Abstract
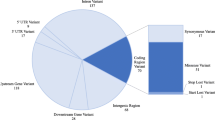
Asthma is a chronic inflammatory disorder of the airways, and a number of genetic loci are associated with the disease. Candidate gene association studies have been regarded as effective tools to study complex traits. Knowledge of the sequence variation and structure of the candidate genes is required for association studies. Thus, we investigated the genetic variants of 32 asthma candidate genes selected by colocalization of positional and functional candidate genes. We screened all exons and promoter regions of those genes using 12 healthy individuals and 12 asthma patients and identified a total of 418 single nucleotide polymorphisms (SNPs), including 270 known SNPs and 148 novel SNPs. Levels of nucleotide diversity varied from gene to gene (0.72×10−4–14.53×10−4), but the average nucleotide diversity between coding SNPs (cSNPs) and noncoding SNPs was roughly equivalent (4.63×10−4 vs 4.69×10−4). However, nucleotide diversity of cSNPs was strongly correlated to codon degeneracy. Nucleotide diversity was much higher at fourfold degenerate sites than at nondegenerate sites (9.42×10−4 vs 3.14×10−4). Gene-based haplotype analysis of asthma-associated genes in this study revealed that common haplotypes (frequency >5%) represented 90.5% of chromosomes, and they could be uniquely identified with five or fewer haplotype-tagging SNPs per gene. Therefore, our results may have important implications for the selection of asthma candidate genes and SNP markers for comprehensive association studies using large sample populations.



Similar content being viewed by others
References
Asher MI, Keil U, Anderson HR, Beasley R, Crane J, Martinez F, Mitchell EA, Pearce N, Sibbald B, Stewart AW (1995) International Study of Asthma and Allergies in Childhood (ISAAC): rationale and methods. Eur Respir J 8:483–491
Botstein D, Risch N (2003) Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease. Nat Genet 33(Suppl):228–237
Cargill M, Altshuler D, Ireland J, Sklar P, Ardlie K, Patil N, Shaw N, Lane CR, Lim EP, Kalyanaraman N, Nemesh J, Ziaugra L, Friedland L, Rolfe A, Warrington J, Lipshutz R, Daley GQ, Lander ES (1999) Characterization of single-nucleotide polymorphisms in coding regions of human genes. Nat Genet 22:231–238
Charlesworth B, Morgan MT, Charlesworth D (1993) The effect of deleterious mutations on neutral molecular variation. Genetics 134:1289–1303
Cookson W (2002) Genetics and genomics of asthma and allergic diseases. Immunol Rev 190:195–206
Halushka MK, Fan JB, Bentley K, Hsie L, Shen N, Weder A, Cooper R, Lipshutz R, Chakravarti A (1999) Patterns of single-nucleotide polymorphisms in candidate genes for blood-pressure homeostasis. Nat Genet 22:239–247
Illig T, Wjst M (2002) Genetics of asthma and related phenotypes. Paediatr Respir Rev 3:47–51
Johnson GC, Todd JA (2000) Strategies in complex disease mapping. Curr Opin Genet Dev 10:330–334
Kruglyak L, Nickerson DA (2001) Variation is the spice of life. Nat Genet 27:234–236
Li WH (1997) Molecular evolution. Sinauer, Sunderland, MA
Livingston RJ, von Niederhausern A, Jegga AG, Crawford DC, Carlson CS, Rieder MJ, Gowrisankar S, Aronow BJ, Weiss RB, Nickerson DA (2004) Pattern of sequence variation across 213 environmental response genes. Genome Res 14:1821–1831
Nachman MW, Bauer VL, Crowell SL, Aquadro CF (1998) DNA variability and recombination rates at X-linked loci in humans. Genetics 150:1133–1141
Nickerson DA, Tobe VO, Taylor SL (1997) PolyPhred: automating the detection and genotyping of single nucleotide substitutions using fluorescence-based resequencing. Nucleic Acids Res 25:2745–2751
Okuda T, Fujioka Y, Kamide K, Kawano Y, Goto Y, Yoshimasa Y, Tomoike H, Iwai N, Hanai S, Miyata T (2002) Verification of 525 coding SNPs in 179 hypertension candidate genes in the Japanese population: identification of 159 SNPs in 93 genes. J Hum Genet 47:387–394
Palmer LJ, Cookson WO (2000) Genomic approaches to understanding asthma. Genome Res 10:1280–1287
Qin ZS, Niu T, Liu JS (2002) Partition-ligation-expectation-maximization algorithm for haplotype inference with single-nucleotide polymorphisms. Am J Hum Genet 71:1242–1247
Risch N, Merikangas K (1996) The future of genetic studies of complex human diseases. Science 273:1516–1517
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Sebastiani P, Lazarus R, Weiss ST, Kunkel LM, Kohane IS, Ramoni MF (2003) Minimal haplotype tagging. Proc Natl Acad Sci U S A 100:9900–9905
Strachan T, Read AP (1999) Human molecular genetics, 2nd edn. The Bath Press Ltd., Bath, UK
Tabor HK, Risch NJ, Myers RM (2002) Opinion: candidate-gene approaches for studying complex genetic traits: practical considerations. Nat Rev, Genet 3:391–397
Toda M, Ono SJ (2002) Genomics and proteomics of allergic disease. Immunology 106:1–10
Wall JD, Pritchard JK (2003) Haplotype blocks and linkage disequilibrium in the human genome. Nat Rev Genet, 4:587–597
Xu J, Meyers DA, Ober C, Blumenthal MN, Mellen B, Barnes KC, King RA, Lester LA, Howard TD, Solway J, Langefeld CD, Beaty TH, Rich SS, Bleecker ER, Cox NJ; Collaborative Study on the Genetics of Asthma (2001) Genomewide screen and identification of gene–gene interactions for asthma-susceptibility loci in three US populations: collaborative study on the genetics of asthma. Am J Hum Genet 68:1437–1446
Acknowledgements
This study was supported by an intramural grant of the National Institute of Health, Korea.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Kim, JJ., Kim, HH., Park, JH. et al. Large-scale identification and characterization of genetic variants in asthma candidate genes. Immunogenetics 57, 636–643 (2005). https://doi.org/10.1007/s00251-005-0024-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-005-0024-y