Abstract
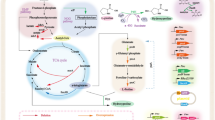
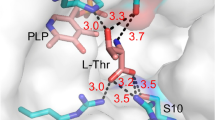
trans-Proline 4-hydroxylases (trans-P4Hs) hydroxylate free L-proline to trans-4-hydroxy-L-proline (trans-4-Hyp) is a valuable chiral synthon for important pharmaceuticals such as carbapenem antibiotics. However, merely few microbial trans-P4Hs have been identified, and trans-4-Hyp fermentations using engineered Escherichia coli strains expressing trans-P4Hs are usually performed at temperatures below 37 °C, which is likely due to poor stability and low activities. In the present study, a new trans-P4H from uncultured bacterium esnapd13 (UbP4H) with potential in the fermentative production of trans-4-Hyp at 37 °C was reported. In order to enhance the activity and thermostability of UbP4H, the replacement of its putative “lid” loop in combination with site-directed mutagenesis was performed. Consequently, four loop hybrids were designed by substituting a loop of UbP4H (A162-K178) with the corresponding sequences of four other known trans-P4Hs, respectively. Among them, UbP4H-Da exhibited a doubled activity when compared to the wild type (81.6 ± 1.9 vs. 40.4 ± 4.6 U/mg) but with reduced thermostability (t1/2, 11 vs. 47 min). Meanwhile, 10 single variants were designed through sequence alignments and folding free energy calculations. Three best point substitutions were respectively combined with UbP4H-Da, resulting in UbP4H-Da-R90G, UbP4H-Da-E112P, and UbP4H-Da-A260P. UbP4H-Da-E112P exhibited a 1.8-fold higher activity (85.2 ± 0.6 vs. 46.6 ± 4.0 U/mg), a 7.6-fold increase in t1/2 (359 vs. 47 min), and a 3 °C rise in Tm (46 vs. 43 °C) when compared to UbP4H. The fed-batch fermentations of trans-4-Hyp at 37 °C using trans-4-Hyp producing chassis cells expressing UbP4H or its variants were evaluated, and a 3.3-fold increase in trans-4-Hyp titer was obtained for UbP4H-Da-E112P (12.9 ± 0.1 vs. 3.9 ± 0.0 g/L for UbP4H). These results demonstrate the potential application of UbP4H-Da-E112P in the industrial production of trans-4-Hyp.








Similar content being viewed by others
References
Afriat-Jurnou L, Jackson CJ, Tawfik DS (2012) Reconstructing a missing link in the evolution of a recently diverged phosphotriesterase by active-site loop remodeling. Biochemistry 51(31):6047–6055. https://doi.org/10.1021/bi300694t
Anbar M, Gul O, Lamed R, Sezerman UO, Bayer EA (2012) Improved thermostability of Clostridium thermocellum endoglucanase Cel8A by using consensus-guided mutagenesis. Appl Environ Microbiol 78(9):3458–3464. https://doi.org/10.1128/AEM.07985-11
Blank LM, Ebert BE, Bühler B, Schmid A (2008) Metabolic capacity estimation of Escherichia coli as a platform for redox biocatalysis: constraint-based modeling and experimental verification. Biotechnol Bioeng 100(6):1050–1065. https://doi.org/10.1002/bit.21837
Boersma YL, Pijning T, Bosma MS, van der Sloot AM, Godinho LF, Dröge MJ, Winter RT, van Pouderoyen G, Dijkstra BW, Quax WJ (2008) Loop grafting of Bacillus subtilis lipase A: inversion of enantioselectivity. Chem Biol 15(8):782–789. https://doi.org/10.1016/j.chembiol.2008.06.009
Chen JJ, Gu DD, Li TY, Ju JS, Xue ZW, Li CH, Yan J, Zhang JX, Wang LA (2015) An efficient procedure for the production of trans-4-hydroxy-L-proline using recombinantly expressed proline hydroxylase. Sci Iran C 22(6):2350–2357
Chen Y, Luo Q, Zhou W, Xie Z, Cai YJ, Liao XR, Guan ZB (2016) Improving the catalytic efficiency of Bacillus pumilus CotA-laccase by site-directed mutagenesis. Appl Microbiol Biotechnol 101(5):1935–1944. https://doi.org/10.1007/s00253-016-7962-1
Dong YH, Li JF, Hu D, Yin X, Wang CJ, Tang SH, Wu MC (2016) Replacing a piece of loop-structure in the substrate-binding groove of Aspergillus usamii β-mannanase, AuMan5A, to improve its enzymatic properties by rational design. Appl Microbiol Biotechnol 100(9):3989–3998. https://doi.org/10.1007/s00253-015-7224-7
Falcioni F, Bühler B, Schmid A (2015) Efficient hydroxyproline production from glucose in minimal media by Corynebacterium glutamicum. Biotechnol Bioeng 112(2):322–330. https://doi.org/10.1002/bit.25442
Fisher CL, Pei GK (1997) Modification of a PCR-based site-directed mutagenesis method. BioTechniques 23(4):570–574
Fulton A, Frauenkron-Machedjou VJ, Skoczinski P, Wilhelm S, Zhu LL, Schwaneberg U, Jaeger KE (2015) Exploring the protein stability landscape: Bacillus subtilis lipase A as a model for detergent tolerance. Chembiochem 16(6):930–936. https://doi.org/10.1002/cbic.201402664
Hawkey PM, Livermore DM (2012) Carbapenem antibiotics for serious infections. BMJ 344:e3236. https://doi.org/10.1136/bmj.e3236
Höppner A, Widderich N, Lenders M, Bremer E, Smits SH (2014) Crystal structure of the ectoine hydroxylase, a snapshot of the active site. J Biol Chem 289(43):29570–29583. https://doi.org/10.1074/jbc.M114.576769
Houwaart S, Youssar L, Hüttel W (2015) Pneumocandin biosynthesis: involvement of a trans-selective proline hydroxylase. Chembiochem 15(16):2365–2369. https://doi.org/10.1002/cbic.201402175
Huang Y, Niu BF, Gao Y, Fu LM, Li WZ (2010) CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics 26(5):680–682. https://doi.org/10.1093/bioinformatics/btq003
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Klein C, Hüttel W (2011) A simple procedure for selective hydroxylation of L-proline and L-pipecolic acid with recombinantly expressed proline hydroxylases. Adv Synth Catal 353(8):1375–1383. https://doi.org/10.1002/adsc.201000863
Knauer SH, Hartl-Spiegelhauer O, Schwarzinger S, Hanzelmann P, Dobbek H (2012) The Fe(II)/α-ketoglutarate-dependent taurine dioxygenases from Pseudomonas putida and Escherichia coli are tetramers. FEBS J 279(5):816–831. https://doi.org/10.1111/j.1742-4658.2012.08473.x
Koketsu K, Shomura Y, Moriwaki K, Hayashi M, Mitsuhashi S, Hara R, Kino K, Higuchi Y (2015) Refined regio- and stereoselective hydroxylation of L-pipecolic acid bu protein engineering of L-proline cis-4-hydroxylase based on the X-ray crystal structure. ACS Synth Biol 4(4):383–392. https://doi.org/10.1021/sb500247a
Lawrence CC, Sobey WJ, Field RA, Baldwin JE, Schofield CJ (1996) Purification and intial characterization of proline 4-hydroxylase from Streptomyces griseoviridus P8648: a 2-oxoacid, ferrous-dependent dioxygenase involved in etamycin biosynthesis. Biochem J 313(1):185–191. https://doi.org/10.1042/bj3130185
Lukat P, Katsuyama Y, Wenzel S, Binz T, König C, Blankenfeldt W, Brönstrup M, Müller R (2017) Biosynthesis of methyl-proline containing griselimycins, natural products with anti-tuberculosis activity. Chem Sci 8(11):7521–7527. https://doi.org/10.1039/c7sc02622f
Markolovic S, Wilkins ES, Schofield JC (2015) Protein hydroxylation catalyzed by 2-oxoglutarate-dependent oxygenases. J Biol Chem 290(34):20712–20722. https://doi.org/10.1074/jbc.R115.662627
Martinez S, Hausinger PR (2015) Catalytic mechanisms of Fe(II)- and 2-oxoglutarate-dependent oxygenases. J Biol Chem 290(34):20702–20711. https://doi.org/10.1074/jbc.R115.648691
Mattay J, Houwaart S, Hüttel W (2018) Cryptic production of trans-3-hydroxyproline in echinocandin B biosynthesis. Appl Environ Microbiol 84(7):1–10. https://doi.org/10.1128/AEM.02370-17
Mihalik SJ, Raiville AM, Watkins PA (1995) Phytanic acid α-oxidation in rat liver peroxisomes production of α-hydroxyphytanoyl-CoA and formate is enhanced by dioxygenase cofactors. Eur J Biochem 232(2):545–551. https://doi.org/10.1111/j.1432-1033.1995.545zz.x
Onishi M, Okumura Y, Okamoto R, Ishikura T (1984) Proline hydroxylation by cell free extract of a Streptomycete. Biochem and Biophys Res Commun 120(1):45–51. https://doi.org/10.1016/0006-291X(84)91411-6
Prajapati RS, Das M, Sreeramulu S, Sirajuddin M, Srinivasan S, Krishnamurthy V, Ranjani R, Ramakrishnan C, Varadarajan R (2007) Thermodynamic effects of proline introduction on protein stability. Proteins 66(2):480–491. https://doi.org/10.1002/prot.21215
Reetz MT, Carballeira JD, Vogel A (2006) Iterative saturation mutagenesis on the basis of B factors as a strategy for increasing protein thermostability. Angew Chem 118(46):7909–7915. https://doi.org/10.1002/ange.200602795
Remuzon P (1996) trans-4-Hydroxy-L-proline, a useful and versatile chiral starting block. Tetrahedron 52(44):13803–13835. https://doi.org/10.1016/0040-4020(96)00822-8
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:W1):320–W1):324. https://doi.org/10.1093/nar/gku316
Shibasaki T, Mori H, Chiba. S, Ozaki A (1999) Microbial proline 4-hydroxylase screening and gene cloning. Appl Environ Microbiol 65(9):4028–4031
Shibasaki T, Hashimoto S, Mori H, Ozaki A (2000a) Construction of a novel hydroxyproline-producing recombinant Escherichia coli by introducing a proline 4-hydroxylase gene. J Biosci Bioeng 90(5):522–525. https://doi.org/10.1016/S1389-1723(01)80033-5
Shibasaki T, Mori H, Ozaki A (2000b) Enzymatic production of trans-4-hydroxy-L-proline by regio- and stereospecific hydroxylation of L-proline. Biosci Biotechnol Biochem 64(4):746–750. https://doi.org/10.1271/bbb.64.746
Theodosiou E, Breisch M, Julsing MK, Falcioni R, Bühler B, Schmid A (2017) An artificial TCA cycle selects for efficient α-ketoglutarate dependent hydroxylase catalysis in engineered Escherichia coli. Biotechnol Bioeng 114(7):1511–1520. https://doi.org/10.1002/bit.26281
Vázquez-Figueroa E, Chaparro-Riggers J, Bommarius AS (2007) Development of a thermostable glucose dehydrogenase by a structure-guided consensus concept. Chembiochem 8:2295–2301. https://doi.org/10.1002/cbic.200700500
Wang JX, Zhang ZY, Liu HD, Sun FF, Yue C, Hu JG, Wang CD (2016) Construction and optimization of trans-4-hydrpxy-L-proline production recombinant E. coli strain taking the glycerol as carbon source. J Chem Technol Biotechnol 91(9):2389–2398. https://doi.org/10.1002/jctb.5024
Wang XC, Liu J, Zhao J, Ni XM, Zheng P, Guo X, Sun CM, Sun JB, Ma YH (2018) Efficient production of trans-4-hydroxy-L-proline from glucose using a new trans-proline 4-hydroxylase in Escherichia coli. J biosci Bioengine doi. https://doi.org/10.1016/j.jbiosc.2018.04.012
Whalen KL, Chang KM, Spies MA (2011) Hybrid steered molecular dynamics-docking: an efficient solution to the problem of ranking inhibitor affinities against a flexible drug target. Mol Inf 30(5):459–471. https://doi.org/10.1002/minf.201100014
Xie Y, An J, Yang GY, Wu G, Zhang Y, Cui L, Feng Y (2014) Enhanced enzyme kinetic stability by increasing rigidity within the active site. J Biol Chem 289(11):7994–8006. https://doi.org/10.1074/jbc.M113.536045
Yi Y, Sheng H, Li Z, Ye Q (2014) Biosynthesis of trans-4-hydroxyproline by recombinant strains of Corynebacterium glutamicum and Escherichia coli. BMC Biotechnol 14:44. https://doi.org/10.1186/1472-6750-14-44
Yu H, Yan Y, Zhang C, Dalby PA (2017) Two strategies to engineer flexible loops for improved enzyme thermostability. Sci Rep 7:41212. https://doi.org/10.1038/srep41212
Acknowledgements
We thank Xingchu Wang (Tianjin Institute of Industrial Biotechnology) for helpful discussions. We thank Dr. Timothy C. Cairns and Taiwo Dele-Osibanjo for critical reading and editing of the manuscript.
Funding
We are grateful for the financial support from Tianjin Natural Science Foundation (18JCQNJC10300), National Natural Science Foundation of China (No. 21606251), the Key Research Program of the Chinese Academy of Sciences (No. KFZD-SW-212), Youth Innovation Promotion Association of CAS (2015137), and Science and Technology Project of Tianjin (Nos. 15PTCYSY00020 and 14ZCZDSY00058).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors. All authors confirm that ethical principles have been followed in the research as well as in manuscript preparation, and approved this submission.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 1014 kb)
Rights and permissions
About this article
Cite this article
Liu, C., Zhao, J., Liu, J. et al. Simultaneously improving the activity and thermostability of a new proline 4-hydroxylase by loop grafting and site-directed mutagenesis. Appl Microbiol Biotechnol 103, 265–277 (2019). https://doi.org/10.1007/s00253-018-9410-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9410-x