Abstract
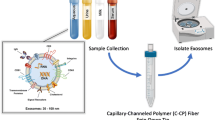
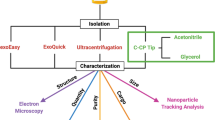
Exosomes are vesicles secreted by cells having a size range from 30 to 150 nm and carrying genetic materials that are important for intercellular functions, including cancer progression. Mounting evidence shows that tumor cells secrete more exosomes than normal cells. Thus, it is important to be able to efficiently isolate and quantify exosomes for potential use in clinical diagnostics, as well as to develop a deeper understanding of their role in intercellular processes. Current methods for exosome isolation and quantification are time-consuming and expensive. Few of these methods are able to combine exosome isolation and quantification into a singular operation scheme. However, a new efficient, rapid, and low-cost isolation and quantification method for exosomes in human urine samples using polyester (PET) capillary-channeled polymer (C-CP) fibers in a hydrophobic interaction chromatography (HIC) protocol has been developed. The process has been verified via scanning electron microscopy (SEM) before and after the capture of exosomes on the fiber surfaces. Sample load and elution rates were optimized to affect high resolution and throughput. Isolated exosomes were quantified based on a UV absorbance response curve created using a commercial human urine-derived exosome standard with an exosome concentration of 7.32 × 1011 mL−1. The loading capacity of a 30-cm C-CP PET column was ~ 7 × 1011 exosomes. An inter-injection washing method with PBS was developed to improve the reproducibility with a 2.9% RSD achieved for 7 complete isolation cycles.

Graphical abstract






Similar content being viewed by others
References
M HR, Bayraktar E, G KH, Abd-Ellah MF, Amero P, Chavez-Reyes A, et al. Exosomes: from garbage bins to promising therapeutic targets. Int J Mol Sci. 2017;18(3):538–63.
Duijvesz D, Luider T, Bangma CH, Jenster G. Exosomes as biomarker treasure chests for prostate cancer. Eur Urol. 2011;59(5):823–31.
Vlassov AV, Magdaleno S, Setterquist R, Conrad R. Exosomes: current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Bba-Gen Subjects. 2012;1820(7):940–8.
Simpson RJ, Lim JW, Moritz RL, Mathivanan S. Exosomes: proteomic insights and diagnostic potential. Expert Rev Proteomics. 2009;6(3):267–83.
Logozzi M, De Milito A, Lugini L, Borghi M, Calabro L, Spada M, et al. High levels of exosomes expressing CD63 and caveolin-1 in plasma of melanoma patients. PLoS One. 2009;4(4):e5219.
Taylor DD, Shah S. Methods of isolating extracellular vesicles impact down-stream analyses of their cargoes. Methods. 2015;87:3–10.
Shao H, Chung J, Issadore D. Diagnostic technologies for circulating tumour cells and exosomes. Biosci Rep. 2015;36(1):e00292.
Antimisiaris SG, Mourtas S, Marazioti A. Exosomes and exosome-inspired vesicles for targeted drug delivery. Pharmaceutics. 2018;10(4):218–58.
Zhang ZY, Wang CX, Li T, Liu Z, Li LJ. Comparison of ultracentrifugation and density gradient separation methods for isolating Tca8113 human tongue cancer cell line-derived exosomes. Oncol Lett. 2014;8(4):1701–6.
Baranyai T, Herczeg K, Onodi Z, Voszka I, Modos K, Marton N, et al. Isolation of exosomes from blood plasma: qualitative and quantitative comparison of ultracentrifugation and size exclusion chromatography methods. PLoS One. 2015;10(12).
Livshts MA, Khomyakova E, Evtushenko EG, Lazarev VN, Kulemin NA, Semina SE, et al. Isolation of exosomes by differential centrifugation: theoretical analysis of a commonly used protocol. Sci Rep-Uk. 2015;5:17319–31.
Jeppesen DK, Hvam ML, Primdahl-Bengtson B, Boysen AT, Whitehead B, Dyrskjot L, et al. Comparative analysis of discrete exosome fractions obtained by differential centrifugation. J Extracell Vesicles. 2014;3(1):25011–26.
Lobb R, Moller A. Size exclusion chromatography: a simple and reliable method for exosome purification. Extracellular Vesicles, Humana Press. 2017;1660:105–10.
Stranska R, Gysbrechts L, Wouters J, Vermeersch P, Bloch K, Dierickx D, et al. Comparison of membrane affinity-based method with size-exclusion chromatography for isolation of exosome-like vesicles from human plasma. J Transl Med. 2018;16:1–9.
Coren LV, Shatzer T, Ott DE. CD45 immunoaffinity depletion of vesicles from Jurkat T cells demonstrates that exosomes contain CD45: no evidence for a distinct exosome/HIV-I budding pathway. Retrovirology. 2008;5:64–8..
Yang F, Liao XZ, Tian Y, Li GY. Exosome separation using microfluidic systems: size-based, immunoaffinity-based and dynamic methodologies. Biotechnol J. 2017;12(4):1600699–706.
Zarovni N, Corrado A, Guazzi P, Zocco D, Lari E, Radano G, et al. Integrated isolation and quantitative analysis of exosome shuttled proteins and nucleic acids using immunocapture approaches. Methods. 2015;87:46–58.
Linares R, Tan S, Gounou C, Arraud N, Brisson AR. High-speed centrifugation induces aggregation of extracellular vesicles. J Extracell Vesicles. 2015;4:29509–15.
Diaz G, Bridges C, Lucas M, Cheng Y, Schorey JS, Dobos KM, et al. Protein digestion, ultrafiltration, and size exclusion chromatography to optimize the isolation of exosomes from human blood plasma and serum. Jove-J Vis Exp. 2018;(134):e57467.
An MR, Wu J, Zhu JH, Lubman DM. Comparison of an optimized ultracentrifugation method versus size-exclusion chromatography for isolation of exosomes from human serum. J Proteome Res. 2018;17(10):3599–605.
Hikita T, Miyata M, Watanabe R, Oneyama C. Sensitive and rapid quantification of exosomes by fusing luciferase to exosome marker proteins. Sci Rep. 2018;8:14035–49.
Kanwar SS, Dunlay CJ, Simeone DM, Nagrath S. Microfluidic device (ExoChip) for on-chip isolation, quantification and characterization of circulating exosomes. Lab Chip. 2014;14(11):1891–900.
He F, Wang J, Yin BC, Ye BC. Quantification of exosome based on a copper-mediated signal amplification strategy. Anal Chem. 2018;90(13):8072–9.
Franquesa M, Hoogduijn MJ, Ripoll E, Luk F, Salih M, Betjes MGH, et al. Update on controls for isolation and quantification methodology of extracellular vesicles derived from adipose tissue mesenchymal stem cells. Front Immunol. 2014;5:525–34.
Bruce TF, Slonecki TJ, Wang L, Huang SS, Powell RR, Marcus RK. Exosome isolation and purification via hydrophobic interaction chromatography using a polyester, capillary-channeled polymer fiber phase. Electrophoresis. 2019;40(4):571–81.
Randunu KM, Marcus RK. Microbore polypropylene capillary channeled polymer (C-CP) fiber columns for rapid reversed-phase HPLC of proteins. Anal Bioanal Chem. 2012;404(3):721–9.
Jiang L, Marcus RK. Microwave-assisted, grafting polymerization preparation of strong cation exchange nylon 6 capillary-channeled polymer fibers and their chromatographic properties. Anal Chim Acta. 2017;977:52–64.
Jiang L, Jin Y, Marcus RK. Polyethylenimine modified poly (ethylene terephthalate) capillary channeled-polymer fibers for anion exchange chromatography of proteins. J Chromatogr A. 2015;1410:200–9.
Stanelle RD, Marcus RK. Nylon-6 capillary-channeled polymer (C-CP) fibers as a hydrophobic interaction chromatography stationary phase for the separation of proteins. Anal Bioanal Chem. 2009;393(1):273–81.
Trang HK, Marcus RK. Application of protein A-modified capillary-channeled polymer polypropylene fibers to the quantitation of IgG in complex matrices. J Pharm Biomed Anal. 2017;142:49–58.
Trang HK, Schadock-Hewitt AJ, Jiang L, Marcus RK. Evaluation of loading characteristics and IgG binding performance of staphylococcal protein A on polypropylene capillary-channeled polymer fibers. J Chromatogr B Analyt Technol Biomed Life Sci. 2016;1015-1016:92–104.
Wang Z, Marcus RK. Determination of pore size distributions in capillary-channeled polymer fiber stationary phases by inverse size-exclusion chromatography and implications for fast protein separations. J Chromatogr A. 2014;1351:82–9.
Randunu KM, Dimartino S, Marcus RK. Dynamic evaluation of polypropylene capillary-channeled fibers as a stationary phase in high-performance liquid chromatography. J Sep Sci. 2012;35(23):3270–80.
Randunu KM, Marcus RK. Initial evaluation of protein throughput and yield characteristics on nylon 6 capillary-channeled polymer (C-CP) fiber stationary phases by frontal analysis. Biotechnol Prog. 2013;29(5):1222–9.
Ju ZH, Hei FL (2016) Introduction to exosomes derived from urine-derived human induced pluripotent stem cells: isolation, characterization and uptake. J Am Coll Cardiol 68 (16):C55-C55.
Belzer S, Romain C, Noerholm M, Skog J, Sinclair I, Fishbeck J, et al. Development of Q beta bacteriophage as an internal process control for isolation and extraction of microvesicles (including exosomes) from urine. J Mol Diagn. 2013;15(6):942.
Gonzales PA, Zhou H, Pisitkun T, Wang NS, Star RA, Knepper MA, et al. Isolation and purification of exosomes in urine. Urinary Proteome, Humana Press. 2010;641:89–99.
Schneider CA, Rasband WS, Eliceiri KW. NIH image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9(7):671–5.
Bhardwaj S, Day RA. Trifluoroethanol removes bound proteins from reversed-phase columns. Lc Gc N Am. 1999;17(4):354–6.
Majors RE. The cleaning and regeneration of reversed-phase HPLC columns. Lc Gc Eur. 2003;16(7):404–9.
Wang L, Marcus RK. Evaluation of protein separations based on hydrophobic interaction chromatography using polyethylene terephthalate capillary-channeled polymer (C-CP) fiber phases. J Chromatogr A. 2019;1585:161–71.
Jiang LW, Marcus RK. Comparison of analytical protein separation characteristics for three amine-based capillary-channeled polymer (C-CP) stationary phases. Anal Bioanal Chem. 2016;408(5):1373–83.
Wang L, Pierson MA, Marcus RK. Coupling of capillary-channeled polymer (C-CP) fibers for reversed phase liquid chromatography and ESI-MS for the determination of proteins in a urine matrix. Anal Method. 2016;8(48):8410–9.
Acknowledgments
The Gibson Foundation, the Prisma Health ITOR Biorepository, and the Greenville Hospital System are gratefully acknowledged. Special thanks are given to Dr. Larry Puls, Lori Allen, and Tina Pettry of Prisma Health for urine collection and storage. Matthew Haney and the Nanomedicines Characterization Core Facility at The University of North Carolina at Chapel Hill are gratefully acknowledged for completion of the NanoSight analysis. Special thanks are given to George Wetzel, Clemson University Electron Microscopy Facility, for assistance with EM; and to Tyler Slonecki for transportation and training.
Funding
This study received financial support for the chromatography development efforts from the National Science Foundation, Division of Chemistry, under grant CHE-1608663, and received financial support for the exosome isolation efforts from the Eppley Foundation for Scientific Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. All of the studies conducted in this manuscript were approved prior to their execution by the Clemson University Institutional Biosafety Committee, and have been performed in compliance with those approved protocols. All human samples were obtained via approved protocols and with informed patient consent as approved and required by the Prisma Hospital System Institutional Review Board (IRB). All patient samples were appropriately blinded for human subject protection.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Huang, S., Wang, L., Bruce, T.F. et al. Isolation and quantification of human urinary exosomes by hydrophobic interaction chromatography on a polyester capillary-channeled polymer fiber stationary phase. Anal Bioanal Chem 411, 6591–6601 (2019). https://doi.org/10.1007/s00216-019-02022-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-02022-7