Abstract
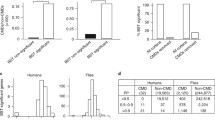
We investigate the extent by which the estimates of the rate of adaptive molecular evolution obtained by extending the McDonald–Kreitman test are biased if the species, subjected to analysis, diverged recently. We show that estimates can be biased if the nucleotide divergence between the species is low relative to within species variation, and that the magnitude of the bias depends on the rate of adaptive evolution and the distribution of fitness effects of new mutations. Bias appears to be because of three factors: (1) misattribution of polymorphism to divergence; (2) the contribution of ancestral polymorphism to divergence; and (3) different rates of fixation of neutral and advantageous mutations. If there is little adaptive molecular evolution, then slightly deleterious mutations inflate estimates of the rate of adaptive evolution, because these contribute proportionately more to polymorphism than to nucleotide divergence than neutral mutations. However, if there is substantial adaptive evolution, polymorphism contributing to apparent divergence may downwardly bias estimates. We propose a simple method for correcting the different contributions of slightly deleterious and neutral mutations to polymorphism and divergence, and apply it to datasets from several species. We find that estimates of the rate of adaptive molecular evolution from closely related species may be underestimates by ~10% or more. However, after the contribution of polymorphism to divergence is removed, the rate of adaptive evolution may still be overestimated as a consequence of ancestral polymorphism and time for fixation effects. This bias may be substantial if branch lengths are less than 10N e generations.




Similar content being viewed by others
References
Akey JM, Eberle MA, Rieder MJ, Carlson CS, Shriver MD (2004) Population history and natural selection shape patterns of genetic variation in 132 genes. PLoS Biol 2:e286
Andolfatto P (2005) Adaptive evolution of non-coding DNA in Drosophila. Nature 437:1149–1152
Bachtrog D (2008) Similar rates of protein adaptation in Drosophila miranda and D. melanogaster, two species with different current effective population sizes. BMC Evol Biol 8:334
Benton MJ, Donoghue PCJ (2007) Paleontological evidence to date the tree of life. Mol Biol Evol 24:26–53
Bierne N, Eyre-Walker A (2004) The genomic rate of adaptive amino acid substitution in Drosophila. Mol Biol Evol 21:1350–1360
Boyko AR, Williamson SH, Indap AR, Degenhardt JD, Hernandez RD, Lohmueller KE et al (2008) Assessing the evolutionary impact of amino acid mutations in the human genome. PLoS Genet 4:e1000083
Charlesworth B (1994) The effect of background selection against deleterious mutations on weakly selected, linked variants. Genet Res 63:213–227
Charlesworth J, Eyre-Walker A (2006) The rate of adaptive evolution in enteric bacteria. Mol Biol Evol 23:1348–1356
Charlesworth J, Eyre-Walker A (2008) The McDonald–Kreitman test and slightly deleterious mutations. Mol Biol Evol 25:1007–1015
Eyre-Walker A, Keightley PD (2009) Estimating the rate of adaptive molecular evolution in the presence of slightly deleterious mutations and population size change. Mol Biol Evol 26:2097–2108
Eyre-Walker A, Keightley PD, Smith NGC, Gaffney D (2002) Quantifying the slightly deleterious model of molecular evolution. Mol Biol Evol 19:2142–2149
Fay J, Wycoff GJ, Wu C-I (2001) Positive and negative selection on the human genome. Genetics 158:1227–1234
Fisher RA (1930) The genetical theory of natural selection. Clarendon Press, Oxford
Gossmann TI, Song B-H, Windsor AJ, Mitchell-Olds T, Dixon CJ, Kapralov MV, Filatov DA, Eyre-Walker A (2010) Genome wide analyses reveal little evidence for adaptive evolution in many plant species. Mol Biol Evol 27:1822–1832
Haddrill PR, Bachtrog D, Andolfatto P (2008) Positive and negative selection on noncoding DNA in Drosophila simulans. Mol Biol Evol 25:1825–1834
Halligan DL, Oliver F, Eyre-Walker A, Harr B, Keightley PD (2010) Evidence for pervasive adaptive protein evolution in wild mice. PLoS Genet 6:e1000825
Ingvarsson PK (2010) Natural selection on synonymous and nonsynonymous mutations shapes patterns of polymorphism in Populus tremula. Mol Biol Evol 27:650–660
Keightley PD, Eyre-Walker A (2007) Joint inference of the distribution of fitness effects of deleterious mutations and population demography based on nucleotide polymorphism frequencies. Genetics 177:2251–2261
Kousathanas A, Oliver F, Halligan DL, Keightley PD (2011) Positive and negative selection on noncoding DNA close to protein-coding genes in wild house mice. Mol Biol Evol 28:1183–1191
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351:652–654
Michaux J, Reyes A, Catzeflis F (2001) Evolutionary history of the most speciose mammals: molecular phylogeny of muroid rodents. Mol Biol Evol 18:2017–2031
Powell JR, DeSalle R (1995) Drosophila molecular phylogenies and their uses. Evol Biol 28:87–138
Sawyer SA, Hartl DL (1992) Population genetics of polymorphism and divergence. Genetics 132:1161–1176
Schneider A, Charlesworth B, Eyre-Walker A, Keightley PD (2011) A method for inferring the rate of occurrence and fitness effects of advantageous mutations. Genetics 189:1427–1437
Shapiro JA, Huang W, Zhang C, Hubisz MJ, Lu J, Turissini DA, Fang S, Wang H-Y, Hudson RR, Nielsen R, Chen Z, Wu C-I (2007) Adaptive genic evolution in the Drosophila genomes. Proc Natl Acad Sci USA 104:2271–2276
Welch JJ (2006) Estimating the genome-wide rate of adaptive protein evolution in Drosophila. Genetics 173:821–837
Acknowledgments
PDK acknowledges the grant support from the BBSRC and Wellcome Trust. The authors thank Dan Halligan for helpful comments in the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Keightley, P.D., Eyre-Walker, A. Estimating the Rate of Adaptive Molecular Evolution When the Evolutionary Divergence Between Species is Small. J Mol Evol 74, 61–68 (2012). https://doi.org/10.1007/s00239-012-9488-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-012-9488-1