Abstract
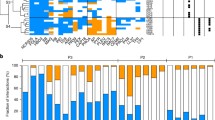
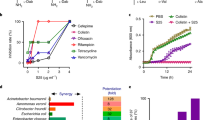
The rising prevalence of antibiotic resistance poses the greatest health threats. Antimicrobial peptides (AMPs) are regarded as the potentially effective therapy. To avoid current crisis of antibiotic resistance, a comprehensive understanding of AMP resistance is necessary before clinical application. In this study, the development of resistance to the anti-Gram-negative bacteria peptide N6NH2 (21 residues, β-sheet) was characterized in E. coli ATCC25922. Three N6NH2-resistant E. coli mutants with 32-fold increase in MIC were isolated by serially passaging bacterial lineages in progressively increasing concentrations of N6NH2 and we mainly focus on the phenotype of N6NH2-resistant bacteria different from sensitive bacteria. The results showed that the resistance mechanism was attributed to synergy effect of multiple mechanisms: (i) increase biofilm formation capacity (3 ~ 4-fold); (ii) weaken the affinity of lipopolysaccharide (LPS) with N6NH2 (3 ~ 8-fold); and (iii) change the cell membrane permeability and potential. Interestingly, a chimeric peptide-G6, also a N6NH2 analog, which keep the same antibacterial activity to both wild-type and resistant clones (MIC value: 16 μg/mL), could curb N6NH2-resistant mutants by stronger inhibition of biofilm formation, stronger affinity with LPS, and stronger membrane permeability and depolarization than that of N6NH2.
Graphical abstract








Similar content being viewed by others
Data availability
All datasets generated for this study are included in the article/Supplementary material.
References
Callaway E (2020) ‘It will change everything’: DeepMind’s AI makes gigantic leap in solving protein structures. Nature 588(7837):203–204
Camilo B, Vincent T, Christian K, Philip R, Robert B, Hinrich S, Gunther J (2017) Alternative evolutionary paths to bacterial antibiotic resistance cause distinct collateral effects. Mol Biol Evol 34(9):2229–2244
Cwiek K, Korzekwa K, Tabis A, Bania J, Bugla-Ploskonska G, Wieliczko A (2020) Antimicrobial resistance and biofilm formation capacity of Salmonella enterica serovar enteritidis strains isolated from poultry and humans in Poland. Pathogens 9(8):643
De Breij A, Riool M, Kwakman PHS, De Boer L, Cordfunke RA, Drijfhout JW, Cohen O, Emanuel N, Zaat SAJ, Nibbering PH (2016) Prevention of Staphylococcus aureus biomaterial-associated infections using a polymer-lipid coating containing the antimicrobial peptide OP-145. J Control Release 222:1–8
Elliott AG, Huang JX, Neve S, Zuegg J, Cooper MA (2020) An amphipathic peptide with antibiotic activity against multidrug-resistant Gram-negative bacteria. Nat Commun 11(1):3184
Fantner GE, Barbero RJ, Gray DS, Belcher AM (2010) Kinetics of antimicrobial peptide activity measured on individual bacterial cells using high-speed atomic force microscopy. Nat Nanotechnol 5(4):280–285
Guozhi B, Desiree Y, Regoes RR, Rolff J (2018) Predicting drug resistance evolution: insights from antimicrobial peptides and antibiotics. Proc R Soc Biol Sci U S A 285(1874):20172687
Han H, Li T, Wang Z, Teng D, Mao R, Hao Y, Yang N, Wang X, Wang J (2020) Improved stability and activity of a marine peptide-N6NH2 against Edwardsiella tarda and its preliminary application in fish. Mar Drugs 18(12):650
Hart EM, Mitchell AM, Konovalova A, Grabowicz M, Silhavy TJ (2019) A small-molecule inhibitor of BamA impervious to efflux and the outer membrane permeability barrier. Proc Natl Acad Sci U S A 116(43):21748–21757
Jochumsen N, Marvig RL, Damkiær SR, Jensen RL, Paulander W, Molin SR, Jelsbak L, Folkesson A (2016) The evolution of antimicrobial peptide resistance in Pseudomonas aeruginosa is shaped by strong epistatic interactions. Nat Commun 7:13002
Kubicek-Sutherland JZ, Hava L, Martin V, Karin H, Hanne I, Andersson DI (2017) Antimicrobial peptide exposure selects for Staphylococcus aureus resistance to human defence peptides. J Antimicrob Chemother 72:115–127
Larock CN, Nizet V (2015) Cationic antimicrobial peptide resistance mechanisms of streptococcal pathogens. Biochim Biophys Acta Biomembr 11:3047–3054
Lazzaro BP, Zasloff M, Rolff J (2020) Antimicrobial peptides: application informed by evolution. Science 368(6490):eaau5480
Li Z, Teng D, Mao R, Wang X, Hao Y, Wang X, Wang J (2018) Improved antibacterial activity of the marine peptide N6 against intracellular Salmonella Typhimurium by conjugating with the cell-penetrating peptide Tat11 via a cleavable linker. J Med Chem 61(17):7991–8000
Li T, Wang Z, Han H, Teng D, Mao R, Hao Y, Yang N, Wang X, Wang J (2020) Dual antibacterial activities and biofilm eradication of a marine peptide-N6NH2 and its analogs against multidrug-resistant Aeromonas veronii. Int J Mol Sci 21(24):9637
Lofton H, Pranting M, Thulin E, Andersson DI (2013) Mechanisms and fitness costs of resistance to antimicrobial peptides LL-37, CNY100HL and wheat germ histones. PLos One 8(7):e68875
Lofton H, Anwar N, Rhen M, Andersson DI (2015) Fitness of Salmonella mutants resistant to antimicrobial peptides. J Antimicrob Chemother 70(2):432–440
Loh B, Grant C, Hancock RE (1984) Use of the fluorescent probe 1-N-phenylnaphthylamine to study the interactions of aminoglycoside antibiotics with the outer membrane of Pseudomonas aeruginosa. Antimicrob Agents Chemother 26(4):546–551
Luther A, Urfer M, Zahn M, Müller M, Obrecht D (2019) Chimeric peptidomimetic antibiotics against Gram-negative bacteria. Nature 576(7787):1–6
Malik SZ, Linkevicius M, Göransson U, Andersson DI (2017) Resistance to the cyclotide cycloviolacin O2 in Salmonella enterica caused by different mutations that often confer cross-resistance or collateral sensitivity to other antimicrobial peptides. Antimicrob Agents Chemother 61(8):e00684-e717
Martin JK, Sheehan JP, Bratton BP, Moore GM, Gitai Z (2020) A dual-mechanism antibiotic kills Gram-negative bacteria and avoids drug resistance. Cell 181(7):1518–1532
Oz T, Guvenek A, Yildiz S, Karaboga E, Tamer YT, Mumcuyan N, Ozan VB, Senturk GH, Cokol M, Yeh PJ (2014) Strength of selection pressure is an important parameter contributing to the complexity of antibiotic resistance evolution. Mol Biol Evol 31(9):2387–2401
Pal C, Papp B, Lazar V (2015) Collateral sensitivity of antibiotic-resistant microbes. Trends Microbiol 23(7):401–407
Pena-Miller R, Laehnemann D, Jansen G, Fuentes-Hernandez A, Rosenstiel P, Schulenburg H, Beardmore R, Read AF (2013) When the most potent combination of antibiotics selects for the greatest bacterial load: the smile-frown transition. PLoS Biol 11(4):e1001540
Puertolasbalint F, Warsi OM, Linkevicius M, Tang P, Andersson DI (2019) Mutations that increase expression of the EmrAB-TolC efflux pump confer increased resistance to nitroxoline in Escherichia coli. J Antimicrob Chemother 75(2):300–308
Rosenkilde C, Munck C, Porse A, Linkevicius M, Andersson DI, Sommer M (2019) Collateral sensitivity constrains resistance evolution of the CTX-M-15 β-lactamase. Nat Commun 10(1):618
Satta G, Witney AA, Begum N, Ortiz Canseco J, Boa AN, McHugh TD (2020) Role of whole-genome sequencing in characterizing the mechanism of action of para-aminosalicylic acid and its resistance. Antimicrob Agents Chemother 64(9):e00675-e720
Shang D, Zhang Q, Dong W, Liang H, Bi X (2016) The effects of LPS on the activity of Trp-containing antimicrobial peptides against Gram-negative bacteria and endotoxin neutralization. Acta Biomater 33:153–165
Sims PJ, Waggoner AS, Wang CH, Hoffman JF (1974) Studies on the mechanism by which cyanine dyes measure membrane potential in red blood cells and phosphatidylcholine vesicles. Biochemistry 13(16):3315–3330
Song M, Liu Y, Huang X, Ding S, Zhu K (2020) A broad-spectrum antibiotic adjuvant reverses multidrug-resistant Gram-negative pathogens. Nat Microbiol 5(8):1–11
Sousa MC (2019) New antibiotics target the outer membrane of bacteria. Nature 576(7787):389–390
Spodsberg N (2006) Polypeptides having antimicrobial activity and polynucleotides encoding same, Patent, 2006 NO. 20060706125
Spohn R, Daruka L, Lazar V, Martins A, Vidovics F, Grezal G, Mehi O, Kintses B, Szamel M, Jangir PK, Csorgo B, Gyorkei A, Bodi Z, Farago A, Bodai L, Foldesi I, Kata D, Maroti G, Pap B, Wirth R, Papp B, Pal C (2019) Integrated evolutionary analysis reveals antimicrobial peptides with limited resistance. Nat Commun 10(1):4538
Wang Z, Liu X, Teng D, Mao R, Wang J (2020) Development of chimeric peptides to facilitate the neutralisation of lipopolysaccharides during bactericidal targeting of multidrug-resistant Escherichia coli. Commun Biol 3(1):41
Wistrand-Yuen E, Knopp M, Hjort K, Koskiniemi S, Berg OG, Andersson DI (2018) Evolution of high-level resistance during low-level antibiotic exposure. Nat Commun 9(1):1599
Yang N, Liu X, Teng D, Li Z, Wang X, Mao R, Wang X, Hao Y, Wang J (2017) Antibacterial and detoxifying activity of NZ17074 analogues with multi-layers of selective antimicrobial actions against Escherichia coli and Salmonella enteritidis. Sci Rep 7(1):3392–3392
Yu I, Meyer KJ, Akira I, Quentin FG, Robert G, Sylvie M, Mariaelena C, Miho M, Samantha N, Meghan G (2020) A new antibiotic selectively kills Gram-negative pathogens. Nature 576(7787):459–464
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415(6870):389–395
Zhang Q, Yang N, Mao R, Hao Y, Ma X, Teng D, Fan H, Wang J (2021) A recombinant fungal defensin-like peptide-P2 combats Streptococcus dysgalactiae and biofilms. Appl Microbiol Biotechnol 105(4):1489–1504
Funding
This work was supported by the National Natural Science Foundation of China (grants No. 31772640, No. 31572444, and No. 31572445) and Direction of Antibiotic Peptides and Alternatives to Antibiotics, the Agricultural Science and Technology Innovation Program (CAAS-ASTIP-2013-FRI-02) and Key Project of Alternatives to Antibiotic for Feed Usages of ASTIP (CAAS-ZDXT2018008) in CAAS.
Author information
Authors and Affiliations
Contributions
Z.L.W., R.Y.M., and J.H.W. conceived and designed the experiments; Z.L.W. conducted the experiments; N.Y. and D.T. supervised the work; N.Y., D.T., Y.H., T.L., and H.H.H. provided some data analysis and revised the final version of the manuscript; Z.L.W., N.Y., R.Y.M., and J.H.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, Z., Yang, N., Teng, D. et al. Resistance response to Arenicin derivatives in Escherichia coli. Appl Microbiol Biotechnol 106, 211–226 (2022). https://doi.org/10.1007/s00253-021-11708-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11708-x