Abstract
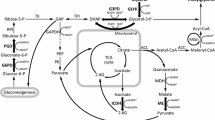
The oleaginous yeast Rhodotorula glutinis has been known to be a potential feedstock for lipid production. In the present study, we investigated the enhancement of expression of malic enzyme (ME; NADP+ dependent; EC 1.1.1.40) from Mucor circinelloides as a strategy to improve lipid content inside the yeast cells. The 26S rDNA and 5.8S rDNA gene fragments isolated from Rhodotorula glutinis were used for homologous integration of ME gene into R. glutinis chromosome under the control of the constitutively highly expressed gene phosphoglycerate kinase 1 to achieve stable expression. We demonstrated that by increasing the expression of the foreign ME gene in R. glutinis, we successfully improved the lipid content by more than twofold. At the end of lipid accumulation phrase (96 h) in the transformants, activity of ME was increased by twofold and lipid content of the yeast cells was increased from 18.74 % of the biomass to 39.35 %. Simultaneously, there were no significant differences in fatty acid profiles between the wild-type strain and the recombinant strain. Over 94 % of total fatty acids were C16:0, C18:0, C16:1, C18:1, and C18:2. Our results indicated that heterologous expression of NADP+-dependent ME involved in fatty acid biosynthesis indeed increased the lipid accumulation in the oleaginous yeast R. glutinis.






Similar content being viewed by others
References
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Chaney AL, Marbach EP (1962) Modified reagents for the determination of ammonium and urea. Clin Chem 8:130–132
Chang GG, Tong L (2003) Structure and function of malic enzymes, a new class of oxidative decarboxylases. Biochemistry 42:12721–12733
Easterling ER, French WT, Hernandez R, Licha M (2009) The effect of glycerol as a sole and secondary substrate on the growth and fatty acid composition of Rhodotorula glutinis. Bioresource Technol 100:356–361
Gong Z, Wang Q, Shen HW, Hu CM, Jin GJ, Zhao ZB (2012) Co-fermentation of cellobiose and xylose by Lipomyces starkeyi for lipid production. Bioresource Technol 117:20–24
Goodridge AG, Klautky SA, Fantozzi DA, Baillie RA, Hodnett DW,Chen WZ, Thurmond DC, Xu G, Roncero C (1996) Nutritional and hormonal regulation of expression of the gene for malic enzyme. Prog Nucleic Acid Re 52:89–122
Handke P, Lynch SA, Gill RT (2011) Application and engineering of fatty acid biosynthesis in Escherichia coli for advanced fuels and chemicals. Metab Eng 13:28–37
Hill S, Winning B, Jenner H, Knorpp C, Leaver C (1996) Role of NAD+-dependent 'malic' enzyme and pyruvate dehydrogenase complex in leaf metabolism. Biochem Soc T 24:743–746
Hsu RY, Lardy HA (1969) Malic enzyme. Methods Enzymol 13:230–235
Klabunde J, Kunze G, Gellissen G, Hollenberg CP (2003) Integration of heterologous genes in several yeast species using vectors containing a Hansenula Polymorpha-derived rDNA-targeting element. FEMS yeast Res 4:185–193
Li YH, Adams IP, Wynn JP (2005) Cloning and characterization of a gene encoding a malic enzyme involved in anaerobic growth in Mucor circinelloides. Mycol Res 109:461–468
Liu B, Zhao ZB (2007) Biodiesel production by direct methanolysis of oleaginous microbial biomass. J Chem Technol Biotechnol 82:775–780
Lopes TS, Klootwijk J, Veenstra AE, van der Aar PC, van Heerikhuizen H, Raue HA, Planta RJ (1989) High-copy-number integration into the ribosomal DNA of Saccharomyces cerevisiae: a new vector for high-level expression. Gene 79:199–206
Lu XF, Vora H, Khosla C (2008) Overproduction of free fatty acids in E. coli: Implications for biodiesel production. Metab Eng 10:333–339
Luo XM (1995) Cloning and characterization of three Aspergillus niger promoters. Gene 163:127–131
Maria CR, Rosa R, Antonio RN (2002) Invertase from a strain of Rhodotorula glutinis. Phytochemistry 61:605–609
Meng X, Yang JM, Cao YJ, Li LZ, Jiang XL, Xu X, Liu W, Xian M, Zhang YW (2011) Increasing fatty acid production in E. coli by simulating the lipid accumulation of oleaginous microorganisms. J Ind Microbiol Biotechnol 38:919–925
Meng X, Yang JM, Xu X, Zhang L, Nie QJ, Xian M (2009) Biodiesel production from oleaginous microorganisms. Renew Energ 34:1–5
Misra S, Grosh A, Dutta J (1984) Production and composition of microbial fat from Rhodotorula glutinis. J Sci Food Agric 35:59–65
Moreadith RW, Lehninger AL (1984) The pathways of glutamate and glutamine oxidation by tumor cell mitochondria. Role of mitochondrial NAD(P)+-dependent malic enzyme. Journal of Biol Chem 259:6215–6221
Ratledge C (1979) Resources conservation by novel biological processes. I Grow fats from wastes. Chem Soc Rev 8:283–296
Ratledge C, Wynn JP (2000) Understanding microbial obesity. SIM News 50:181–185
Rittmann BE (2008) Opportunities for renewable bioenergy using microorganisms. Biotechnol Bioeng 100:203–212
Rodriguez-Frometa RA, Gutierrez A, Torres-Martinez S, Garre V (2012) Malic enzyme activity is not the only bottleneck for lipid accumulation in the oleaginous fungus Mucor circinelloides. Appl Microbiol Biotechnol. doi:10.1007/s00253-012-4432-2
Rose A, Harrison J (1970) Rhodotorula glutinis, fat synthesizing yeast. Yeast 3:433
Ruenwai R, Cheevadhanarak S, Laoteng K (2009) Overexpression of acetyl-CoA carboxylase gene of Mucor rouxii enhanced fatty acid content in Hansenula polymorpha. Mol Biotechnol 42:327–332
Sambrook J, Russell DW (2001) Molecular cloning a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schiestl RH, Gietz RD (1989) High efficiency transformation of intact yeast cells using single stranded nucleic acids as a carrier. Curr Genet 16:339–346
Scopione RC, de Camargo SS, Astolfi-Filho S, Schenberg ACG (1993) A new promoter probe vector for Saccharomyces cerevisiae using fungal glucoamylase cDNA as the reporter gene. Yeast 9:599–605
Song YD, Wynn JP, Li YH, Grantham D, Ratledge C (2001) A pre-genetic study of the isoforms of malic enzyme associated with lipid accumulation in Mucor circinelloides. Microbiology 147:1507–1515
Struhl K, Stinchcomb DT, Scherer S, Davis RW (1979) High frequency transformation of yeast: autonomous replication of hybrid DNA molecules. PNAS 76:1035–1039
Tamano K, Bruno KS, Karagiosis SA, Cully DE, Deng S, Collett JR, Umemura M, Koike H, Baker SE, Machida M (2012) Increased production of fatty acids and triglycerides in Aspergillus oryzae by enhancing expressions of fatty acid synthesis-related genes. Appl Microbiol Biotechnol. doi:10.1007/s00253-012-4193-y
Visser H, Vreugdenhil S, Bont JAM, Verdoes JC (2000) Cloning and characterization of epoxide hydrolase-encoding gene from Rhodotorula glutinis. Appl Microbiol Biotechnol 53:415–419
Vongsangnak W, Zhang YT, Chen W, Ratledge C, Song YD (2012) Annotation and analysis of malic enzyme genes encoding for multiple isoforms in the fungus Mucor circinelloides CBS 277.49. Biotechnol Lett 34:941–947
Wynn JP, Hamid AA, Ratledge C (1999) The role of malic enzyme in the regulation of lipid accumulation in filamentous fungi. Microbiology 145:1911–1917
Wynn JP, Kendrick A, Ratledge C (1997) Sesamol as an inhibitor of growth and lipid metabolism in Mucor circinelloides via its action on malic enzyme. Lipids 32:605–610
Wynn JP, Ratledge C (1997) Malic enzyme is a major source of NADPH for lipid accumulation by Aspergillus nidulans. Microbiology 143:253–257
Wynn JP, Ratledge C (2000) Evidence that the rate-limiting step for the biosynthesis of arachidonic acid in Mortierella alpina is at the level of the 18:3 to 20:3 elongase. Microbiology 146:2325–2331
Yech Y (1996) Single-cell protein of Rhodotorula sp. Y38 from ethanol, acetic acid and acetaldehyde. Biotechnol Lett 18:411–416
Zhang T, Gong F, Chi Z, Liu GL, Chi ZM, Sheng J, Li J, Wang XH (2009) Cloning and characterization of the inulinase gene from a marine yeast Pichia guilliermondii and its expression in Pichia pastoris. Anton Leeuw Int JG 95:13–22
Zhang Y, Adams P, Ratledge C (2007) Malic enzyme: the controlling activity for lipid production? Overexpression of malic enzyme in Mucor circinelloides leads to a 2.5-fold increase in lipid accumulation. Microbiology 153:2013–2025
Acknowledgments
This work was sponsored by the National Science Foundation of China (NSFC; No.3087221) and Nanyang Qiwei Microecology Gene Science and Technology Development Co., Ltd.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Z., Sun, H., Mo, X. et al. Overexpression of malic enzyme (ME) of Mucor circinelloides improved lipid accumulation in engineered Rhodotorula glutinis . Appl Microbiol Biotechnol 97, 4927–4936 (2013). https://doi.org/10.1007/s00253-012-4571-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4571-5