Abstract
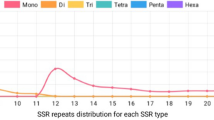
Agaricus subrufescens is one of the most important culinary-medicinal cultivable mushrooms with potentially high-added-value products and extended agronomical valorization. The development of A. subrufescens-related technologies is hampered by, among others, the lack of suitable molecular tools. Thus, this mushroom is considered as a genomic orphan species with a very limited number of available molecular markers or sequences. To fill this gap, this study reports the generation and analysis of the first set of expressed sequence tags (EST) for A. subrufescens. cDNA fragments obtained from young sporophores (SP) and vegetative mycelium in liquid culture (CL) were sequenced using 454 pyrosequencing technology. After assembly process, 4,989 and 5,125 sequences were obtained in SP and CL libraries, respectively. About 87 % of the EST had significant similarity with Agaricus bisporus-predicted proteins, and 79 % correspond to known proteins. Functional categorization according to Gene Ontology could be assigned to 49 % of the sequences. Some gene families potentially involved in bioactive compound biosynthesis could be identified. A total of 232 simple sequence repeats (SSRs) were identified, and a set of 40 EST-SSR polymorphic markers were successfully developed. This EST dataset provides a new resource for gene discovery and molecular marker development. It constitutes a solid basis for further genetic and genomic studies in A. subrufescens.




Similar content being viewed by others
References
Akiyama H, Endo M, Matsui T, Katsuda I, Emi N, Kawamoto Y, Koike T, Beppu H (2011) Agaritine from Agaricus blazei Murrill induces apoptosis in the leukemic cell line U937. Biochim Biophys Acta 1810(5):519–525
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST:a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Annadurai RS, Jayakumar V, Mugasimangalam RC, Katta MA, Anand S, Gopinathan S, Sarma SP, Fernandes SJ, Mullapudi N, Murugesan S, Rao SN (2012) Next generation sequencing and de novo transcriptome analysis of Costus pictus D. Don, a non-model plant with potent anti-diabetic properties. BMC Genomics 13:663
Baumgartner D, Hoesch L, Rast DM (1998) The biogenesis of β-N-(gamma-glutamyl)-4-hydroxymethylphenylhydrazine (agaritine) in Agaricus bisporus. Phytochemistry 49(2):465–474
Buée M, Reich M, Murat C, Morin E, Nilsson RH, Uroz S, Martin F (2009) 454 Pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity. New Phytol 184(2):449–456
Callac P, Theochari I, Kerrigan RW (2002) The germplasm of Agaricus bisporus: main results after ten years of collecting in France, in Greece and in North America. Acta Hortic 579:49–55
Chang ST, Wasser SP (2012) The role of culinary-medicinal mushrooms on human welfare with a pyramid model for human health. IntJ Med Mushrooms 14(2):95–134
Cho SM, Park JS, Kim KP, Cha DY, Kim HM, Yoo ID (1999) Chemical features and purification of immunostimulating polysaccharides from the fruiting bodies of Agaricus blazei. Korean J Mycol 27:170–174
Chum WW, Ng KT, Shih RS, Au CH, Kwan HS (2008) Gene expression studies of the dikaryotic mycelium and primordium of Lentinula edodes by serial analysis of gene expression. Mycol Res 112(Pt 8):950–964
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO:a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676
Cornman RS, Bennett AK, Murray KD, Evans JD, Elsik CG, Aronstein K (2012) Transcriptome analysis of the honey bee fungal pathogen, Ascosphaera apis:implications for host pathogenesis. BMC Genomics 13:285
Dashtban M, Schraft H, Qin W (2009) Fungal bioconversion of lignocellulosic residues; opportunities and perspectives. Int J Biol Sci 5(6):578–595
Diguistini S, Liao NY, Platt D, Robertson G, Seidel M, Chan SK, Docking TR, Birol I, Holt RA, Hirst M, Mardis E, Marra MA, Hamelin RC, Bohlmann J, Breuil C, Jones SJ (2009) De novo genome sequence assembly of a filamentous fungus using Sanger, 454 and Illumina sequence data. Genome Biol 10(9):R94
Duran C, Edwards D, Batley J (2009) Genetic maps and the use of synteny. Methods Mol Biol 513:41–55
Ekblom R, Galindo J (2011) Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity (Edinb) 107(1):1–15
Ellegren H (2004) Microsatellites:simple sequences with complex evolution. Nat Rev Genet 5(6):435–445
Ellis JR, Burke JM (2007) EST-SSRs as a resource for population genetic analyses. Heredity (Edinb) 99(2):125–132
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2(4):953–971
Endo M, Beppu H, Akiyama H, Wakamatsu K, Ito S, Kawamoto Y, Shimpo K, Sumiya T, Koike T, Matsui T (2010) Agaritine purified from Agaricus blazei Murrill exerts anti-tumor activity against leukemic cells. Biochim Biophys Acta 1800(7):669–673
Feau N, Bergeron MJ, Joly DL, Roussel F, Hamelin RC (2007) Detection and validation of EST-derived SNPs for poplar leaf rust Melampsora medusae f. sp.deltoidae. Mol Ecol Notes 7(6):1222–1228
Firenzuoli F, Gori L, Lombardo G (2008) The medicinal mushroom Agaricus blazei Murrill:review of literature and pharmaco-toxicological problems. Evid Based Complement Alternat Med 5(1):3–15
Forgetta V, Leveque G, Dias J, Grove D, Lyons RJ, Genik S, Wright C, Singh S, Peterson N, Zianni M, Kieleczawa J, Steen R, Perera A, Bintzler D, Adams S, Hintz W, Jacobi V, Bernier L, Levesque R, Dewar K (2013) Sequencing of the Dutch elm disease fungus genome using the Roche/454 GS-FLX Titanium System in a comparison of multiple genomics core facilities. J Biomol Tech 24(1):39–49
Foulongne-Oriol M, Spataro C, Moinard M, Cabannes D, Callac P, Savoie JM (2012) Development of polymorphic microsatellite markers issued from pyrosequencing technology for the medicinal mushroom Agaricus subrufescens. FEMS Microbiol Lett 334(2):119–126
Foulongne-Oriol M, Murat C, Castanera R, Ramirez L, Sonnenberg AS (2013) Genome-wide survey of repetitive DNA elements in the button mushroom Agaricus bisporus. Fungal Genet Biol 55(1):6–21
Fujimiya Y, Suzuki Y, Oshiman K, Kobori H, Moriguchi K, Nakashima H, Matumoto Y, Takahara S, Ebina T, Katakura R (1998) Selective tumoricidal effect of soluble proteoglucan extracted from the basidiomycete, Agaricus blazei Murill, mediated via natural killer cell activation and apoptosis. Cancer Immunol Immunother 46(3):147–159
Gonzaga MLC, Ricardo NMPS, Heatley F, Soares SD (2005) Isolation and characterization of polysaccharides from Agaricus blazei Murill. Carbohydr Polym 60(1):43–49
Hammond JBW, Nichols R (1976) Carbohydrate metabolism in Agaricus bisporus(Lange) Sing.; changes in soluble carbohydrates during growth of mycelium and sporophore. J Gen Microbiol 93:309–320
Hikichi M, Hiroe E, Okubo S (1999) Protein polysaccharide 0041. European Patent 0939082
Itoh H, Ito H, Hibasami H (2008) Blazein of a new steroid isolated from Agaricus blazei Murrill (himematsutake) induces cell death and morphological change indicative of apoptotic chromatin condensation in human lung cancer LU99 and stomach cancer KATO III cells. Oncol Rep 20(6):1359–1361
Joh JH, Lee SH, Lee JS, Kim KH, Jeong SJ, Youn WH, Kim NK, Son ES, Cho YS, Yoo YB, Lee CS, Kim BG (2007) Isolation of genes expressed during the developmental stages of the oyster mushroom, Pleurotus ostreatus, using expressed sequence tags. FEMS Microbiol Lett 276(1):19–25
Joh JH, Kim KY, Lim JH, Son ES, Park HR, Park YJ, Kong WS, Yoo YB, Lee CS (2009) Comparative analysis of expressed sequence tags from Flammulina velutipes at different developmental stages. J Microbiol Biotechnol 19(8):774–780
Kaur S, Cogan NO, Pembleton LW, Shinozuka M, Savin KW, Materne M, Forster JW (2011) Transcriptome sequencing of lentil based on second-generation technology permits large-scale unigene assembly and SSR marker discovery. BMC Genomics 12:265
Kawagishi H, Kanao T, Inagaki R, Mizuno T, Shimura K, Ito H, Hagiwara T, Nakamura T (1990) Formolysis of a potent antitumor (1→6)-β-d-glucan protein complex from Agaricus blazei fruiting bodies and antitumor-activity of the resulting products. Carbohydr Polym 12(4):393–403
Kent WJ (2002) BLAT–the BLAST-like alignment tool. Genome Res 12(4):656–664
Kimura Y, Kido T, Takaku T, Sumiyoshi M, Baba K (2004) Isolation of an anti-angiogenic substance from Agaricus blazei Murill:its antitumor and antimetastatic actions. Cancer Sci 95(9):758–764
Kirby J, Keasling JD (2009) Biosynthesis of plant isoprenoids:perspectives for microbial engineering. Annu Rev Plant Biol 60:335–355
Koo HJ, McDowell ET, Ma X, Greer KA, Kapteyn J, Xie Z, Descour A, Kim H, Yu Y, Kudrna D, Wing RA, Soderlund CA, Gang DR (2013) Ginger and turmeric expressed sequence tags identify signature genes for rhizome identity and development and the biosynthesis of curcuminoids, gingerols and terpenoids. BMC Plant Biol 13:27
Kubartova A, Ottosson E, Dahlberg A, Stenlid J (2012) Patterns of fungal communities among and within decaying logs, revealed by 454 sequencing. Mol Ecol 21(18):4514–4532
Largeteau M, Llarena-Hernandez RC, Regnault-Roger C, Savoie JM (2011) The medicinal Agaricus mushroom cultivated in Brazil:biology, cultivation and non-medicinal valorisation. Appl Microbiol Biotechnol 92(5):897–907
Lee SH, Kim BG, Kim KJ, Lee JS, Yun DW, Hahn JH, Kim GH, Lee KH, Suh DS, Kwon ST, Lee CS, Yoo YB (2002) Comparative analysis of sequences expressed during the liquid-cultured mycelia and fruit body stages of Pleurotus ostreatus. Fungal Genet Biol 35(2):115–134
Li W, Godzik A (2006) Cd-hit:a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22(13):1658–1659
Li YC, Korol AB, Fahima T, Nevo E (2004) Microsatellites within genes:structure, function, and evolution. Mol Biol Evol 21(6):991–1007
Liu K, Muse SV (2005) PowerMarker:an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129
Llarena-Hernandez RC, Largeteau ML, Farnet AM, Foulongne-Oriol M, Ferrer N, Regnault-Roger C, Savoie JM (2013) Potential of European wild strains of Agaricus subrufescens for productivity and quality on wheat straw based compost. World J Microbiol Biotechnol 29(7):1243–1253
Luo H, Sun C, Song J, Lan J, Li Y, Li X, Chen S (2010) Generation and analysis of expressed sequence tags from a cDNA library of the fruiting body of Ganoderma lucidum. Chin Med 5:9
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer ML, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Metzker ML (2010) Sequencing technologies - the next generation. Nat Rev Genet 11(1):31–46
Mizuno T, Hagiwara T, Nakamura T, Ito H, Shimura K, Sumiya T, Asakura A (1990) Antitumor activity and some properties of water-soluble polysaccharides from “Himematsutake”, the fruiting body of Agaricus blazei murrill. Agric Biol Chem 54:2897–2906
Morin E, Kohler A, Baker AR, Foulongne-Oriol M, Lombard V, Nagy LG, Ohm RA, Patyshakuliyeva A, Brun A, Aerts AL, Bailey AM, Billette C, Coutinho PM, Deakin G, Doddapaneni H, Floudas D, Grimwood J, Hildén K, Kües U, Labutti KM, Lapidus A, Lindquist EA, Lucas SM, Murat C, Riley RW, Salamov AA, Schmutz J, Subramanian V, Wösten HA, Xu J, Eastwood DC, Foster GD, Sonnenberg AS, Cullen D, de Vries RP, Lundell T, Hibbett DS, Henrissat B, Burton KS, Kerrigan RW, Challen MP, Grigoriev IV, Martin F (2012) Genome sequence of the button mushroom Agaricus bisporus reveals mechanisms governing adaptation to a humic-rich ecological niche. Proc Natl Acad Sci U S A 109(43):17501–17506
Nagaoka MH, Nagaoka H, Kondo K, Akiyama H, Maitani T (2006) Measurement of a genotoxic hydrazine, agaritine, and its derivatives by HPLC with fluorescence derivatization in the Agaricus mushroom and its products. Chem Pharm Bull 54(6):922–924
Ohm RA, de Jong JF, de Bekker C, Wösten HAB, Lugones LG (2011) Transcription factor genes of Schizophyllum commune involved in regulation of mushroom formation. Mol Microbiol 81(6):1433–1445
Opik M, Metsis M, Daniell TJ, Zobel M, Moora M (2009) Large-scale parallel 454 sequencing reveals host ecological group specificity of arbuscular mycorrhizal fungi in a boreonemoral forest. New Phytol 184(2):424–437
Ospina-Giraldo MD, Collopy PD, Romaine CP, Royse DJ (2000) Classification of sequences expressed during the primordial and basidiome stages of the cultivated mushroom Agaricus bisporus. Fungal Genet Biol 29(2):81–94
Parkinson J, Blaxter M (2009) Expressed sequence tags:an overview. Methods Mol Biol 533:1–12
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0:discriminating signal peptides from transmembrane regions. Nat Methods 8(10):785–786
Peter-Valence F, Llarena-Hernandez C, Largeteau M, Savoie JM, Ruaudel F, Ziarelli F, Ferre E, Farnet AM (2011) Chemical characterization of the biomass of an edible medicinal mushroom, Agaricus subrufescens, via solid-state 13C NMR. J Agric Food Chem 59(16):8939–8943
Pop M, Salzberg SL (2008) Bioinformatics challenges of new sequencing technology. Trends Genet 24(3):142–149
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan:proteindomains identifier. Nucleic Acids Res 33(Web Server issue):W116–W120
Ramirez L, Oguiza JA, Perez G, Lavin JL, Omarini A, Santoyo F, Alfaro M, Castanera R, Parenti A, Muguerza E, Pisabarro AG (2011) Genomics and transcriptomics characterization of genes expressed during postharvest at 4 °C by the edible basidiomycete Pleurotus ostreatus. Int Microbiol 14(2):111–120
Rice P, Longden I, Bleasby A (2000) EMBOSS:the European Molecular Biology Open Software Suite. Trends Genet 16(6):276–277
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Sato S, Feltus FA, Iyer P, Tien M (2009) The first genome-level transcriptome of the wood-degrading fungus Phanerochaete chrysosporium grown on red oak. Curr Genet 55(3):273–286
Savoie JM, Foulongne-Oriol M, Barroso G, Callac P (2013) Genetics and genomics of cultivated mushrooms, application to breeding of Agarics. In: Kempken F (ed) The Mycota, Agricultural Applications, vol 11. Springer, Berlin, pp 3–33
Schultz J, Milpetz F, Bork P, Ponting CP (1998) SMART, a simple modular architecture research tool:Identification of signaling domains. Proc Natl Acad Sci U S A 95:5857–5864
Sharma V, Sarkar IN (2013) Bioinformatics opportunities for identification and study of medicinal plants. Brief Bioinform 14(2):238–250
Soanes DM, Skinner W, Keon J, Hargreaves J, Talbot NJ (2002) Genomics of phytopathogenic fungi and the development of bioinformatic resources. Mol Plant Microbe Interact 15(5):421–427
Sonnhammer EL, von Heijne G, Krogh A (1998) A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6:175–182
Stein LD, Mungall C, Shu S, Caudy M, Mangone M, Day A, Nickerson E, Stajich JE, Harris TW, Lewis S (2002) The generic genome browser:a building block for a model organism system database. Genome Res 12(10):1599–1610
Stukenbrock EH (2013) Evolution, selection and isolation:a genomic view of speciation in fungal plant pathogens. New Phytol 199(4):895–907
Stussi H, Rast DM (1981) The biosynthesis and possible function of gamma-glutaminyl-4-hydroxybenzene in Agaricus bisporus. Phytochemistry 20(10):2347–2352
Takaku T, Kimura Y, Okuda H (2001) Isolation of an antitumor compound from Agaricus blazei Murill and its mechanism of action. J Nutr 131(5):1409–1413
Tedersoo L, Nilsson RH, Abarenkov K, Jairus T, Sadam A, Saar I, Bahram M, Bechem E, Chuyong G, Koljalg U (2010) 454 Pyrosequencing and Sanger sequencing of tropical mycorrhizal fungi provide similar results but reveal substantial methodological biases. New Phytol 188(1):291–301
Varshney RK, Mahendar T, Aggarwal RK, Börner A (2007) Genic molecular markers in plants:development and applications. In: Varshney RK, Tuberosa R (eds) Genomics-assisted crop improvment:genomics approaches and platforms, vol 1. Springer, Dordrecht, pp 13–29
Varshney RK, Nayak SN, May GD, Jackson SA (2009) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotechnol 27(9):522–530
Vera JC, Wheat CW, Fescemyer HW, Frilander MJ, Crawford DL, Hanski I, Marden JH (2008) Rapid transcriptome characterization for a nonmodel organism using 454 pyrosequencing. Mol Ecol 17(7):1636–1647
Wall PK, Leebens-Mack J, Chanderbali AS, Barakat A, Wolcott E, Liang H, Landherr L, Tomsho LP, Hu Y, Carlson JE, Ma H, Schuster SC, Soltis DE, Soltis PS, Altman N, dePamphilis CW (2009) Comparison of next generation sequencing technologies for transcriptome characterization. BMC Genomics 10:347
Ward JA, Ponnala L, Weber CA (2012) Strategies for transcriptome analysis in nonmodel plants. Am J Bot 99(2):267–276
Wawrzyn GT, Bloch SE, Schmidt-Dannert C (2012) Discovery and characterization of terpenoid biosynthetic pathways of fungi. Methods Enzymol 515:83–105
Wessling R, Schmidt SM, Micali CO, Knaust F, Reinhardt R, Neumann U, Loren V, van Themaat E, Panstruga R (2012) Transcriptome analysis of enriched Golovinomyces orontii haustoria by deep 454 pyrosequencing. Fungal Genet Biol 49(6):470–482
Wisitrassameewong K, Karunarathna SC, Thongklang N, Zhao R, Callac P, Moukha S, Férandon C, Chukeatirote E, Hyde K (2012) Agaricus subrufescens: a review. Saudi J Biol Sci 19(2):131–146
Xiang L, Li Y, Zhu Y, Luo H, Li C, Xu X, Sun C, Song J, Shi L, He L, Sun W, Chen S (2014) Transcriptome analysis of the Ophiocordyceps sinensis fruiting body reveals putative genes involved in fruiting body development and cordycepin biosynthesis. Genomics 103(1):154–159
Zeng S, Xiao G, Guo J, Fei Z, Xu Y, Roe BA, Wang Y (2010) Development of a EST dataset and characterization of EST-SSRs in a traditional Chinese medicinal plant, Epimedium sagittatum (Sieb. Et Zucc.) Maxim. BMC Genomics 11:94
Zhao RL, Karunarathna S, Raspé O, Parra LA, Guinberteau J, Moinard M, De Kesel A, Barroso G, Courtecuisse R, Hyde K, Guelly AK, Desjardin DE, Callac P (2011) Major clades in tropical Agaricus. Fungal Divers 51:279–296
Acknowledgments
This work was financially supported by a research project funded by a bilateral cooperation between Mexico (project 115790 CONACYT) and France (ANR-09-BLAN-0391-01). We acknowledge the URGI platform and particularly people involved in the GnpIS information system development and system administration.
Author information
Authors and Affiliations
Corresponding author
Additional information
Joelle Amselem and Jean-Michel Savoie were joint senior authors on this work.
Rights and permissions
About this article
Cite this article
Foulongne-Oriol, M., Lapalu, N., Férandon, C. et al. The first set of expressed sequence tags (EST) from the medicinal mushroom Agaricus subrufescens delivers resource for gene discovery and marker development. Appl Microbiol Biotechnol 98, 7879–7892 (2014). https://doi.org/10.1007/s00253-014-5844-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5844-y