Abstract
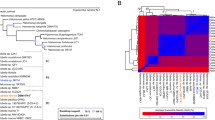
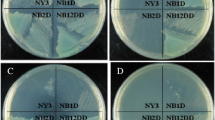
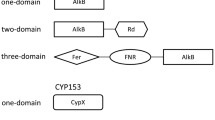
CYP153 and AlkB-like hydroxylases were recently discovered in Gram-positive alkane-degrading bacteria. However, it is unclear whether they cooperate with each other in alkane degradation as they do in Gram-negative bacteria. In this paper, we cloned the CYP153 gene from a representative Gram-positive alkane-degrading bacterium, Dietzia sp. DQ12-45-1b. The CYP153 gene transcription in Dietzia sp. DQ12-45-1b and heterologous expression in alkB gene knockout mutant strain Pseudomonas fluorescens KOB2∆1 both confirmed the functions of CYP153 on C6-C10 n-alkanes degradation, but not on longer chain-length n-alkanes. In addition, substrate-binding analysis of the purified CYP153 protein revealed different substrate affinities to C6-C16 n-alkanes, confirming n-alkanes binding to CYP153 protein. Along with AlkW1, an AlkB-like alkane hydroxylase in Dietzia sp. DQ12-45-1b, a teamwork pattern was found in n-alkane degradation, i.e. CYP153 was responsible for hydroxylating n-alkanes shorter than C10 while AlkW1 was responsible for those longer than C14. Further sequence analysis suggested that the high horizontal gene transfer (HGT) potential of CYP153 genes may be universal in Gram-positive alkane-degrading actinomycetes that contain both alkB and CYP153 genes.






Similar content being viewed by others
References
Alonso-Gutierrez J, Teramoto M, Yamazoe A, Harayama S, Figueras A, Novoa B (2011) Alkane-degrading properties of Dietzia sp. H0B, a key player in the Prestige oil spill biodegradation (NW Spain). J Appl Microbiol 111(4):800–810. doi:10.1111/j.1365-2672.2011.05104.x
Bihari Z, Szabo Z, Szvetnik A, Balazs M, Bartos P, Tolmacsov P, Zombori Z, Kiss I (2010) Characterization of a novel long-chain n-alkane-degrading strain, Dietzia sp. E1. Z Naturforsch C 65(11–12):693–700
Bihari Z, Szvetnik A, Szabó Z, Blastyák A, Zombori Z, Balázs M, Kiss I (2011) Functional analysis of long-chain n-alkane degradation by Dietzia spp. FEMS Microbiol Lett 316:100–107. doi:10.1111/j.1574-6968.2010.02198.x
Bodtker G, Hvidsten IV, Barth T, Torsvik T (2009) Hydrocarbon degradation by Dietzia sp. A14101 isolated from an oil reservoir model column. Antonie Van Leeuwenhoek 96(4):459–469. doi:10.1007/s10482-009-9359-y
Cai M, Chen WM, Nie Y, Chi CQ, Wang YN, Tang YQ, Li GY, Wu XL (2011) Complete genome sequence of Amycolicicoccus subflavus DQS3-9A1T, an actinomycete isolated from crude oil-polluted soil. J Bacteriol 193(17):4538–4539. doi:10.1128/jb.05388-11
Feng L, Wang W, Cheng J, Ren Y, Zhao G, Gao C, Tang Y, Liu X, Han W, Peng X, Liu R, Wang L (2007) Genome and proteome of long-chain alkane degrading Geobacillus thermodenitrificans NG80-2 isolated from a deep-subsurface oil reservoir. Proc Natl Acad Sci U S A 104(13):5602–5607. doi:10.1073/pnas.0609650104
Funhoff EG, Bauer U, Garcia-Rubio I, Witholt B, van Beilen JB (2006) CYP153A6, a soluble P450 oxygenase catalyzing terminal-alkane hydroxylation. J Bacteriol 188(14):5220–5227. doi:10.1128/jb.00286-06
Gallegos MT, Schleif R, Bairoch A, Hofmann K, Ramos JL (1997) Arac/XylS family of transcriptional regulators. Microbiol Mol Biol Rev 61(4):393–410
Ishikawa J, Yamashita A, Mikami Y, Hoshino Y, Kurita H, Hotta K, Shiba T, Hattori M (2004) The complete genomic sequence of Nocardia farcinica IFM 10152. Proc Natl Acad Sci U S A 101(41):14925–14930. doi:10.1073/pnas.0406410101
Kim J, Roh SW, Bae JW (2011) Draft genome sequence of Dietzia alimentaria 72T, belonging to the family Dietziaceae, isolated from a traditional Korean food. J Bacteriol 193(23):6791. doi:10.1128/jb.06229-11
Lageveen RG, Huisman GW, Preusting H, Ketelaar P, Eggink G, Witholt B (1988) Formation of Polyesters by Pseudomonas oleovorans: effect of substrates on formation and composition of poly-(R)-3-hydroxyalkanoates and poly-(R)-3-hydroxyalkenoates. Appl Environ Microbiol 54(12):2924–2932
Langille MGI, Brinkman FSL (2009) IslandViewer: an integrated interface for computational identification and visualization of genomic islands. Bioinformatics 25(5):664–665
Lazar I, Petrisor IG, Yen TF (2007) Microbial enhanced oil recovery (MEOR). Pet Sci Technol 25(11):1353–1366. doi:10.1080/10916460701287714
Liu C, Wang W, Wu Y, Zhou Z, Lai Q, Shao Z (2011) Multiple alkane hydroxylase systems in a marine alkane degrader, Alcanivorax dieselolei B-5. Environ Microbiol 13(5):1168–1178. doi:10.1111/j.1462-2920.2010.02416.x
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25(4):402–408
Lo Piccolo L, De Pasquale C, Fodale R, Puglia AM, Quatrini P (2011) Involvement of an alkane hydroxylase system of Gordonia sp. strain SoCg in degradation of solid n-alkanes. Appl Environ Microbiol 77(4):1204–1213. doi:10.1128/AEM.02180-10
Merkx M, Kopp DA, Sazinsky MH, Blazyk JL, Müller J, Lippard SJ (2001) Dioxygen activation and methane hydroxylation by soluble methane monooxygenase: a tale of two irons and three proteins. Angew Chem 40(15):2782–2807. doi:10.1002/1521-3773(20010803)40:15<2782::aid-anie2782>3.0.co;2-p
Miles J, Munro A, Rospendowski B, Smith W, McKnight J, Thomson A (1992) Domains of the catalytically self-sufficient cytochrome P-450 BM-3. Genetic construction, overexpression, purification and spectroscopic characterization. Biochem J 288(Pt 2):503
Nie Y, Liang J, Fang H, Tang YQ, Wu XL (2011) Two novel alkane hydroxylase-rubredoxin fusion genes isolated from a Dietzia bacterium and the functions of fused rubredoxin domains in long-chain n-alkane degradation. Appl Environ Microbiol 77(20):7279–7288. doi:10.1128/aem.00203-11
Procopio L, Alvarez VM, Jurelevicius DA, Hansen L, Sorensen SJ, Cardoso JS, Padula M, Leitao AC, Seldin L, van Elsas JD (2012) Insight from the draft genome of Dietzia cinnamea P4 reveals mechanisms of survival in complex tropical soil habitats and biotechnology potential. Antonie Van Leeuwenhoek 101(2):289–302. doi:10.1007/s10482-011-9633-7
Rainey FA, Klatte S, Kroppenstedt RM, Stackebrandt E (1995) Dietzia, a new genus including Dietzia maris comb. nov., formerly Rhodococcus maris. Int J Syst Bacteriol 45(1):32–36
Ramos JL, Martínez-Bueno M, Molina-Henares AJ, Terán W, Watanabe K, Zhang X, Gallegos MT, Brennan R, Tobes R (2005) The TetR family of transcriptional repressors. Microbiol Mol Biol Rev 69(2):326–356
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Schneiker S, dos Santos VAPM, Bartels D, Bekel T, Brecht M, Buhrmester J, Chernikova TN, Denaro R, Ferrer M, Gertler C, Goesmann A, Golyshina OV, Kaminski F, Khachane AN, Lang S, Linke B, McHardy AC, Meyer F, Nechitaylo T, Pühler A, Regenhardt D, Rupp O, Sabirova JS, Selbitschka W, Yakimov MM, Timmis KN, Vorhölter F-J, Weidner S, Kaiser O, Golyshin PN (2006) Genome sequence of the ubiquitous hydrocarbon-degrading marine bacterium Alcanivorax borkumensis. Nat Biotechnol 24(8):997–1004. doi:10.1038/nbt1232
Smits THM, Balada SB, Witholt B, van Beilen JB (2002) Functional analysis of alkane hydroxylases from gram-negative and gram-positive bacteria. J Bacteriol 184(6):1733–1742. doi:10.1128/jb.184.6.1733-1742.2002
Smits THM, Seeger M, Witholt B, van Beilen JB (2001) New alkane-responsive expression vectors for Escherichia coli and Pseudomonas. Plasmid 46(1):16–24. doi:10.1006/plas.2001.1522
Stinear TP, Seemann T, Harrison PF, Jenkin GA, Davies JK, Johnson PDR, Abdellah Z, Arrowsmith C, Chillingworth T, Churcher C, Clarke K, Cronin A, Davis P, Goodhead I, Holroyd N, Jagels K, Lord A, Moule S, Mungall K, Norbertczak H, Quail MA, Rabbinowitsch E, Walker D, White B, Whitehead S, Small PLC, Brosch R, Ramakrishnan L, Fischbach MA, Parkhill J, Cole ST (2008) Insights from the complete genome sequence of Mycobacterium marinum on the evolution of Mycobacterium tuberculosis. Genome Res 18(5):729–741. doi:10.1101/gr.075069.107
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599
Throne-Holst M, Wentzel A, Ellingsen TE, Kotlar HK, Zotchev SB (2007) Identification of novel genes involved in long-chain n-alkane degradation by Acinetobacter sp. Strain DSM 17874. Appl Environ Microbiol 73(10):3327–3332. doi:10.1128/aem.00064-07
van Beilen JB, Funhoff EG, van Loon A, Just A, Kaysser L, Bouza M, Holtackers R, Rothlisberger M, Li Z, Witholt B (2006) Cytochrome P450 alkane hydroxylases of the CYP153 family are common in alkane-degrading eubacteria lacking integral membrane alkane hydroxylases. Appl Environ Microbiol 72(1):59–65. doi:10.1128/aem.72.1.59-65.2006
van Beilen JB, Wubbolts MG, Witholt B (1994) Genetics of alkane oxidation by Pseudomonas oleovorans. Biodegradation 5(3–4):161–174
von der Weid I, Marques JM, Cunha CD, Lippi RK, Dos Santos SC, Rosado AS, Lins U, Seldin L (2007) Identification and biodegradation potential of a novel strain of Dietzia cinnamea isolated from a petroleum-contaminated tropical soil. Syst Appl Microbiol 30(4):331–339. doi:10.1016/j.syapm.2006.11.001
Wang W, Shao Z (2012) Genes involved in alkane degradation in the Alcanivorax hongdengensis strain A-11-3. Appl Microbiol Biotechnol 94:437–448. doi:10.1007/s00253-011-3818-x
Wang XB, Chi CQ, Nie Y, Tang YQ, Tan Y, Wu G, Wu XL (2011) Degradation of petroleum hydrocarbons (C6-C40) and crude oil by a novel Dietzia strain. Bioresour Technol 102(17):7755–7761. doi:10.1016/j.biortech.2011.06.009
Wang YN, Chi CQ, Cai M, Lou ZY, Tang YQ, Zhi XY, Li WJ, Wu XL, Du X (2010) Amycolicicoccus subflavus gen. nov., sp. nov., an actinomycete isolated from a saline soil contaminated by crude oil. Int J Syst Evol Microbiol 60(Pt 3):638–643. doi:10.1099/ijs.0.010546-0
Whyte LG, Smits THM, Labbe D, Witholt B, Greer CW, van Beilen JB (2002) Gene cloning and characterization of multiple alkane hydroxylase systems in Rhodococcus Strains Q15 and NRRL B-16531. Appl Environ Microbiol 68(12):5933–5942. doi:10.1128/aem.68.12.5933-5942.2002
Acknowledgments
This study was supported by the National Natural Sncience Foundation of China (31070107, 31225001). The authors are indebted to Dr. Theo Smits for his kind providing P. fluorescens KOB2∆1 and pCom8 plasmid.
Author information
Authors and Affiliations
Corresponding author
Additional information
Yong Nie, Jie-Liang Liang and Hui Fang equally contributed to the work.
Rights and permissions
About this article
Cite this article
Nie, Y., Liang, JL., Fang, H. et al. Characterization of a CYP153 alkane hydroxylase gene in a Gram-positive Dietzia sp. DQ12-45-1b and its “team role” with alkW1 in alkane degradation. Appl Microbiol Biotechnol 98, 163–173 (2014). https://doi.org/10.1007/s00253-013-4821-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-4821-1