Abstract
Purpose
Telomerase reverse transcriptase (TERT) promoter mutation status is an important biomarker for the precision diagnosis and prognosis prediction of lower grade glioma (LGG). This study aimed to construct a radiomic signature to noninvasively predict the TERT promoter status in LGGs.
Methods
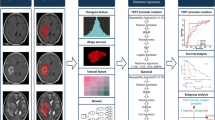
Eighty-three local patients with pathology-confirmed LGG were retrospectively included as a training cohort, and 33 patients from The Cancer Imaging Archive (TCIA) were used as for independent validation. Three types of regions of interest (ROIs), which covered the tumor, peri-tumoral area, and tumor plus peri-tumoral area, were delineated on three-dimensional contrast-enhanced T1 (3D-CE-T1)-weighted and T2-weighted images. One hundred seven shape, first-order, and texture radiomic features from each modality under each ROI were extracted and selected through least absolute shrinkage and selection operator. Radiomic signatures were constructed with multiple classifiers and evaluated using receiver operating characteristic (ROC) analysis. The tumors were also stratified according to IDH status.
Results
Three radiomic signatures, namely, tumoral radiomic signature, tumoral plus peri-tumoral radiomic signature, and fusion radiomic signature, were built, all of which exhibited good accuracy and balanced sensitivity and specificity. The tumoral signature displayed the best performance, with area under the ROC curves (AUC) of 0.948 (0.903–0.993) in the training cohort and 0.827 (0.667–0.988) in the validation cohort. In the IDH subgroups, the AUCs of the tumoral signature ranged from 0.750 to 0.940.
Conclusion
The MRI-based radiomic signature is reliable for noninvasive evaluation of TERT promoter mutations in LGG regardless of the IDH status. The inclusion of peri-tumoral area did not significantly improve the performance.




Similar content being viewed by others
References
Lapointe S, Perry A, Butowski NA (2018) Primary brain tumours in adults. Lancet 392(10145):432–446. https://doi.org/10.1016/s0140-6736(18)30990-5
Arita H, Narita Y, Takami H, Fukushima S, Matsushita Y, Yoshida A, Miyakita Y, Ohno M, Shibui S, Ichimura K (2013) TERT promoter mutations rather than methylation are the main mechanism for TERT upregulation in adult gliomas. Acta Neuropathol 126(6):939–941. https://doi.org/10.1007/s00401-013-1203-9
Eckel-Passow JE, Lachance DH, Molinaro AM, Walsh KM, Decker PA, Sicotte H, Pekmezci M, Rice T, Kosel ML, Smirnov IV, Sarkar G, Caron AA, Kollmeyer TM, Praska CE, Chada AR, Halder C, Hansen HM, McCoy L, Bracci PM, Marshall R, Zheng S, Reis GF, Pico AR, O'Neill BP, Buckner JC, Giannini C, Huse JT, Perry A, Tihan T, Berger MS, Chang SM, Prados MD, Wiemels J, Wiencke JK, Wrensch MR, Jenkins RB (2015) Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N Engl J Med 372(26):2499–2508. https://doi.org/10.1056/NEJMoa1407279
Yang P, Cai J, Yan W, Zhang W, Wang Y, Chen B, Li G, Li S, Wu C, Yao K, Li W, Peng X, You Y, Chen L, Jiang C, Qiu X, Jiang T, CGGA project (2016) Classification based on mutations of TERT promoter and IDH characterizes subtypes in grade II/III gliomas. Neuro-oncology 18(8):1099–1108. https://doi.org/10.1093/neuonc/now021
Tefferi A, Lasho TL, Begna KH, Patnaik MM, Zblewski DL, Finke CM et al (2015) A pilot study of the telomerase inhibitor imetelstat for myelofibrosis. N Engl J Med 373(10):908–919. https://doi.org/10.1056/NEJMoa1310523
Ersoy TF, Keil VC, Hadizadeh DR, Gielen GH, Fimmers R, Waha A, Heidenreich B, Kumar R, Schild HH, Simon M (2017) New prognostic factor telomerase reverse transcriptase promotor mutation presents without MR imaging biomarkers in primary glioblastoma. Neuroradiology 59(12):1223–1231. https://doi.org/10.1007/s00234-017-1920-1
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, Sanduleanu S, Larue RTHM, Even AJG, Jochems A, van Wijk Y, Woodruff H, van Soest J, Lustberg T, Roelofs E, van Elmpt W, Dekker A, Mottaghy FM, Wildberger JE, Walsh S (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14(12):749–762. https://doi.org/10.1038/nrclinonc.2017.141
Hwan-Ho C, Hyunjin P (2017) Classification of low-grade and high-grade glioma using multi-modal image radiomics features. Conf Proc Ann Int Conf IEEE Eng Med Biol Soc IEEE Eng Med Biol Soc Ann Conf 2017:3081–3084. https://doi.org/10.1109/embc.2017.8037508
De Looze C, Beausang A, Cryan J, Loftus T, Buckley PG, Farrell M et al (2018) Machine learning: a useful radiological adjunct in determination of a newly diagnosed glioma’s grade and IDH status. J Neuro-Oncol 139(2):491–499. https://doi.org/10.1007/s11060-018-2895-4
Yu J, Shi Z, Lian Y, Li Z, Liu T, Gao Y, Wang Y, Chen L, Mao Y (2017) Noninvasive IDH1 mutation estimation based on a quantitative radiomics approach for grade II glioma. Eur Radiol 27(8):3509–3522. https://doi.org/10.1007/s00330-016-4653-3
Arita H, Kinoshita M, Kawaguchi A, Takahashi M, Narita Y, Terakawa Y, Tsuyuguchi N, Okita Y, Nonaka M, Moriuchi S, Takagaki M, Fujimoto Y, Fukai J, Izumoto S, Ishibashi K, Nakajima Y, Shofuda T, Kanematsu D, Yoshioka E, Kodama Y, Mano M, Mori K, Ichimura K, Kanemura Y (2018) Lesion location implemented magnetic resonance imaging radiomics for predicting IDH and TERT promoter mutations in grade II/III gliomas. Scientific reports 8. doi:https://doi.org/10.1038/s41598-018-30273-4
Grossmann P, Gutman DA, Dunn WD Jr, Holder CA, Aerts HJ (2016) Imaging-genomics reveals driving pathways of MRI derived volumetric tumor phenotype features in glioblastoma. BMC Cancer 16:611. https://doi.org/10.1186/s12885-016-2659-5
Li ZC, Bai H, Sun Q, Li Q, Liu L, Zou Y et al (2018) Multiregional radiomics features from multiparametric MRI for prediction of MGMT methylation status in glioblastoma multiforme: a multicentre study. Eur Radiol 28(9):3640–3650. https://doi.org/10.1007/s00330-017-5302-1
Liu Z, Wang Y, Liu X, Du Y, Tang Z, Wang K et al (2018) Radiomics analysis allows for precise prediction of epilepsy in patients with low-grade gliomas. NeuroImage Clin 19:271–278. https://doi.org/10.1016/j.nicl.2018.04.024
Zhou H, Vallières M, Bai HX, Su C, Tang H, Oldridge D, Zhang Z, Xiao B, Liao W, Tao Y, Zhou J, Zhang P, Yang L (2017) MRI features predict survival and molecular markers in diffuse lower-grade gliomas. Neuro-oncology 19(6):862–870. https://doi.org/10.1093/neuonc/now256
Kickingereder P, Neuberger U, Bonekamp D, Piechotta PL, Gotz M, Wick A et al (2018) Radiomic subtyping improves disease stratification beyond key molecular, clinical, and standard imaging characteristics in patients with glioblastoma. Neuro-Oncol 20(6):848–857. https://doi.org/10.1093/neuonc/nox188
Fan X, Wang Y, Liu Y, Liu X, Zhang C, Wang L, Li S, Ma J, Jiang T (2016) Brain regions associated with telomerase reverse transcriptase promoter mutations in primary glioblastomas. J Neuro-Oncol 128(3):455–462. https://doi.org/10.1007/s11060-016-2132-y
Yamashita K, Hatae R, Hiwatashi A, Togao O, Kikuchi K, Momosaka D et al (2019) Predicting TERT promoter mutation using MR images in patients with wild-type IDH1 glioblastoma. Diagn Interv Imaging. https://doi.org/10.1016/j.diii.2019.02.010
Suh HB, Choi YS, Bae S, Ahn SS, Chang JH, Kang S-G, Kim EH, Kim SH, Lee SK (2018) Primary central nervous system lymphoma and atypical glioblastoma: differentiation using radiomics approach. Eur Radiol 28(9):3832–3839
Kickingereder P, Wiestler B, Sahm F, Heiland S, Roethke M, Schlemmer H-P, Wick W, Bendszus M, Radbruch A (2014) Primary central nervous system lymphoma and atypical glioblastoma: multiparametric differentiation by using diffusion-, perfusion-, and susceptibility-weighted MR imaging. Radiology 272(3):843–850
Chan AK, Yao Y, Zhang Z, Chung NY, Liu JS, Li KK et al (2015) TERT promoter mutations contribute to subset prognostication of lower-grade gliomas. Mod Pathol Off J U S Canadian Acad Pathol Inc 28(2):177–186. https://doi.org/10.1038/modpathol.2014.94
Horbinski C, Kofler J, Kelly LM, Murdoch GH, Nikiforova MN (2009) Diagnostic use of IDH1/2 mutation analysis in routine clinical testing of formalin-fixed, paraffin-embedded glioma tissues. J Neuropathol Exp Neurol 68(12):1319–1325. https://doi.org/10.1097/NEN.0b013e3181c391be
Duffau H, Taillandier L (2015) New concepts in the management of diffuse low-grade glioma: proposal of a multistage and individualized therapeutic approach. Neuro-Oncol 17(3):332–342. https://doi.org/10.1093/neuonc/nou153
Suchorska B, Weller M, Tabatabai G, Senft C, Hau P, Sabel MC, Herrlinger U, Ketter R, Schlegel U, Marosi C, Reifenberger G, Wick W, Tonn JC, Wirsching HG (2016) Complete resection of contrast-enhancing tumor volume is associated with improved survival in recurrent glioblastoma-results from the DIRECTOR trial. Neuro-Oncol 18(4):549–556. https://doi.org/10.1093/neuonc/nov326
Jenkinson M, Beckmann CF, Behrens TE, Woolrich MW, Smith SM (2012) Fsl. NeuroImage 62(2):782–790. https://doi.org/10.1016/j.neuroimage.2011.09.015
Smith SM, Jenkinson M, Woolrich MW, Beckmann CF, Behrens TE, Johansen-Berg H et al (2004) Advances in functional and structural MR image analysis and implementation as FSL. NeuroImage 23(Suppl 1):S208–S219. https://doi.org/10.1016/j.neuroimage.2004.07.051
Woolrich MW, Jbabdi S, Patenaude B, Chappell M, Makni S, Behrens T, Beckmann C, Jenkinson M, Smith SM (2009) Bayesian analysis of neuroimaging data in FSL. NeuroImage 45(1 Suppl):S173–S186. https://doi.org/10.1016/j.neuroimage.2008.10.055
van Griethuysen JJM, Fedorov A, Parmar C, Hosny A, Aucoin N, Narayan V, Beets-Tan RGH, Fillion-Robin JC, Pieper S, Aerts HJWL (2017) Computational radiomics system to decode the radiographic phenotype. Cancer Res 77(21):e104–e107. https://doi.org/10.1158/0008-5472.can-17-0339
Tang J, Alelyani S, Liu H (2014) Feature selection for classification: a review. Data classification: algorithms and applications:37
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O et al (2011) Scikit-learn: machine learning in python. J Mach Learn Res 12:2825–2830
DeLong ER, DeLong DM, Clarke-Pearson DL (1988) Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44(3):837–845
Vittinghoff E, McCulloch CE (2007) Relaxing the rule of ten events per variable in logistic and cox regression. Am J Epidemiol 165(6):710–718. https://doi.org/10.1093/aje/kwk052
Nonoguchi N, Ohta T, Oh JE, Kim YH, Kleihues P, Ohgaki H (2013) TERT promoter mutations in primary and secondary glioblastomas. Acta Neuropathol 126(6):931–937. https://doi.org/10.1007/s00401-013-1163-0
Brat DJ, Verhaak RG, Aldape KD, Yung WK, Salama SR, Cooper LA et al (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372(26):2481–2498. https://doi.org/10.1056/NEJMoa1402121
Ceccarelli M, Barthel FP, Malta TM, Sabedot TS, Salama SR, Murray BA, Morozova O, Newton Y, Radenbaugh A, Pagnotta SM, Anjum S, Wang J, Manyam G, Zoppoli P, Ling S, Rao AA, Grifford M, Cherniack AD, Zhang H, Poisson L, Carlotti CG Jr, Tirapelli DP, Rao A, Mikkelsen T, Lau CC, Yung WK, Rabadan R, Huse J, Brat DJ, Lehman NL, Barnholtz-Sloan JS, Zheng S, Hess K, Rao G, Meyerson M, Beroukhim R, Cooper L, Akbani R, Wrensch M, Haussler D, Aldape KD, Laird PW, Gutmann DH, TCGA Research Network, Noushmehr H, Iavarone A, Verhaak RG (2016) Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164(3):550–563. https://doi.org/10.1016/j.cell.2015.12.028
Diplas BH, He X, Brosnan-Cashman JA, Liu H, Chen LH, Wang Z, Moure CJ, Killela PJ, Loriaux DB, Lipp ES, Greer PK, Yang R, Rizzo AJ, Rodriguez FJ, Friedman AH, Friedman HS, Wang S, He Y, McLendon R, Bigner DD, Jiao Y, Waitkus MS, Meeker AK, Yan H (2018) The genomic landscape of TERT promoter wildtype-IDH wildtype glioblastoma. Nat Commun 9(1):2087. https://doi.org/10.1038/s41467-018-04448-6
Li Y, Liu X, Qian Z, Sun Z, Xu K, Wang K, Fan X, Zhang Z, Li S, Wang Y, Jiang T (2018) Genotype prediction of ATRX mutation in lower-grade gliomas using an MRI radiomics signature. Eur Radiol 28(7):2960–2968. https://doi.org/10.1007/s00330-017-5267-0
Ren Y, Zhang X, Rui W, Pang H, Qiu T, Wang J, Xie Q, Jin T, Zhang H, Chen H, Zhang Y, Lu H, Yao Z, Zhang J, Feng X (2019) Noninvasive prediction of IDH1 mutation and ATRX expression loss in low-grade gliomas using multiparametric MR radiomic features. Journal of magnetic resonance imaging : JMRI 49(3):808–817. https://doi.org/10.1002/jmri.26240
Chen B, Wang H, Ge P, Zhao J, Li W, Gu H, Wang G, Luo Y, Chen D (2012) Gross total resection of glioma with the intraoperative fluorescence-guidance of fluorescein sodium. Int J Med Sci 9(8):708–714. https://doi.org/10.7150/ijms.4843
Verburg N, Baayen JC, Idema S, Klitsie MA, Claus S, de Jonge CS et al (2016) In vivo accuracy of a frameless stereotactic drilling technique for diagnostic biopsies and stereoelectroencephalography depth electrodes. World Neurosurg 87:392–398. https://doi.org/10.1016/j.wneu.2015.11.041
Wolpert F, Lareida A, Terziev R, Grossenbacher B, Neidert MC, Roth P et al (2019) Risk factors for the development of epilepsy in patients with brain metastasis. Neuro-Oncology. https://doi.org/10.1093/neuonc/noz172
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins GJ, Friedman H, Friedman A, Reardon D, Herndon J, Kinzler KW, Velculescu VE, Vogelstein B, Bigner DD (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360(8):765–773. https://doi.org/10.1056/NEJMoa0808710
Buckner JC, Shaw EG, Pugh SL, Chakravarti A, Gilbert MR, Barger GR et al (2016) Radiation plus procarbazine, CCNU, and vincristine in low-grade glioma. N Engl J Med 374(14):1344–1355. https://doi.org/10.1056/NEJMoa1500925
Mellinghoff IK, Cloughesy TF, Wen PY, Taylor JW, Maher EA, Arrillaga I et al. (2019) A phase I, open label, perioperative study of AG-120 and AG-881 in recurrent IDH1 mutant, low-grade glioma: results from cohort 1. American Society of Clinical Oncology
Brat DJ, Aldape K, Colman H, Holland EC, Louis DN, Jenkins RB, Kleinschmidt-DeMasters B, Perry A, Reifenberger G, Stupp R, von Deimling A, Weller M (2018) cIMPACT-NOW update 3: recommended diagnostic criteria for “diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma, WHO grade IV”. Acta Neuropathol 136(5):805–810. https://doi.org/10.1007/s00401-018-1913-0
Ivanidze J, Lum M, Pisapia D, Magge R, Ramakrishna R, Kovanlikaya I, Fine HA, Chiang GC (2019) MRI features associated with TERT promoter mutation status in glioblastoma. J Neuroimaging Off J Am Soc Neuroimaging 29(3):357–363. https://doi.org/10.1111/jon.12596
Funding
This study was funded by the Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (Grant number 2016-I2M-2-001), the Fundamental Research Funds for the Central Universities (Grant number 3332018029), and the National Natural Science Foundation of China (Grant number 81601485).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. For this type of study, formal consent is not required.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jiang, C., Kong, Z., Zhang, Y. et al. Conventional magnetic resonance imaging–based radiomic signature predicts telomerase reverse transcriptase promoter mutation status in grade II and III gliomas. Neuroradiology 62, 803–813 (2020). https://doi.org/10.1007/s00234-020-02392-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00234-020-02392-1