Abstract
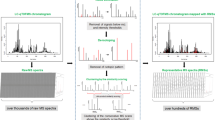
Medicinal plants are complex chemical systems containing thousands of secondary metabolites. The rapid classification and characterization of the components in medicinal plants using mass spectrometry (MS) remains an immense challenge. Herein, a novel strategy is presented for MS through the combination of solid-phase extraction (SPE), multiple mass defect filtering (MMDF) and molecular networking (MN). This strategy enables efficient classification and annotation of natural products. When combined with SPE and MMDF, the improved analytical method of MN can perform the rapid annotation of diverse natural products in Citrus aurantium according to the tandem mass spectrometry (MS/MS) fragments. In MN, MS2LDA can be initially applied to recognize substructures of natural products, according to the common fragmentation patterns and neutral losses in multiple MS/MS spectra. MolNetEnhancer was adopted here to obtain chemical classifications provided by ClassyFire. The results suggest that the integrated SPE-MMDF-MN method was capable of rapidly annotating a greater number of natural products from Citrus aurantium than the classical MN strategy alone. Moreover, SPE and MMDF enhanced the effectiveness of MN for annotating, classifying and distinguishing different types of natural products. Our workflow provides the foundation for the automated, high-throughput structural classification and annotation of secondary metabolites with various chemical structures. The developed approach can be widely applied in the analysis of constituents in natural products.
Graphical abstract








Similar content being viewed by others
References
Goodier J. Handbook of pharmaceutical natural products. Ref Rev. 2010;25(3):42–3.
Månsson M, Phipps RK, Gram L, Munro MHG, Larsen TO, Nielsen KF. Explorative solid-phase extraction (E-SPE) for accelerated microbial natural product discovery, dereplication, and purification. J Nat Prod. 2010;73(6):1126–32. https://doi.org/10.1021/np100151y.
Allard P-M, Péresse T, Bisson J, Gindro K, Marcourt L, Pham VC, et al. Integration of molecular networking and in-silico MS/MS fragmentation for natural products dereplication. Anal Chem. 2016;88(6):3317–23. https://doi.org/10.1021/acs.analchem.5b04804.
Shi Y, Zhan H, Zhong L, Yan F, Feng F, Liu W, et al. Total ion chromatographic fingerprints combined with chemometrics and mass defect filter to predict antitumor components of Picrasma quassioids. J Sep Sci. 2016;39(13):2633–41. https://doi.org/10.1002/jssc.201501410.
Shang Z, Cai W, Cao Y, Wang F, Wang Z, Lu J, et al. An integrated strategy for rapid discovery and identification of the sequential piperine metabolites in rats using ultra high-performance liquid chromatography/high resolution mass spectrometery. J Pharm Biomed Anal. 2017;146:387. https://doi.org/10.1016/j.jpba.2017.09.012.
Xing J, Zang M, Liu H. The application of a novel high-resolution mass spectrometry-based analytical strategy to rapid metabolite profiling of a dual drug combination in humans. Anal Chim Acta. 2017;993:38–46. https://doi.org/10.1016/j.aca.2017.08.047.
Gu Z-M, Wang L-Q, Wu J. Mass defect filter - a new tool to expedite screening and dereplication of natural products and generate natural product profiles. Nat Products J. 2011;1(2):135–45. https://doi.org/10.2174/2210315511101020135.
Wang M, Carver JJ, Phelan VV, Sanchez LM, Bandeira N. Sharing and community curation of mass spectrometry data with global natural products social molecular networking. Nat Biotechnol. 2016;34(8):828–37. https://doi.org/10.1038/nbt.3597.
Watrous J, Roach P, Alexandrov T, Heath BS, Yang JY, Kersten RD, et al. Mass spectral molecular networking of living microbial colonies. Proc Natl Acad Sci U S A. 2012;109(26):E1743–E52. https://doi.org/10.1073/pnas.1203689109.
Hautbergue T, Jamin EL, Costantino R, Tadrist S, Meneghetti L, Tabet J-C, et al. Combination of isotope labeling and molecular networking of tandem mass spectrometry data to reveal 69 unknown metabolites produced by Penicillium nordicum. Anal Chem. 2019;91(19):12191–202. https://doi.org/10.1021/acs.analchem.9b01634.
Nothias L-F, Nothias-Esposito M, da Silva R, Wang M, Protsyuk I, Zhang Z, et al. Bioactivity-based molecular networking for the discovery of drug leads in natural product bioassay-guided fractionation. J Nat Prod. 2018;81(4):758–67. https://doi.org/10.1021/acs.jnatprod.7b00737.
Reher R, Kuschak M, Heycke N, Annala S, Kehraus S, Dai H-F, et al. Applying molecular networking for the detection of natural sources and analogues of the selective Gq protein inhibitor FR900359. J Nat Prod. 2018;81(7):1628–35. https://doi.org/10.1021/acs.jnatprod.8b00222.
Ridder L, van der Hooft JJJ, Verhoeven S, de Vos RCH, Vervoort J, Bino RJ. In silico prediction and automatic LC–MSn annotation of green tea metabolites in urine. Anal Chem. 2014;86(10):4767–74. https://doi.org/10.1021/ac403875b.
Yu JS, Seo H, Kim GB, Hong J, Yoo HH. MS-based molecular networking of designer drugs as an approach for the detection of unknown derivatives for forensic and doping applications: a case of NBOMe derivatives. Anal Chem. 2019;91(9):5483–8. https://doi.org/10.1021/acs.analchem.9b00294.
Melnik AV, da Silva RR, Hyde ER, Aksenov AA, Vargas F, Bouslimani A, et al. Coupling targeted and untargeted mass spectrometry for metabolome-microbiome-wide association studies of human fecal samples. Anal Chem. 2017;89(14):7549–59. https://doi.org/10.1021/acs.analchem.7b01381.
Quinn RA, Nothias LF, Vining O, Meehan M, Esquenazi E, Dorrestein PC. Molecular networking as a drug discovery, drug metabolism, and precision medicine strategy. Trends Pharmacol Sci. 2017;38(2):143–54. https://doi.org/10.1016/j.tips.2016.10.011.
Ernst M, Kang KB, Caraballo Rodríguez A, Nothias L-F, Wandy J, Wang M, et al. MolNetEnhancer: enhanced molecular networks by integrating metabolome mining and annotation tools. Metabolites. 2019;9(7):144. https://doi.org/10.3390/metabo9070144.
Zhang H, Zhang D, Ray K, Zhu M. Mass defect filter technique and its applications to drug metabolite identification by high-resolution mass spectrometry. J Mass Spectrom. 2009;44(7):999–1016. https://doi.org/10.1002/jms.1610.
Wandy J, Zhu Y, van der Hooft JJJ, Daly R, Barrett MP, Rogers S. Ms2lda.org: web-based topic modelling for substructure discovery in mass spectrometry. Bioinformatics (Oxford, England). 2018;34(2):317–8. https://doi.org/10.1093/bioinformatics/btx582.
Nothias-Esposito M, Nothias LF, Da Silva RR, Retailleau P, Zhang Z, Leyssen P, et al. Investigation of premyrsinane and myrsinane esters in euphorbia cupanii and euphobia pithyusa with MS2LDA and combinatorial molecular network annotation propagation. J Nat Prod. 2019;82(6):1459–70. https://doi.org/10.1021/acs.jnatprod.8b00916.
Pilon AC, Gu H, Raftery D, Bolzani VDS, Lopes NP, Castro-Gamboa I, et al. Mass spectral similarity networking and gas-phase fragmentation reactions in the structural analysis of flavonoid glycoconjugates. Anal Chem. 2019;91(16):10413–23. https://doi.org/10.1021/acs.analchem.8b05479.
Fox Ramos AE, Pavesi C, Litaudon M, Dumontet V, Poupon E, Champy P, et al. CANPA: computer-assisted natural products anticipation. Anal Chem. 2019;91(17):11247–52. https://doi.org/10.1021/acs.analchem.9b02216.
da Silva RR, Dorrestein PC, Quinn RA. Illuminating the dark matter in metabolomics. Proc Natl Acad Sci U S A. 2015;112(41):12549–50. https://doi.org/10.1073/pnas.1516878112.
Aron A, Gentry E, McPhail K, Nothias L-F, Esposito M, Bouslimani A, et al. Reproducible molecular networking of untargeted mass spectrometry data using GNPS. Nat Protoc. 2020;15(6):1954–91. https://doi.org/10.1038/s41596-020-0317-5.
Funding
This work was supported by the National Key Research and Development Program of China (2017YFC1700906, 2017YFC1702900) and the Double Thousand Program of Jiangxi Province (jxsq2018102022).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 1.53 mb)
Rights and permissions
About this article
Cite this article
Wang, YK., Xiao, XR., Zhou, ZM. et al. A strategy combining solid-phase extraction, multiple mass defect filtering and molecular networking for rapid structural classification and annotation of natural products: characterization of chemical diversity in Citrus aurantium as a case study. Anal Bioanal Chem 413, 2879–2891 (2021). https://doi.org/10.1007/s00216-021-03201-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-021-03201-1