Abstract
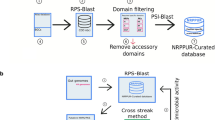
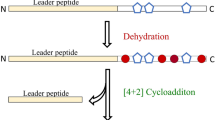
Identification of secondary metabolites produced by cryptic gene in bacteria may be difficult, but in the case of nonribosomal peptide (NRP)-type secondary metabolites, this study can be facilitated by bioinformatic analysis of the biosynthetic gene cluster and tandem mass spectrometry analysis. To illustrate this concept, we used mass spectrometry-guided bioinformatic analysis of genomic sequences to identify an NRP-type secondary metabolite from Streptomyces peucetius ATCC 27952. Five putative NRPS biosynthetic gene clusters were identified in the S. peucetius genome by DNA sequence analysis. Of these, the sp970 gene cluster encoded a complete NRPS domain structure, viz., C-A-T-C-A-T-E-C-A-T-C-A-T-C domains. Tandem mass spectrometry revealed that the functional siderophore peptide produced by this cluster had a molecular weight of 644.4 Da. Further analysis demonstrated that the siderophore peptide has a cyclic structure and an amino acid composition of AchfOrn–Arg–hOrn–hfOrn. The discovery of functional cryptic genes by analysis of the secretome, especially of NRP-type secondary metabolites, using mass spectrometry together with genome mining may contribute significantly to the development of pharmaceuticals such as hybrid antibiotics.







Similar content being viewed by others
References
Ansari MZ, Yadav G, Gokhale RS, Mohanty D (2004) NRPS-PKS: a knowledge-based resource for analysis of NRPS/PKS megasynthases. Nucleic Acids Res 32(Web Server issue):W405–W413
Challis GL, Ravel J (2000) Coelichelin, a new peptide siderophore encoded by the Streptomyces coelicolor genome: structure prediction from the sequence of its non-ribosomal peptide synthetase. FEMS Microbiol Lett 187(2):111–114
Challis GL, Ravel J, Townsend CA (2000) Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem Biol 7(3):211–224
Essen SA, Bylund D, Holmstrom SJ, Moberg M, Lundstrom US (2006) Quantification of hydroxamate siderophores in soil solutions of podzolic soil profiles in Sweden. Biometals 19(3):269–282
Felnagle EA, Barkei JJ, Park H, Podevels AM, McMahon MD, Drott DW, Thomas MG (2010) MbtH-like proteins as integral components of bacterial nonribosomal peptide synthetases. Biochemistry 49(41):8815–8817
Fuchs R, Budzikiewicz H (2001) Rearrangement reactions in the electrospray ionization mass spectra of pyoverdins. Int J Mass Spectrom 210(1–3):603–612
Gillam AH, Lewis AG, Andersen RJ (1981) Quantitative-determination of hydroxamic acids. Anal Chem 53(6):841–844
Gross H, Stockwell VO, Henkels MD, Nowak-Thompson B, Loper JE, Gerwick WH (2007) The genomisotopic approach: a systematic method to isolate products of orphan biosynthetic gene clusters. Chem Biol 14(1):53–63
Hayen H, Volmer DA (2005) Rapid identification of siderophores by combined thin-layer chromatography/matrix-assisted laser desorption/ionization mass spectrometry. Rapid Commun Mass Spectrom 19(5):711–720
Lautru S, Challis GL (2004) Substrate recognition by nonribosomal peptide synthetase multi-enzymes. Microbiology 150(Pt 6):1629–1636
Lautru S, Deeth RJ, Bailey LM, Challis GL (2005) Discovery of a new peptide natural product by Streptomyces coelicolor genome mining. Nat Chem Biol 1(5):265–269
Malla S, Niraula NP, Liou K, Sohng JK (2009) Enhancement of doxorubicin production by expression of structural sugar biosynthesis and glycosyltransferase genes in Streptomyces peucetius. J Biosci Bioeng 108(2):92–98
McClerren AL, Cooper LE, Quan C, Thomas PM, Kelleher NL, van der Donk WA (2006) Discovery and in vitro biosynthesis of haloduracin, a two-component lantibiotic. Proc Natl Acad Sci USA 103(46):17243–17248
Muller G, Raymond KN (1984) Specificity and mechanism of ferrioxamine-mediated iron transport in Streptomyces pilosus. J Bacteriol 160(1):304–312
Qi W, Jia CX, He ZM, Qiao B (2007) Multistep tandem mass spectrometry for sequencing cyclic peptides axinastatin 1 and deducing fragmentation pathways. Acta Chimica Sin 65(3):233–238
Robbel L, Knappe TA, Linne U, Xie X, Marahiel MA (2010) Erythrochelin—a hydroxamate-type siderophore predicted from the genome of Saccharopolyspora erythraea. FEBS J 277(3):663–676
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160(1):47–56
Sharman GJ, Williams DH, Ewing DF, Ratledge C (1995a) Determination of the structure of exochelin MN, the extracellular siderophore from Mycobacterium neoaurum. Chem Biol 2(8):553–561
Sharman GJ, Williams DH, Ewing DF, Ratledge C (1995b) Isolation, purification and structure of exochelin MS, the extracellular siderophore from Mycobacterium smegmatis. Biochem J 305(Pt 1):187–196
Silakowski B, Kunze B, Muller R (2001) Multiple hybrid polyketide synthase/non-ribosomal peptide synthetase gene clusters in the myxobacterium Stigmatella aurantiaca. Gene 275(2):233–240
Simionato AV, de Souza GD, Rodrigues-Filho E, Glick J, Vouros P, Carrilho E (2006) Tandem mass spectrometry of coprogen and deferoxamine hydroxamic siderophores. Rapid Commun Mass Spectrom 20(2):193–199
Stegmann E, Rausch C, Stockert S, Burkert D, Wohlleben W (2006) The small MbtH-like protein encoded by an internal gene of the balhimycin biosynthetic gene cluster is not required for glycopeptide production. FEMS Microbiol Lett 262(1):85–92
Sudek S, Haygood MG, Youssef DT, Schmidt EW (2006) Structure of trichamide, a cyclic peptide from the bloom-forming cyanobacterium Trichodesmium erythraeum, predicted from the genome sequence. Appl Environ Microbiol 72(6):4382–4387
Wenzel SC, Muller R (2005) Formation of novel secondary metabolites by bacterial multimodular assembly lines: deviations from textbook biosynthetic logic. Curr Opin Chem Biol 9(5):447–458
Wenzel SC, Kunze B, Hofle G, Silakowski B, Scharfe M, Blocker H, Muller R (2005) Structure and biosynthesis of myxochromides S1-3 in Stigmatella aurantiaca: evidence for an iterative bacterial type I polyketide synthase and for module skipping in nonribosomal peptide biosynthesis. Chembiochem 6(2):375–385
Wong GB, Kappel MJ, Raymond KN, Matzanke B, Winkelmann G (1983) Coordination chemistry of microbial iron transport compounds. 24. Characterization of coprogen and ferricrocin, 2 ferric hydroxamate siderophores. J Am Chem Soc 105(4):810–815
Wong DK, Gobin J, Horwitz MA, Gibson BW (1996) Characterization of exochelins of Mycobacterium avium: evidence for saturated and unsaturated and for acid and ester forms. J Bacteriol 178(21):6394–6398
Zhang W, Heemstra JR Jr, Walsh CT, Imker HJ (2010) Activation of the pacidamycin PacL adenylation domain by MbtH-like proteins. Biochemistry 49(46):9946–9947
Acknowledgments
This work was supported by the Intelligent Synthetic Biology Center of Global Frontier Project funded by the Ministry of Education, Science and Technology (2011-0031960) and Basic Science Research Program (2009-0071169) and Priority Research Centers Program (20110031388) through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science, and Technology.
Conflict of interest
The authors have no conflicts of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, HM., Kim, BG., Chang, D. et al. Genome-based cryptic gene discovery and functional identification of NRPS siderophore peptide in Streptomyces peucetius . Appl Microbiol Biotechnol 97, 1213–1222 (2013). https://doi.org/10.1007/s00253-012-4268-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4268-9