I Segmentation and surface reconstruction. Neuroimage. 1999;9:179–94. https://doi.org/10.1006/nimg.1998.0395.
Wang Z, Bovik AC, Sheikh HR, Simoncelli EP. Image quality assessment: from error visibility to structural similarity. IEEE Trans Image Process. 2004;13:600–12.
Desikan RS, Segonne F, Fischl B, Quinn BT, Dickerson BC, Blacker D, et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage. 2006;31:968–80. https://doi.org/10.1016/j.neuroimage.2006.01.021.
DeLong ER, DeLong DM, Clarke-Pearson DL. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics. 1988;44:837–45.
Cohen J. Statistical power analysis for the behavioral sciences. 2nd ed. Hillsdale: L. Erlbaum Associates; 1988.
Ye JC, Han Y, Cha E. Deep convolutional framelets: a general deep learning framework for inverse problems. SIAM J Imaging Sci. 2018;11:991–1048. https://doi.org/10.1137/17M1141771.
French RM. Catastrophic forgetting in connectionist networks. Trends Cogn Sci. 1999;3:128–35.
Kirkpatrick J, Pascanu R, Rabinowitz N, Veness J, Desjardins G, Rusu AA, et al. Overcoming catastrophic forgetting in neural networks. Proc Natl Acad Sci U S A. 2017;114:3521–6. https://doi.org/10.1073/pnas.1611835114.
Acknowledgments
This project was made possible by the NIH grants P41-EB015891 and P50-AG047366 (Stanford Alzheimer’s Disease Research Center), GE Healthcare, the Michael J. Fox Foundation for Parkinson’s Disease Research, the Foundation of the ASNR, and Life Molecular Imaging. The authors would also like to thank Tie Liang, EdD, for the statistical analysis.
Availability of data and material
Data was collected at the authors’ institutions and is available when requested for review.
Funding
This project was made possible by the NIH grants P41-EB015891 and P50-AG047366 (Stanford Alzheimer’s Disease Research Center), GE Healthcare, the Michael J. Fox Foundation for Parkinson’s Disease Research, the Foundation of the ASNR, and Life Molecular Imaging.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Outside submitted work: GZ-Subtle Medical Inc., co-founder and equity relationship. No other potential conflicts of interest relevant to this article exist.
Ethics approval
All procedures involving human participants were in accordance with the ethical standards of the Stanford University Institutional Review Board and the Leipzig University Ethics Committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Consent to participate
Written informed consent for imaging was obtained from all participants or an authorized surrogate decision-maker.
Code availability
Custom code was used for this project.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Advanced Image Analyses (Radiomics and Artificial Intelligence)
Electronic supplementary material
ESM 1
(DOCX 1.50 mb)
Rights and permissions
About this article
Cite this article
Chen, K.T., Schürer, M., Ouyang, J. et al. Generalization of deep learning models for ultra-low-count amyloid PET/MRI using transfer learning. Eur J Nucl Med Mol Imaging 47, 2998–3007 (2020). https://doi.org/10.1007/s00259-020-04897-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-020-04897-6