Abstract
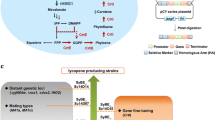
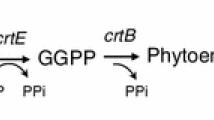
Lycopene is a red carotenoid pigment with strong antioxidant activity. Saccharomyces cerevisiae is considered a promising host to produce lycopene, but lycopene toxicity is one of the limiting factors for high-level production. In this study, we used heterologous lycopene biosynthesis genes crtE and crtI from Xanthophyllomyces dendrorhous and crtB from Pantoea agglomerans for lycopene production in S. cerevisiae. The crtE, crtB, and crtI genes were integrated into the genome of S. cerevisiae CEN.PK2-1C strain, while deleting DPP1 and LPP1 genes to inhibit a competing pathway producing farnesol. Lycopene production was further improved by inhibiting ergosterol production via downregulation of ERG9 expression and by deleting ROX1 or MOT3 genes encoding transcriptional repressors for mevalonate and sterol biosynthetic pathways. To further increase lycopene production, CrtE and CrtB mutants with improved activities were isolated by directed evolution, and subsequently, the mutated genes were randomly integrated into the engineered lycopene-producing strains via delta-integration. To relieve lycopene toxicity by increasing unsaturated fatty acid content in cell membranes, the OLE1 gene encoding stearoyl-CoA 9-desaturase was overexpressed. In combination with the overexpression of STB5 gene encoding a transcription factor involved in NADPH production, the final strain produced up to 41.8 mg/gDCW of lycopene, which is approximately 74.6-fold higher than that produced in the initial strain.






Similar content being viewed by others
References
Asadollahi MA, Maury J, Moller K, Nielsen KF, Schalk M, Clark A, Nielsen J (2008) Production of plant sesquiterpenes in Saccharomyces cerevisiae: effect of ERG9 repression on sesquiterpene biosynthesis. Biotechnol Bioeng 99(3):666–677. https://doi.org/10.1002/bit.21581
Asadollahi MA, Maury J, Schalk M, Clark A, Nielsen J (2010) Enhancement of farnesyl diphosphate pool as direct precursor of sesquiterpenes through metabolic engineering of the mevalonate pathway in Saccharomyces cerevisiae. Biotechnol Bioeng 106(1):86–96. https://doi.org/10.1002/bit.22668
Baek SH, Kwon EY, Bae SJ, Cho BR, Kim SY, Hahn JS (2017) Improvement of D-lactic acid production in Saccharomyces cerevisiae under acidic conditions by evolutionary and rational metabolic engineering. Biotechnol J 12:1700015. https://doi.org/10.1002/biot.201700015
Cadiere A, Galeote V, Dequin S (2010) The Saccharomyces cerevisiae zinc factor protein Stb5p is required as a basal regulator of the pentose phosphate pathway. FEMS Yeast Res 10(7):819–827. https://doi.org/10.1111/j.1567-1364.2010.00672.x
Chen Y, Xiao W, Wang Y, Liu H, Li X, Yuan Y (2016) Lycopene overproduction in Saccharomyces cerevisiae through combining pathway engineering with host engineering. Microb Cell Factories 15(1):113. https://doi.org/10.1186/s12934-016-0509-4
DiCarlo JE, Norville JE, Mali P, Rios X, Aach J, Church GM (2013) Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems. Nucleic Acids Res 41(7):4336–4343. https://doi.org/10.1093/nar/gkt135
Donald KA, Hampton RY, Fritz IB (1997) Effects of overproduction of the catalytic domain of 3-hydroxy-3-methylglutaryl coenzyme A reductase on squalene synthesis in Saccharomyces cerevisiae. Appl Environ Microbiol 63(9):3341–3344
Fang Z, Chen Z, Wang S, Shi P, Shen Y, Zhang Y, Xiao J, Huang Z (2017) Overexpression of OLE1 enhances cytoplasmic membrane stability and confers resistance to cadmium in Saccharomyces cerevisiae. Appl Environ Microbiol 83:e02319–e02316. https://doi.org/10.1128/aem.02319-16
Faulkner A, Chen X, Rush J, Horazdovsky B, Waechter CJ, Carman GM, Sternweis PC (1999) The LPP1 and DPP1 gene products account for most of the isoprenoid phosphate phosphatase activities in Saccharomyces cerevisiae. J Biol Chem 274(21):14831–14837
Gueldener U, Heinisch J, Koehler GJ, Voss D, Hegemann JH (2002) A second set of loxP marker cassettes for Cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res 30(6):e23
Hector RE, Bowman MJ, Skory CD, Cotta MA (2009) The Saccharomyces cerevisiae YMR315W gene encodes an NADP(H)-specific oxidoreductase regulated by the transcription factor Stb5p in response to NADPH limitation. New Biotechnol 26(3–4):171–180. https://doi.org/10.1016/j.nbt.2009.08.008
Kang MJ, Lee YM, Yoon SH, Kim JH, Ock SW, Jung KH, Shin YC, Keasling JD, Kim SW (2005) Identification of genes affecting lycopene accumulation in Escherichia coli using a shot-gun method. Biotechnol Bioeng 91(5):636–642. https://doi.org/10.1002/bit.20539
Kennedy MA, Bard M (2001) Positive and negative regulation of squalene synthase (ERG9), an ergosterol biosynthetic gene, in Saccharomyces cerevisiae. Biochim Biophys Acta 1517(2):177–189
Kim S, Hahn JS (2015) Efficient production of 2,3-butanediol in Saccharomyces cerevisiae by eliminating ethanol and glycerol production and redox rebalancing. Metab Eng 31:94–101. https://doi.org/10.1016/j.ymben.2015.07.006
Larochelle M, Drouin S, Robert F, Turcotte B (2006) Oxidative stress-activated zinc cluster protein Stb5 has dual activator/repressor functions required for pentose phosphate pathway regulation and NADPH production. Mol Cell Biol 26(17):6690–6701. https://doi.org/10.1128/mcb.02450-05
Liu P, Sun L, Sun Y, Shang F, Yan G (2016) Decreased fluidity of cell membranes causes a metal ion deficiency in recombinant Saccharomyces cerevisiae producing carotenoids. J Ind Microbiol Biotechnol 43(4):525–535. https://doi.org/10.1007/s10295-015-1728-0
Meadows AL, Hawkins KM, Tsegaye Y, Antipov E, Kim Y, Raetz L, Dahl RH, Tai A, Mahatdejkul-Meadows T, Xu L, Zhao L, Dasika MS, Murarka A, Lenihan J, Eng D, Leng JS, Liu CL, Wenger JW, Jiang H, Chao L, Westfall P, Lai J, Ganesan S, Jackson P, Mans R, Platt D, Reeves CD, Saija PR, Wichmann G, Holmes VF, Benjamin K, Hill PW, Gardner TS, Tsong AE (2016) Rewriting yeast central carbon metabolism for industrial isoprenoid production. Nature 537(7622):694–697. https://doi.org/10.1038/nature19769
Miyagi H, Kawai S, Murata K (2009) Two sources of mitochondrial NADPH in the yeast Saccharomyces cerevisiae. J Biol Chem 284(12):7553–7560. https://doi.org/10.1074/jbc.M804100200
Montanes FM, Pascual-Ahuir A, Proft M (2011) Repression of ergosterol biosynthesis is essential for stress resistance and is mediated by the Hog1 MAP kinase and the Mot3 and Rox1 transcription factors. Mol Microbiol 79(4):1008–1023. https://doi.org/10.1111/j.1365-2958.2010.07502.x
Mumberg D, Muller R, Funk M (1995) Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene 156(1):119–122
Nasution O, Lee YM, Kim E, Lee Y, Kim W, Choi W (2017) Overexpression of OLE1 enhances stress tolerance and constitutively activates the MAPK HOG pathway in Saccharomyces cerevisiae. Biotechnol Bioeng 114(3):620–631. https://doi.org/10.1002/bit.26093
Otero JM, Vongsangnak W, Asadollahi MA, Olivares-Hernandes R, Maury J, Farinelli L, Barlocher L, Osteras M, Schalk M, Clark A, Nielsen J (2010) Whole genome sequencing of Saccharomyces cerevisiae: from genotype to phenotype for improved metabolic engineering applications. BMC Genomics 11:723. https://doi.org/10.1186/1471-2164-11-723
Ozaydin B, Burd H, Lee TS, Keasling JD (2013) Carotenoid-based phenotypic screen of the yeast deletion collection reveals new genes with roles in isoprenoid production. Metab Eng 15:174–183. https://doi.org/10.1016/j.ymben.2012.07.010
Paradise EM, Kirby J, Chan R, Keasling JD (2008) Redirection of flux through the FPP branch-point in Saccharomyces cerevisiae by down-regulating squalene synthase. Biotechnol Bioeng 100(2):371–378. https://doi.org/10.1002/bit.21766
Paramasivan K, Mutturi S (2017) Regeneration of NADPH coupled with HMG-CoA reductase activity increases squalene synthesis in Saccharomyces cerevisiae. J Agric Food Chem 65(37):8162–8170. https://doi.org/10.1021/acs.jafc.7b02945
Pegklidou K, Mantzouridou F, Tsimidou MZ (2008) Lycopene production using Blakeslea trispora in the presence of 2-methyl imidazole: yield, selectivity, and safety aspects. J Agric Food Chem 56(4482):4490
Pereira FB, Guimaraes PM, Teixeira JA, Domingues L (2010) Selection of Saccharomyces cerevisiae strains for efficient very high gravity bio-ethanol fermentation processes. Biotechnol Lett 32(11):1655–1661. https://doi.org/10.1007/s10529-010-0330-9
Ro D-K, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM, Ho KA, Eachus RA, Ham TS, Kirby J, Chang MCY, Withers ST, Shiba Y, Sarpong R, Keasling JD (2006) Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440:940. https://doi.org/10.1038/nature04640
Scalcinati G, Partow S, Siewers V, Schalk M, Daviet L, Nielsen J (2012) Combined metabolic engineering of precursor and co-factor supply to increase alpha-santalene production by Saccharomyces cerevisiae. Microb Cell Factories 11:117. https://doi.org/10.1186/1475-2859-11-117
Schmidt-Dannert C (2000) Engineering novel carotenoids in microorganisms. Curr Opin Biotechnol 11(3):255–261
Shi YQ, Xin XL, Yuan QP (2012) Improved lycopene production by Blakeslea trispora with isopentenyl compounds and metabolic precursors. Biotechnol Lett 34(5):849–852. https://doi.org/10.1007/s10529-011-0839-6
Sung WS, Lee IS, Lee DG (2007) Damage to the cytoplasmic membrane and cell death caused by lycopene in Candida albicans. J Microbiol Biotechnol 17(11):1797–1804
Takahashi S, Yeo Y, Greenhagen BT, McMullin T, Song L, Maurina-Brunker J, Rosson R, Noel JP, Chappell J (2007) Metabolic engineering of sesquiterpene metabolism in yeast. Biotechnol Bioeng 97(1):170–181. https://doi.org/10.1002/bit.21216
Tippmann S, Scalcinati G, Siewers V, Nielsen J (2016) Production of farnesene and santalene by Saccharomyces cerevisiae using fed-batch cultivations with RQ-controlled feed. Biotechnol Bioeng 113(1):72–81. https://doi.org/10.1002/bit.25683
van Dijken JP, Bauer J, Brambilla L, Duboc P, Francois JM, Gancedo C, Giuseppin ML, Heijnen JJ, Hoare M, Lange HC, Madden EA, Niederberger P, Nielsen J, Parrou JL, Petit T, Porro D, Reuss M, van Riel N, Rizzi M, Steensma HY, Verrips CT, Vindelov J, Pronk JT (2000) An interlaboratory comparison of physiological and genetic properties of four Saccharomyces cerevisiae strains. Enzym Microb Technol 26(9–10):706–714
Verwaal R, Jiang Y, Wang J, Daran JM, Sandmann G, van den Berg JA, van Ooyen AJ (2010) Heterologous carotenoid production in Saccharomyces cerevisiae induces the pleiotropic drug resistance stress response. Yeast 27(12):983–998. https://doi.org/10.1002/yea.1807
Xie W, Lv X, Ye L, Zhou P, Yu H (2015) Construction of lycopene-overproducing Saccharomyces cerevisiae by combining directed evolution and metabolic engineering. Metab Eng 30:69–78. https://doi.org/10.1016/j.ymben.2015.04.009
Yamada R, Taniguchi N, Tanaka T, Ogino C, Fukuda H, Kondo A (2010) Cocktail delta-integration: a novel method to construct cellulolytic enzyme expression ratio-optimized yeast strains. Microb Cell Factories 9:32. https://doi.org/10.1186/1475-2859-9-32
Yoon SH, Lee YM, Kim JE, Lee SH, Lee JH, Kim JY, Jung KH, Shin YC, Keasling JD, Kim SW (2006) Enhanced lycopene production in Escherichia coli engineered to synthesize isopentenyl diphosphate and dimethylallyl diphosphate from mevalonate. Biotechnol Bioeng 94(6):1025–1032. https://doi.org/10.1002/bit.20912
Zhang Y, Nielsen J, Liu Z (2017) Engineering yeast metabolism for production of terpenoids for use as perfume ingredients, pharmaceuticals and biofuels. FEMS Yeast Res 17(8). https://doi.org/10.1093/femsyr/fox080
Zhao X, Shi F, Zhan W (2015) Overexpression of ZWF1 and POS5 improves carotenoid biosynthesis in recombinant Saccharomyces cerevisiae. Lett Appl Microbiol 61(4):354–360. https://doi.org/10.1111/lam.12463
Funding
This research was supported by the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (2017R1E1A1A01073523 and 2016M3D3A01913245).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 426 kb)
Rights and permissions
About this article
Cite this article
Hong, J., Park, SH., Kim, S. et al. Efficient production of lycopene in Saccharomyces cerevisiae by enzyme engineering and increasing membrane flexibility and NAPDH production. Appl Microbiol Biotechnol 103, 211–223 (2019). https://doi.org/10.1007/s00253-018-9449-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9449-8