Abstract
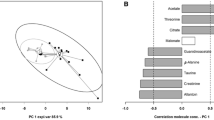
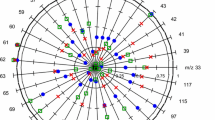
Exercise modulates the metabolome in urine or blood as demonstrated previously for humans and animal models. Using nuclear magnetic resonance (NMR) metabolomics, the present study compares the metabolic consequences of an exhaustive exercise at peak velocity (Vp) and at critical velocity (Vc) on mice. Since small-volume samples (blood and urine) were collected, dilution was necessary to acquire NMR spectra. Consequently, specific processing methods were applied before statistical analysis. According to the type of exercise (control group, Vp group and Vc group), 26 male mice were divided into three groups. Mice were sacrificed 2 h after the end of exercise, and urine and blood samples were drawn from each mouse. Proton NMR spectra were acquired with urine and deproteinized blood. The NMR data were aligned with the icoshift method and normalised using the probabilistic quotient method. Finally, data were analysed with the orthogonal projection of latent-structure analysis. The spectra obtained with deproteinized blood can neither discriminate the control mice from exercised mice nor discriminate according to the duration of the exercise. With urine samples, a significant statistical model can be estimated when comparing the control mice to both groups, Vc and Vp. The best model is obtained according to the exercise duration with all mice. Taking into account the spectral regions having the highest correlations, the discriminant metabolites are allantoin, inosine and branched-chain amino acids. In conclusion, metabolomic profiles assessed with NMR are highly dependent on the exercise. These results show that urine samples are more informative than blood samples and that the duration of the exercise is a more important parameter to influence the metabolomic status than the exercise velocity.


Similar content being viewed by others
References
Nicholson JK, Lindon JC (2008) Systems biology: metabonomics. Nature 455(7216):1054–1056. doi:10.1038/4551054a
Pan Z, Raftery D (2007) Comparing and combining NMR spectroscopy and mass spectrometry in metabolomics. Anal Bioanal Chem 387(2):525–527. doi:10.1007/s00216-006-0687-8
Enea C, Seguin F, Petitpas-Mulliez J, Boildieu N, Boisseau N, Delpech N, Diaz V, Eugene M, Dugue B (2010) (1)H NMR-based metabolomics approach for exploring urinary metabolome modifications after acute and chronic physical exercise. Anal Bioanal Chem 396(3):1167–1176. doi:10.1007/s00216-009-3289-4
Lehmann R, Zhao X, Weigert C, Simon P, Fehrenbach E, Fritsche J, Machann J, Schick F, Wang J, Hoene M, Schleicher ED, Haring HU, Xu G, Niess AM (2010) Medium chain acylcarnitines dominate the metabolite pattern in humans under moderate intensity exercise and support lipid oxidation. PLoS One 5(7):e11519. doi:10.1371/journal.pone.0011519
Lewis GD, Farrell L, Wood MJ, Martinovic M, Arany Z, Rowe GC, Souza A, Cheng S, McCabe EL, Yang E, Shi X, Deo R, Roth FP, Asnani A, Rhee EP, Systrom DM, Semigran MJ, Vasan RS, Carr SA, Wang TJ, Sabatine MS, Clish CB, Gerszten RE (2010) Metabolic signatures of exercise in human plasma. Sci Transl Med 2(33):33ra37. doi:10.1126/scitranslmed.3001006
Yan B, Jiye A, Wang G, Lu H, Huang X, Liu Y, Zha W, Hao H, Zhang Y, Liu L, Gu S, Huang Q, Zheng Y, Sun J (2009) Metabolomic investigation into variation of endogenous metabolites in professional athletes subject to strength-endurance training. J Appl Physiol 106(2):531–538. doi:10.1152/japplphysiol.90816.2008
Duggan GE, Hittel DS, Sensen CW, Weljie AM, Vogel HJ, Shearer J (2011) Metabolomic response to exercise training in lean and diet-induced obese mice. J Appl Physiol 110(5):1311–1318. doi:10.1152/japplphysiol.00701.2010
Huang CC, Lin WT, Hsu FL, Tsai PW, Hou CC (2010) Metabolomics investigation of exercise-modulated changes in metabolism in rat liver after exhaustive and endurance exercises. Eur J Appl Physiol 108(3):557–566. doi:10.1007/s00421-009-1247-7
Barba I, de Leon G, Martin E, Cuevas A, Aguade S, Candell-Riera J, Barrabes JA, Garcia-Dorado D (2008) Nuclear magnetic resonance-based metabolomics predicts exercise-induced ischemia in patients with suspected coronary artery disease. Magn Reson Med 60(1):27–32. doi:10.1002/mrm.21632
Hill DW, Poole DC, Smith JC (2002) The relationship between power and the time to achieve.VO(2max). Med Sci Sports Exerc 34(4):709–714
Froelicher VF Jr, Brammell H, Davis G, Noguera I, Stewart A, Lancaster MC (1974) A comparison of the reproducibility and physiologic response to three maximal treadmill exercise protocols. Chest 65(5):512–517
Taylor HL, Buskirk E, Henschel A (1955) Maximal oxygen intake as an objective measure of cardio-respiratory performance. J Appl Physiol 8(1):73–80
Day JR, Rossiter HB, Coats EM, Skasick A, Whipp BJ (2003) The maximally attainable VO2 during exercise in humans: the peak vs. maximum issue. J Appl Physiol 95(5):1901–1907. doi:10.1152/japplphysiol.00024.2003
Ferreira JC, Rolim NP, Bartholomeu JB, Gobatto CA, Kokubun E, Brum PC (2007) Maximal lactate steady state in running mice: effect of exercise training. Clin Exp Pharmacol Physiol 34(8):760–765. doi:10.1111/j.1440-1681.2007.04635.x
Moritani TA, Nagata HA, deVries HA, Muro M (1981) Critical power as measure of physical work capacity and anaerobic threshold. Ergonomics 24:339–350
Jones AM, Vanhatalo A, Burnley M, Morton RH, Poole DC (2010) Critical power: implications for determination of VO2max and exercise tolerance. Med Sci Sports Exerc 42(10):1876–1890. doi:10.1249/MSS.0b013e3181d9cf7f
Billat VL, Mouisel E, Roblot N, Melki J (2005) Inter- and intrastrain variation in mouse critical running speed. J Appl Physiol 98(4):1258–1263. doi:10.1152/japplphysiol.00991.2004
Savorani F, Tomasi G, Engelsen SB (2010) icoshift: a versatile tool for the rapid alignment of 1D NMR spectra. J Magn Reson 202(2):190–202. doi:10.1016/j.jmr.2009.11.012
Dieterle F, Ross A, Schlotterbeck G, Senn H (2006) Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in 1H NMR metabonomics. Anal Chem 78(13):4281–4290. doi:10.1021/ac051632c
Lerman I, Harrison BC, Freeman K, Hewett TE, Allen DL, Robbins J, Leinwand LA (2002) Genetic variability in forced and voluntary endurance exercise performance in seven inbred mouse strains. J Appl Physiol 92(6):2245–2255. doi:10.1152/japplphysiol.01045.2001
Pechlivanis A, Kostidis S, Saraslanidis P, Petridou A, Tsalis G, Mougios V, Gika HG, Mikros E, Theodoridis GA (2010) (1)H NMR-based metabonomic investigation of the effect of two different exercise sessions on the metabolic fingerprint of human urine. J Proteome Res 9(12):6405–6416. doi:10.1021/pr100684t
Miccheli A, Marini F, Capuani G, Miccheli AT, Delfini M, Di Cocco ME, Puccetti C, Paci M, Rizzo M, Spataro A (2009) The influence of a sports drink on the postexercise metabolism of elite athletes as investigated by NMR-based metabolomics. J Am Coll Nutr 28(5):553–564
Mikami T, Kitagawa J (2006) Intense exercise induces the degradation of adenine nucleotide and purine nucleotide synthesis via de novo pathway in the rat liver. Eur J Appl Physiol 96(5):543–550. doi:10.1007/s00421-005-0106-4
Shing CM, Peake JM, Ahern SM, Strobel NA, Wilson G, Jenkins DG, Coombes JS (2007) The effect of consecutive days of exercise on markers of oxidative stress. Appl Physiol Nutr Metab 32(4):677–685. doi:10.1139/H07-051
Stathis CG, Carey MF, Hayes A, Garnham AP, Snow RJ (2006) Sprint training reduces urinary purine loss following intense exercise in humans. Appl Physiol Nutr Metab 31(6):702–708. doi:10.1139/h06-074
Almstetter MF, Appel IJ, Gruber MA, Lottaz C, Timischl B, Spang R, Dettmer K, Oefner PJ (2009) Integrative normalization and comparative analysis for metabolic fingerprinting by comprehensive two-dimensional gas chromatography-time-of-flight mass spectrometry. Anal Chem 81(14):5731–5739. doi:10.1021/ac900528b
Yuan J, Doucette CD, Fowler WU, Feng XJ, Piazza M, Rabitz HA, Wingreen NS, Rabinowitz JD (2009) Metabolomics-driven quantitative analysis of ammonia assimilation in E. coli. Mol Syst Biol 5:302. doi:10.1038/msb.2009.60
Wightman PJ, Santer R, Ribes A, Dougherty F, McGill N, Thorburn DR, FitzPatrick DR (2003) MLYCD mutation analysis: evidence for protein mistargeting as a cause of MLYCD deficiency. Hum Mutat 22(4):288–300. doi:10.1002/humu.10264
Rodríguez D, Alcarraz-Vizán G, Díaz-Moralli S, Reed M, Gómez F, Falciani F, Günther U, Roca J, Cascante M (2011) Plasma metabolic profile in COPD patients: effects of exercise and endurance training. Metabolomics 8:508–516. doi:10.1007/s11306-011-0336-x
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Le Moyec, L., Mille-Hamard, L., Triba, M.N. et al. NMR metabolomics for assessment of exercise effects with mouse biofluids. Anal Bioanal Chem 404, 593–602 (2012). https://doi.org/10.1007/s00216-012-6165-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-012-6165-6