Abstract
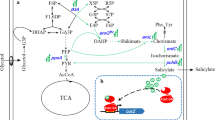
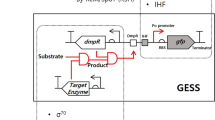
High-throughput screening is a popular tool for collating biological data which would otherwise require the use of excessive resources. In this study, an artificial genetic circuit sensor responding to dimethylallyl diphosphate (DMAPP) was constructed based on a modified L-arabinose operon for high-throughput screening and isoprene synthase (ispS) evolution in Escherichia coli (E. coli). As a first step, the DNA sequence of the L-arabinose ligand-binding domain (LBD) was replaced with an ispS gene to enable the AraC operon responding to DMAPP, which is the substrate of the IspS enzyme. Then, an enhanced GFP (eGFP) was also introduced as a reporter for pBAD promoter. The expression level of the reporter was monitored using either of the two tools: flow cytometer (FCM) and microplate reader. Sequentially, we observed that a high DMAPP concentration led to low eGFP fluorescence, and the overexpression of ispS gene, which consumes DMAPP, resulted in a high eGFP expression. These results demonstrated that the artificial genetic circuit sensor responded directly to the intracellular concentration of DMAPP, and the expression of IspS enzyme could be positively correlated to the expression level of eGFP. Finally, we identified two IspS mutants with different activities from an ispS gene library and further validated the screening method.





Similar content being viewed by others
References
Adam KP, Zapp J (1998) Biosynthesis of the isoprene units of chamomile sesquiterpenes. Phytochemistry 48(6):953–959. https://doi.org/10.1016/S0031-9422(97)00992-8
Berry A, Dodge TC, Pepsin M, Weyler W (2002) Application of metabolic engineering to improve both the production and use of biotech indigo. J Ind Microbiol Biotech 28(3):127–133. https://doi.org/10.1038/sj.jim.7000228
Bustos SA, Schleif RF (1993) Functional domains of the AraC protein. Proc Natl Acad Sci USA 90(12):5638–5642
Chou HH, Keasling JD (2013) Programming adaptive control to evolve increased metabolite production. Nat Commun 4:2595. https://doi.org/10.1038/ncomms3595
Dunn TM, Schleif RF (1984) An operator at −280 base pairs that is required for repression of araBAD operon promoter: addition of DNA helical turns between the operator and promoter cyclically hinders repression. Proc Natl Acad Sci USA 81(16):5017–5020. https://doi.org/10.1073/pnas.81.16.5017
Eustance RJ, Schleif RF (1997) The linker region of AraC protein. J Bacteriol 178(24):7025–7030
Eustance RJ, Bustos SA, Schleif RF (1994) Reaching out. Locating and lengthening the interdomain linker in AraC protein. J Mol Biol 242(4):330–338
Floras N, Xiao J, Berry A, Bolivar F, Valle F (1996) Pathway engineering for the production of aromatic compounds in Escherichia coli. Nat Biotechnol 14(5):620–623. https://doi.org/10.1038/nbt0596-620
Furubayashi M, Ikezumi M, Jun K, Iwasaki M, Fujii A, Li L, Saito K, Umeno D (2014) A high-throughput colorimetric screening assay for terpene synthase activity based on substrate consumption. PLoS One 9(3):11
Hemmerlin A, Harwood JL, Bach TJ (2012) A raison d’etre for two distinct pathways in the early steps of plant isoprenoid biosynthesis? Prog Lipid Res 51(2):95–148. https://doi.org/10.1016/j.plipres.2011.12.001
Hendrickson W, Schleif RF (1984) Regulation of the Escherichia coli l -arabinose operon studied by gel electrophoresis DNA binding assay. J Mol Biol 178(3):611–628. https://doi.org/10.1016/0022-2836(84)90241-9
Hendrickson W, Schleif R (1985) A dimer of AraC protein contacts three adjacent major groove regions of the araI DNA site. Proc Natl Acad Sci USA 82(10):3129–3133. https://doi.org/10.1073/pnas.82.10.3129
Hong SY, Zurbriggen AS, Melis A (2012) Isoprene hydrocarbons production upon heterologous transformation of Saccharomyces cerevisiae. J Appl Microbiol 113(1):52–65. https://doi.org/10.1111/j.1365-2672.2012.05319.x
Julsing MK, Rijpkema M, Woerdenbag HJ, Quax WJ, Kayser O (2007) Functional analysis of genes involved in the biosynthesis of isoprene in Bacillus subtilis. Appl Microbiol Biotech 75(6):1377
Koksal M, Zimmer I, Schnitzler JP, Christianson DW (2010) Structure of isoprene synthase illuminates the chemical mechanism of teragram atmospheric carbon emission. J Mol Biol 402(2):363–373. https://doi.org/10.1016/j.jmb.2010.07.009
Lauble H, Georgalis Y, Heinemann U (1989) Studies on the domain structure of the Salmonella typhimurium AraC protein. Eur J Biochemist 185(2):319–325. https://doi.org/10.1111/j.1432-1033.1989.tb15118.x
Lewis M, Chang G, Horton NC, Kercher MA, Pace HC, Schumacher MA, Brennan RG, Lu P (1996) Crystal structure of the lactose operon repressor and its complexes with DNA and inducer. Science 271(5253):1247–1254. https://doi.org/10.1126/science.271.5253.1247
Liu C-L, Fan L-H, Liu L, Tan T-W (2014) Combinational biosynthesis of isoprene by engineering the MEP pathway in Escherichia coli. Proc Biochemist 49(12):2078–2085
Liu C-L, Lv Q, Tan T-W (2015) Joint antisense RNA strategies for regulating isoprene production in Escherichia coli. RSC Adv 5(91):74892–74898. https://doi.org/10.1039/C5RA12161B
Lobell RB, Schleif RF (1990) DNA looping and unlooping by AraC protein. Science (N.Y.) 250(4980):528–532
Mahmoud SS, Croteau RB (2001) Metabolic engineering of essential oil yield and composition in mint by altering expression of deoxyxylulose phosphate reductoisomerase and menthofuran synthase. Proc Natl Aacd Sci USA 98(15):8915–8920. https://doi.org/10.1073/pnas.141237298
Martin K, Huo L, Schleif RF (1986) The DNA loop model for ara repression: AraC protein occupies the proposed loop sites in vivo and repression-negative mutations lie in these same sites. Proc Natl Aacd Sci USA 83(11):3654–3658. https://doi.org/10.1073/pnas.83.11.3654
Martin VJJ, Pitera DJ, Withers ST, Newman JD, Keasling JD (2003) Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat Biotechnol 21(7):796–802. https://doi.org/10.1038/nbt833
Ozaydin B, Burd H, Lee TS, Keasling JD (2013) Carotenoid-based phenotypic screen of the yeast deletion collection reveals new genes with roles in isoprenoid production. Metab Eng 15:174–183. https://doi.org/10.1016/j.ymben.2012.07.010
Sasaki K, Ohara K, Yazaki K (2005) Gene expression and characterization of isoprene synthase from Populus alba. Febs Lett 579(11):2514–2518. https://doi.org/10.1016/j.febslet.2005.03.066
Sharkey TD, Chen X, Yeh S (2001) Isoprene increases thermotolerance of fosmidomycin-fed leaves. Plant Physiol 125(4):2001
Sivy TL, Fall R, Rosenstiel TN (2011) Evidence of isoprenoid precursor toxicity in Bacillus subtilis. Biosci Biotechnol Biochem 75(12):2376–2383. https://doi.org/10.1271/bbb.110572
Takeno S, Bamba T, Nakazawa Y, Fukusaki E, Okazawa A, Kobayashi A (2010) High-throughput and highly sensitive analysis method for polyisoprene in plants by pyrolysis-gas chromatography/mass spectrometry. Biosci Biotech Biochemist 74(1):13–17
Wang F, Lv X, Xie W, Zhou P, Zhu Y, Zhen Y, Yang C, Yang X, Ye L, Yu H (2017) Combining Gal4p-mediated expression enhancement and directed evolution of isoprene synthase to improve isoprene production in Saccharomyces cerevisiae. Metab Eng 39:257–266. https://doi.org/10.1016/j.ymben.2016.12.011
Whited MG, Feher JF, Benko AD, Cervin AM, Chotani KG, McAuliffe CJ, LaDuca JR, Ben-Shoshan AE, Sanford JK (2010) Development of a gas-phase bioprocess for isoprene-monomer production using metabolic pathway engineering. Industrial Biotech 6(3):152–163
Yang J, Xian M, Su S, Zhao G, Nie Q, Jiang X, Zheng Y, Liu W (2012) Enhancing production of bio-isoprene using hybrid MVA pathway and isoprene synthase in E. coli. PLoS One 7(4):e33509. https://doi.org/10.1371/journal.pone.0033509
Yoon SH, Lee YM, Kim JE, Lee SH, Lee JH, Kim JY, Jung KH, Shin YC, Keasling JD, Kim SW (2006) Enhanced lycopene production in Escherichia coli engineered to synthesize isopentenyl diphosphate and dimethylallyl diphosphate from mevalonate. Biotechnol Bioeng 94(6):1025–1032. https://doi.org/10.1002/bit.20912
Acknowledgements
This work was supported by the State key laboratory of organic–inorganic composites, the National Basic Research Program of China (973 program) (2013CB733600), and the National Nature Science Foundation of China (21390202, 21476017). The authors also thank David Luo (Lawrence Berkeley National Lab, Joint Bio-Energy Institute) and Jan-Christer Janson (Uppsala University) for revising the language of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author(s) declare that they have no competing interests
Ethical statement
This article does not contain any studies with human participants performed by any of the authors.
Electronic supple material
ESM 1
(PDF 376 kb)
Rights and permissions
About this article
Cite this article
Liu, CL., Cai, JY., Bi, HR. et al. A novel DMAPP-responding genetic circuit sensor for high-throughput screening and evolving isoprene synthase. Appl Microbiol Biotechnol 102, 1381–1391 (2018). https://doi.org/10.1007/s00253-017-8676-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8676-8