Abstract
Pancreatic diseases, such as pancreatitis and pancreatic cancer, remain the most threatening gastrointestinal diseases with a high mortality due to atypical symptoms. MicroRNA plays crucial roles in regulating metastasis and cell proliferation of pancreatic cancer, constituting important biomarkers for the early diagnosis of pancreatic cancers. Herein, we develop a sensitive and simple exosomal miRNA detection method with only a dual-hairpin-probe. In detail, the dual-hairpin-probe is constructed through combination of two functional sections for both target miRNA identification and signal amplification. With only one probe, the method possesses the capability to avoid interferences from concentration changes of other probes, and exhibits a higher stability which is demonstrated through the obtained low coefficients of variation (CV) of 6.73%. With let-7a as detection target, the LOD of the established method is determined to be 243 aM, while maintaining a high discriminating capability towards let-7a homogenous miRNAs.
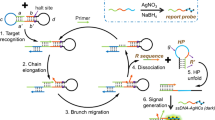
Graphical abstract






Similar content being viewed by others
References
Heckler M, Hackert T, Hu K, Halloran CM, Buchler MW, Neoptolemos JP. Severe acute pancreatitis: surgical indications and treatment. Langenbecks Arch Surg. 2021;406(3):521–35. https://doi.org/10.1007/s00423-020-01944-6.
Liu XH, Chen H, Tan RY, Luo C. Acute pancreatitis due to tacrolimus in kidney transplant and review of the literature. J Clin Pharm Ther. 2021;46(1):230–5. https://doi.org/10.1111/jcpt.13269.
Morrison AH, Byrne KT, Vonderheide RH. Immunotherapy and prevention of pancreatic cancer. Trends Cancer. 2018;4(6):418–28. https://doi.org/10.1016/j.trecan.2018.04.001.
Wu J, Cai J. Dilemma and challenge of immunotherapy for pancreatic cancer. Dig Dis Sci. 2021;66(2):359–68. https://doi.org/10.1007/s10620-020-06183-9.
Zhang L, Sanagapalli S, Stoita A. Challenges in diagnosis of pancreatic cancer. World J Gastroenterol. 2018;24(19):2047–60. https://doi.org/10.3748/wjg.v24.i19.2047.
Ariston Gabriel AN, Wang F, Jiao Q, Yvette U, Yang X, Al-Ameri SA, Du L, Wang YS, Wang C. The involvement of exosomes in the diagnosis and treatment of pancreatic cancer. Mol Cancer. 2020;19(1):132. https://doi.org/10.1186/s12943-020-01245-y.
Kamerkar S, LeBleu VS, Sugimoto H, Yang S, Ruivo CF, Melo SA, Lee JJ, Kalluri R. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature. 2017;546(7659):498–503. https://doi.org/10.1038/nature22341.
Zhou W, Zhou Y, Chen X, Ning T, Chen H, Guo Q, Zhang Y, Liu P, Zhang Y, Li C, Chu Y, Sun T, Jiang C. Pancreatic cancer-targeting exosomes for enhancing immunotherapy and reprogramming tumor microenvironment. Biomaterials. 2021;268:120546. https://doi.org/10.1016/j.biomaterials.2020.120546.
Lan B, Zeng S, Grutzmann R, Pilarsky C (2019) The role of exosomes in pancreatic cancer. Int J Mol Sci 20(18). https://doi.org/10.3390/ijms20184332.
Richards KE, Zeleniak AE, Fishel ML, Wu J, Littlepage LE, Hill R. Cancer-associated fibroblast exosomes regulate survival and proliferation of pancreatic cancer cells. Oncogene. 2017;36(13):1770–8. https://doi.org/10.1038/onc.2016.353.
Chen C, Tan R, Wong L, Fekete R, Halsey J. Quantitation of microRNAs by real-time RT-qPCR. Methods Mol Biol. 2011;687:113–34. https://doi.org/10.1007/978-1-60761-944-4_8.
Deng R, Zhang K, Li J. Isothermal amplification for microRNA detection: from the test tube to the cell. Acc Chem Res. 2017;50(4):1059–68. https://doi.org/10.1021/acs.accounts.7b00040.
Xu S, Chang Y, Wu Z, Li Y, Yuan R, Chai Y. One DNA circle capture probe with multiple target recognition domains for simultaneous electrochemical detection of miRNA-21 and miRNA-155. Biosens Bioelectron. 2020;149:111848. https://doi.org/10.1016/j.bios.2019.111848.
Wang F, Wang H, Zhang P, Su F, Wang H, Li Z. Ultrasensitive multiplexed detection of miRNA targets of interest based on encoding probe extension in improved cDNA library. Anal Chim Acta. 2021;1152:338281. https://doi.org/10.1016/j.aca.2021.338281.
He C, Wang M, Sun X, Zhu Y, Zhou X, Xiao S, Zhang Q, Liu F, Yu Y, Liang H, Zou G. Integrating PDA microtube waveguide system with heterogeneous CHA amplification strategy towards superior sensitive detection of miRNA. Biosens Bioelectron. 2019;129:50–7. https://doi.org/10.1016/j.bios.2019.01.003.
Masud MK, Umer M, Hossain MSA, Yamauchi Y, Nguyen NT, Shiddiky MJA. Nanoarchitecture frameworks for electrochemical miRNA detection. Trends Biochem Sci. 2019;44(5):433–52. https://doi.org/10.1016/j.tibs.2018.11.012.
Koo KM, Carrascosa LG, Shiddiky MJ, Trau M. Poly(A) Extensions of miRNAs for amplification-free electrochemical detection on screen-printed gold electrodes. Anal Chem. 2016;88(4):2000–5. https://doi.org/10.1021/acs.analchem.5b04795.
Wang R, Zhao X, Chen X, Qiu X, Qing G, Zhang H, Zhang L, Hu X, He Z, Zhong D, Wang Y, Luo Y. Rolling circular amplification (RCA)-assisted CRISPR/Cas9 cleavage (RACE) for highly specific detection of multiple extracellular vesicle microRNAs. Anal Chem. 2020;92(2):2176–85. https://doi.org/10.1021/acs.analchem.9b04814.
Zhang G, Zhang L, Tong J, Zhao X, Ren J. CRISPR-Cas12a enhanced rolling circle amplification method for ultrasensitive miRNA detection. Microchemical Journal. 2020;158(2020):105239. https://doi.org/10.1016/j.microc.2020.105239.
Li J, Liu J, Bi Y, Sun M, Bai J, Zhou M. Ultrasensitive electrochemiluminescence biosensing platform for miRNA-21 and MUC1 detection based on dual catalytic hairpin assembly. Anal Chim Acta. 2020;1105:87–94. https://doi.org/10.1016/j.aca.2020.01.034.
Gu J, Qiao Z, He X, Yu Y, Lei Y, Tang J, Shi H, He D, Wang K. Enzyme-free amplified detection of miRNA based on target-catalyzed hairpin assembly and DNA-stabilized fluorescent silver nanoclusters. Analyst. 2020;145(15):5194–9. https://doi.org/10.1039/d0an00545b.
Zhang Y, Zhang Q, Weng X, Du Y, Zhou X. NEase-based amplification for detection of miRNA, multiple miRNAs and circRNA. Anal Chim Acta. 2021;1145:52–8. https://doi.org/10.1016/j.aca.2020.12.024.
Lu W, Wang Y, Song S, Chen C, Yao B, Wang M. A fishhook probe-based rolling circle amplification (FP-RCA) assay for efficient isolation and detection of microRNA without total RNA extraction. Analyst. 2018;143(20):5046–53. https://doi.org/10.1039/c8an01544a.
Zhang M, Wang H, Wang H, Wang F, Li Z. CRISPR/Cas12a-assisted ligation-initiated loop-mediated isothermal amplification (CAL-LAMP) for highly specific detection of microRNAs. Anal Chem. 2021;93(22):7942–8. https://doi.org/10.1021/acs.analchem.1c00686.
Zhao X, Zhang L, Gao W, Yu X, Gu W, Fu W, Luo Y. Spatiotemporally controllable microRNA imaging in living cells via a near-infrared light-activated nanoprobe. ACS Appl Mater Interfaces. 2020;12(32):35958–66. https://doi.org/10.1021/acsami.0c10962.
Liu Y, Li S, Zhang L, Zhao Q, Li N, Wu Y. Catalytic hairpin assembly-assisted rolling circle amplification for high-sensitive telomerase activity detection. ACS Omega. 2020;5(20):11836–41. https://doi.org/10.1021/acsomega.0c01459.
Zhao X, Yuan Y, Liu X, Mao F, Xu G, Liu Q. A versatile platform for sensitive and label-free identification of biomarkers through an exo-III-assisted cascade signal amplification strategy. Anal Chem. 2022;94(4):2298–304. https://doi.org/10.1021/acs.analchem.1c05012.
Song J, Lee CY, Park HG. Ultrasensitive multiplexed miRNA detection based on a self-priming hairpin-triggered isothermal cascade reaction. Chem Commun (Camb). 2022;58(14):2279–82. https://doi.org/10.1039/d1cc06282d.
Wang J, Sun Y, Lau C, Lu J. Target-fueled catalytic hairpin assembly for sensitive and multiplex microRNA detection. Anal Bioanal Chem. 2020;412(13):3019–27. https://doi.org/10.1007/s00216-020-02531-w.
Povedano E, Ruiz-Valdepenas Montiel V, Gamella M, Serafin V, Pedrero M, Moranova L, Bartosik M, Montoya JJ, Yanez-Sedeno P, Campuzano S, Pingarron JM. A novel zinc finger protein-based amperometric biosensor for miRNA determination. Anal Bioanal Chem. 2020;412(21):5031–41. https://doi.org/10.1007/s00216-019-02219-w.
Funding
The project is funded by the Natural Science Foundation of Chongqing Science and Technology Commission (cstc2021jcyj-msxm2671 and cstc2018jcyjAX0758); the Natural Science Foundation of Chongqing Medical and Pharmaceutical College (ygz2019302).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ning, L., Cheng, H., Yu, F. et al. Construction of simple and sensitive pancreatitis related microRNA detection strategy via self-priming triggered cascade signal amplification. Anal Bioanal Chem 414, 5837–5844 (2022). https://doi.org/10.1007/s00216-022-04147-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04147-8