Abstract
The family of C3-like exoenzymes comprises seven bacterial ADP-ribosyltransferases of different origin. The common hallmark of these exoenzymes is the selective N-ADP-ribosylation of the low molecular mass GTP-binding proteins RhoA, B, and C and inhibition of signal pathways controlled by Rho GTPases. Therefore, C3-like exoenzymes were applied as pharmacological tools for analyses of cellular functions of Rho protein in numerous studies. Recent structural and functional analyses of C3-like exoenzymes provide detailed information on the molecular mechanisms and functional consequences of ADP-ribosylation catalyzed by these toxins. More recently additional non-enzymatic actions of C3-like ADP-ribosyltransferases have been identified showing that C3 transferases from Clostridium botulinum and Clostridium limosum form a GDI-like complex with the Ras-like low molecular mass GTPase Ral without ADP-ribosylation. These results add novel information on the molecular mode of action(s) of C3-like exoenzymes and are discussed in this review.
Similar content being viewed by others
Introduction
ADP-ribosylation is a biologically relevant protein modification for physiological (e.g., DNA repair, intracellular signaling, and extracellular regulation of immune cell signaling) and pathophysiological processes since many bacterial toxins were identified as mono-ADP-ribosyltransferases. After entry into target cells, which often involves highly efficient uptake mechanisms, the toxins modify eukaryotic target proteins with high specificity. In most cases, they grossly affect the biological functions of their targets. These properties are the reason for using the toxins as pharmacological tools (Aktories 2000). This applies especially to C3-like exoenzymes, whose pathogenetic role is still not well understood. Because C3 toxins modify selectively RhoA, B, and C, and thereby inactivate the molecular switches, they have been applied as pharmacological tools to elucidate the cellular functions of Rho GTPases in hundreds of papers for more than 20 years. To obtain more insights into the mode of action of these highly efficient toxins enhanced structural and biochemical analyses of C3 exoenzymes have been performed over the last few years. In this short review we will focus on the structure and function of C3-like ADP-ribosyltransferases. Moreover, the recently identified non-enzymatic interaction of C3 toxins with Ral GTPases is discussed and application of C3-like ADP-ribosyltransferases as pharmacological and cell biological tools is briefly reviewed.
Origin of C3-like ADP-ribosyltransferases
Clostridium botulinum C3 transferase (“C3bot1”), the prototype of C3-like exoenzymes, was initially discovered in strains of type C and D nearly 20 years ago (Aktories et al. 1987, 1988; Rubin et al. 1988). The term C3 indicates the third protein toxin beside the C1 neurotoxin and the binary actin-modifying C2 toxin, which are produced by Clostridium botulinum. C3bot2, an isoform of the C3 transferase with an identity of ∼65% was identified at nearly the same time (Nemoto et al. 1991). Together with botulinum neurotoxin C1 or D, C3 transferase (e.g., C3bot1) from Clostridium botulinum (e.g., strain C-468) is phage-encoded (Popoff et al. 1990; Sakaguchi et al. 2005). A second clostridial C3-like exoenzyme was found in Clostridium limosum some years later (Just et al. 1992).
However, expression of C3 transferases is not restricted to clostridia. Four of the seven toxins identified so far are produced by Gram-positive pathogens other than clostridia: Bacillus cereus produces the C3cer transferase with a sequence identity of ∼35% to C3bot1 (Just et al. 1995a). Three C3-like enzymes (originally designated EDIN) have been described from Staphylococcus aureus and termed in analogy with the other C3-like exoenzymes C3stau1, 2, and 3 (Inoue et al. 1991; Wilde et al. 2001; Yamaguchi et al. 2001). These exoenzymes are ∼35% identical with C3bot1 and 66–77% identical among each other (Fig. 1).
Alignment of C3-like transferases. The sequences of the seven C3-like ADP-ribosyltransferases are presented. Note: numbering of the C3-like transferases is without signal sequences. Clostridium botulinum C3 transferase type I (C3bot1; Acc. No. P15879), Clostridium botulinum C3 transferase type II (C3bot2; Acc. No. Q00901), Clostridium limosum C3 transferase (C3lim; Acc. No. Q46134), Bacillus cereus C3 transferase (C3cer; Acc. No. AJ429241.1), Staphylococcus aureus C3 transferase A, B, and C (C3stau1; Acc. No. P24121; C3stau2; Acc. No. BAC22946, C3stau3; Acc. No.;. NP-478345; also termed EDIN A, B, and C). Important motifs are highlighted: the ADP-ribosylation toxin-turn-turn (ARTT) loop is involved in protein substrate recognition and harbors conserved residues essential for the enzymatic activity (C3botQ172 and C3botE174); the phosphate-nicotinamide loop (PN-loop), which is involved in the binding of NAD; the STS-motif, which is conserved within the family of ADP-ribosyltransferases (C3stau isoforms possess an STQ motif), is considered to construct a β-strand-α-helix structure to maintain the reaction cavity; finally, several arginine residues involved in interaction with NAD are shown (C3botR51, C3botR88 and C3botR146). Comparing the sequences of prokaryotic and eukaryotic ADP-ribosyltransferases three residues are highly conserved among all these transferases: an NAD-interacting arginine residue (R), the first serine residues from the STS motif (S), and the catalytic glutamate (E) forming the RSE-motif of ADP-ribosyltransferases
Enzymatic activity and substrate specificity
C3-like exoenzymes catalyze the transfer of the ADP-ribose moiety from the co-substrate NAD to their substrate proteins RhoA, B, and C of the Rho family of low molecular mass GTPases (Fig. 2). RhoN41 was identified as acceptor amino acid for the ADP-ribosylation of RhoA, B, and C (Sekine et al. 1989). Other Rho-family members are poor (Rac for C3bot) or marginal substrates (Cdc42 for C3lim) for C3-like ADP-ribosyltransferases (Just et al. 1992; Wilde et al. 2003). The highest specificity for RhoA, B, and C was observed with C3cer (Wilde et al. 2003). The asparagine modification is a unique property of C3 toxins, since many bacterial ADP-ribosyltransferases modify arginine residues (e.g., cholera toxin, Pseudomonas exoenzyme S and T, C. botulinum C2 toxin; Moss and Vaughan 1977; Barbieri and Sun 2004; Vandekerckhove et al. 1988) or a cysteine residue (e.g., pertussis toxin; West et al. 1985). Recently, it was shown that the three isoforms C3stau1, 2, and 3 of Staphylococcus aureus differ in their substrate specificity. In addition to RhoA, B, and C they also modify the atypical Rho family member RhoE/Rnd3 (Wilde et al. 2001). RhoE and Rnd3 are constitutively active isoforms, which differ only in an addition of 15 amino acids in the N-terminus of Rnd3 and function as a RhoA antagonist (Foster et al. 1996; Wennerberg et al. 2003; Riento et al. 2003). However, the modification of RhoE/Rnd3 has a very slow reaction velocity compared with the ADP-ribosylation of RhoA and the functional relevance is not clear so far.
ADP-ribosylation of RhoA by C3. C3 transferases ADP-ribosylate RhoA at RhoAN41 (scheme). RhoA (pdb code 1FTN) is shown as a ribbon plot (left) and as a surface representation (right) with GDP as a space-filling model. The acceptor RhoAN41 (blue) is located adjacent to the switch I region in the “effector loop”, a region involved in the interaction with effector proteins of RhoA. RhoAR5, K6, E40, E47, E54, W58, and RhoAV38, F39, V43, which are involved in substrate–toxin interaction, are shown in pink. Further residues of RhoA, which are modified by bacterial toxins, are shown in green: RhoAT37, which is located in the switch I region, is mono-O-glucosylated by clostridial glucosylating toxins (e.g., C. difficile toxin A and B). This leads to inactivation of Rho GTPases. Deamidation of RhoAQ63, which is located in the switch II region, by E. coli cytotoxic necrotizing factors (CNFs) leads to constitutive activation of RhoA
In the absence of the protein substrates, Rho C3-like ADP-ribosyltransferases hydrolyze NAD into ADP-ribose and nicotinamide (Wilde et al. 2002b). NAD-glycohydrolase activity is also observed with other bacterial ADP-ribosylating toxins. However, its pathophysiological role is not clear.
The targets of C3-like exoenzymes are molecular switches
Rho proteins represent a discrete branch of the Ras superfamily of low molecular mass GTP-binding proteins and act as molecular switches in various signaling pathways (Jaffe and Hall 2005; Etienne-Manneville and Hall 2002; Burridge and Wennerberg 2004). The Rho subfamily consists of more than 20 members, which share 50% sequence identity and are ubiquitously expressed (Ridley 2000; Wennerberg and Der 2004). After binding of GTP Rho proteins are in the active conformation and inactive when bound to GDP (Fig. 3). Activation occurs by nucleotide exchange of Rho-GDP, which is catalyzed by guanine nucleotide exchange factors (GEFs; Rossman et al. 2005) or inhibited by the binding of guanine nucleotide dissociation inhibitors (GDIs; DerMardirossian and Bokoch 2005). Up to now, more than 60 GEF and only 3 GDI proteins for Rho have been identified. In the active GTP-bound form Rho proteins mediate various signals by interaction with a specific set of effector proteins. Rho proteins are master regulators of the actin cytoskeleton (Burridge and Wennerberg 2004; Hall 1998). Moreover, they are involved in cell and smooth muscle contraction, phagocytosis, polarity, activation of transcription, cell cycle progression and cell transformation (Bishop and Hall 2000; Etienne-Manneville and Hall 2002; Jaffe and Hall 2002; Wennerberg and Der 2004). The signal transduction of Rho is terminated by intrinsic hydrolysis of Rho-GTP, which is catalyzed by GTPase activating proteins (GAPs). More than 70 GAPs have been identified up to now. In the inactive GDP-bound form Rho proteins are extracted from the membrane by GDI, resulting in cytosolic accumulation of an inactive Rho–RhoGDI complex (Moon and Zheng 2003).
RhoA GTPase cycle and functional consequences of ADP-ribosylation by C3. Rho GTPases are inactive in the GDP-bound form and activated by GDP/GTP exchange, which is facilitated by guanine nucleotide exchange factors (GEFs). In the active form RhoA GTP interacts with a large variety of effectors to induce various cellular effects indicated. The active form of RhoA is terminated by hydrolysis of the bound GTP, which is enhanced by GTPase-activating proteins (GAPs). The inactive form is extracted from the membrane by guanine nucleotide exchange factors (GDI), which stabilize Rho GTPases in an inactive Rho–RhoGDI complex in the cytosol. RhoA is ADP-ribosylated by C3 in the GDI-free form and is then inactivated by a two-step mechanism: 1. ADP-ribosylation inhibits the activation of RhoA by GEF. 2. Binding of ADP-ribosylated RhoA to GDI is increased, which leads to a cytosolic accumulation of inactive Rho–RhoGDI complex (“trapping effect”)
Studies from recent years identified Rho GTPases as the preferred targets of several bacterial toxins. In addition to the ADP-ribosylation by C3, Rho GTPases are covalently modified by glucosylation (e.g., by clostridial glucosylating toxins; Aktories and Barbieri 2005; Just et al. 1995b; Aktories and Just 2005; Voth and Ballard 2005; Just and Gerhard 2004), deamidation (e.g., by E. coli cytotoxic necrotizing factor; Schmidt et al. 1997; Flatau et al. 1997), and transglutamination (Bordetella dermonecrotizing factor; Masuda et al. 2000) resulting in inactivation or activation of the GTPases respectively (Fig. 2). Other toxins influence the activity of Rho GTPases by mimicry of endogenous GAP activity, e.g., Yersinia YopE (von Pawel-Rammingen et al. 2000), Salmonella SptP (Fu and Galan 1999), Pseudomonas aeruginosa ExoS (Goehring et al. 1999) or endogenous GEF activity, e.g., Salmonella SopE (Hardt et al. 1998). Finally, it has been shown recently that Rho GTPase were cleaved proteolytically at the C-terminal isoprenylated cysteine by Yersinia YopT and thereby inactivated (Shao et al. 2002, 2003).
Functional consequence of the ADP-ribosylation
C3-like exoenzymes catalyze the transfer of the ADP-ribose moiety of NAD on a conserved asparagine residue in Rho (e.g., RhoAN41), which is located directly adjacent to the so-called switch I region (Fig. 2; Sekine et al. 1989; Maehama et al. 1994). The switch regions are structural hallmarks of guanine nucleotide-binding proteins and undergo dramatic conformational changes upon nucleotide exchange, which is required for the downstream signaling (Vetter and Wittinghofer 2001). ADP-ribosylation of this asparagine residue renders Rho biologically inactive and leads to a dramatic and lethal redistribution of actin in the target cell as a consequence (Chardin et al. 1989; Paterson et al. 1990; Wiegers et al. 1991). The mechanism underlying the inactivation of Rho by C3-mediated ADP-ribosylation was analyzed in great detail. First, it was shown that the ADP-ribosylation does not influence nucleotide-binding and hydrolysis (Sehr et al. 1998). This finding is in line with the crystal structure of RhoA-GTP, where RhoAN41 is surface located and does not interact with the nucleotide. Moreover, several crystal structures of Rho in complex with effector or regulator proteins respectively have revealed RhoAN41 as part of a conserved interaction site of low molecular GTP binding proteins (Dvorsky and Ahmadian 2004). It was thus reasonable to assume that ADP-ribosylation inactivates RhoA by inhibition of downstream signaling (i.e., blocking Rho-effector interaction) or by affecting the Rho-regulator interaction. From the follow-up studies it turned out that ADP-ribosylation inactivates RhoA by at least two steps: first by inhibition of the GEF-mediated nucleotide exchange of RhoA (e.g., Lbc; Sehr et al. 1998) and second by the trapping of ADP-ribosylated RhoA in the inactive RhoA–RhoGDI complex in the cytosol (Fig. 3; Genth et al. 2003a; Fujihara et al. 1997). Accordingly, translocation and activation of RhoA under physiological conditions is effectively blocked after ADP-ribosylation by C3-like transferases. However, inactivation of Rho by ADP-ribosylation can be overcome by treatment with the cytotoxic necrotizing factor CNF1. CNF1 deamidates RhoA at RhoAQ63, which is located in the switch II region, and thereby constitutively activates RhoA (Barth et al. 1999). In contrast, no evidence was found for the hypothesis that ADP-ribosylation prevents binding and activation of downstream effectors (Sehr et al. 1998; Genth et al. 2003b) or influences the GAP-mediated Rho inactivation (Sehr et al. 1998).
The pathogenetic role of C3-like exoenzymes
The pathogenetic role of C3-like exoenzymes is not well understood. One reason is the enigmatic up-take mechanism of the exoenzymes. Their structure is not in line with the composition of typical AB toxins, which possess an enzyme domain and a cell binding and translocation domain. C3-like toxins lack a specific transportation unit and, therefore, their cell accessibility is poor in many cell lines.
At least three possibilities for an effective uptake into their target cells are discussed. First, the concentration of C3 in the surrounding of the toxin-producing bacterium is very high allowing entry of the toxin into target cells over an extended time by unspecific up-take. Second, C3 toxins may be taken up by means of an additional bacterial pore-forming factor according to the mechanism proposed for the action of the Streptococcus pyogenes NAD glycohydrolase SPN. SPN appears to be specifically translocated into host cells by the pore-forming cytolysin Streptolysin O (Madden et al. 2001; Meehl and Caparon 2004). It remains to be clarified whether pore-forming cytolysins, which are frequently synthesized by C3-producing clostridia, are involved in the translocation of C3. Finally, at least for C3-producing Staphylococcus aureus an alternative mechanism has been suggested. For these bacteria, which are classically considered as extracellular pathogens, it was shown that they can act as intracellular pathogens, capable of invading and surviving within a broad range of non-phagocytic cells, including enterocytes, and endothelial and epithelial cells (Hess et al. 2003; Lowy 2000). Accordingly, recent studies indicate that these bacteria are able to invade host cells and secrete the C3 toxins directly into the host cytosol. Thus, it was shown that C3stau2 reaches the cytosol of the target cell during the staphylococcal invasion process without the need of a cell membrane translocation machinery (Molinari et al. 2006).
Once in the cytosol, C3 toxins should affect numerous Rho-dependent signal pathways in a manner as mentioned above. From a pathophysiological point of view it is important that C3 was shown to alter for example epithelial and endothelial barrier functions (Nusrat et al. 1995; Stamatovic et al. 2003), the signaling of immune cells including phagocytosis (Caron and Hall 1998), the production of cytokines (Chen et al. 2002; Dreikhausen et al. 2001), adhesion (Laudanna et al. 1996), de-adhesion (Liu et al. 2002), and migration of immune cells (Worthylake et al. 2001; Millan and Ridley 2005). A very recent study reported that C3stau-producing S. aureus form transcellular tunnels by inactivation of Rho, so-called macroapertures, which lead to the loss of barrier function in endothelial cells (Boyer et al. 2006). Other functions of Rho GTPases in immune cells, which are potentially targeted by bacterial toxins including C3 exoenzymes, have been reviewed recently (Aktories and Barbieri 2005; Cantrell 2003; Bokoch 2005).
C3 toxins are pharmacological tools
Because of the high specificity for Rho GTPases and their effective inactivation by ADP-ribosylation, C3-like exoenzymes were widely used as Rho inhibitors in many pharmacological and cell biological studies (Aktories et al. 2004). The application of C3-like ADP-ribosyltransferases as tools is limited by their poor cell accessibility. To overcome the poor cell accessibility of C3-like ADP-ribosyltransferase several strategies were successfully applied. Quite often the toxins were introduced by microinjection (Paterson et al. 1990; Watanabe et al. 1997). Furthermore, several C3 toxin chimeras were generated. In one of the earlier approaches, C3bot was fused to the binding and translocation subunit of diphtheria toxin resulting in the construct DC3B (Aullo et al. 1993). In another approach the binary actin-ADP-ribosylating C2 toxin from Clostridium botulinum was employed to construct a chimeric fusion toxin. C2 toxin consists of the actin-ADP-ribosylating enzyme component C2I and the binding and translocation component C2II, which are both separated proteins (Ohishi et al. 1980; Aktories et al. 1986; Aktories and Wegner 1989). The N-terminal part (C2IN) of C2I, which interacts with the binding component C2II, was fused to full length C3lim or C3stau respectively (Haug et al. 2006; Barth et al. 1998). All mammalian cells studied so far are sensitive towards the C2–C3 fusion toxin (e.g., CHO, Hela and NIH-3T3 cells; Valderrama et al. 2001; Wahl et al. 2000; Meyer et al. 2000; Vischer et al. 2000). Typical cytotoxic effects were observed at 200 ng/ml C2IN-C3lim/C3stau and 200 ng/ml C2II after 1–2 h of incubation (Barth et al. 1998). Several studies have shown that the C-terminal fusion of short transport peptide sequences enhances the cellular uptake of C3 (Sauzeau et al. 2001; Park et al. 2003; Winton et al. 2002; Sahai and Olson 2006). In many studies, the C3 gene was introduced into eukaryotic target cells by transient and stable transfection using plasmids or by viral infection (Fujisawa et al. 1998; Hill et al. 1995; Caron and Hall 1998; Henning et al. 1997; Meacci et al. 1999; Genot et al. 1996).
Structural analysis of C3-like exoenzymes
Crystal structure analysis using X-ray diffraction is one of the most advanced methods available for obtaining high resolution structural information about biological macromolecules. In case of the C3-like exoenzymes, the structures of two family members have been solved so far.
C3bot1 was the first C3-like exoenzyme, which was successfully crystallized by Han et al. (2001). The overall structure of the toxin has a mixed α, β-fold: a central β-sandwich, which is formed by a perpendicular packing of a five-stranded β-sheet against a three stranded β-sheet surrounding the NAD-binding pocket (Fig. 4). Important for the architecture of this pocket are three residues C3botS134, T135 and C3botS136, which are conserved in all C3-like transferases (Note: numbering of C3bot is without the signal sequence). Together they form the so-called STS motif (in C3stau transferases an ST motif), which connects strand β3 with the PN loop and maintains the reaction cavity. This central cleft of C3bot is terminated on one side by a so called ARTT loop (Han et al. 2001), a double turn motif consisting of 10 residues. The ARTT loop was proposed as part of the active site center and is conserved in other ADP-ribosyltransferases (e.g., actin-modifying Bacillus cereus VIP 2). It contains the invariant catalytic glutamate C3botE174 as well as C3botQ172 and C3botF169 residues, which are necessary for the activity of C3-like transferases. In their model for the ADP-ribosylation of Rho by C3bot, Tainer and coworkers predicted that C3 recognizes its substrate mainly via the ARTT loop (Han et al. 2001). In that context C3botF169 was proposed to contribute to the binding of Rho via a hydrophobic interaction with a patch of Rho (RhoAV38,F39,V43), whereas C3botQ172 was proposed to interact with RhoN41, the acceptor for the ADP-ribose moiety.
Structure of C. botulinum C3 transferase (C3bot1). C3bot in complex with NAD (pdb code 1GZF) is displayed as ribbon plot (left). Secondary structure elements are color-coded (α-helices, blue, β-sheets, orange) and labeled according to Han et al. (2001). ARTT loop and PN loop, which enclose the active site are shown in yellow, with the catalytic residues C3botQ172 and C3botE174 as stick models; the newly defined helix-loop-helix motif (HLH motif, green) mediates the non-enzymatic interaction of C3bot with Ral. A surface representation (right) indicates the Ral binding site (green) and three regions that are important for Rho recognition (yellow)
Ménétrey and coworkers compared structures of monomeric C3bot and C3bot complexed to NAD (Ménétrey et al. 2002). NAD binds in an elongated conformation into the central cleft of C3bot and contacts the ARTT loop as well as a further structural element, the so-called phosphate PN loop (Fig. 4). The PN loop (C3bot137–146) stabilizes NAD by interacting with its nicotinamide moiety and by compensating the charge of one phosphate group with a conserved arginine residue. The structural comparison of the monomeric C3bot and the C3bot-NAD complex revealed that binding of the cosubstrate NAD induces conformational changes in the C3 exoenzyme. The ARTT loop underwent an “ample swinging motion“ into the NAD-binding pocket. Most prominently, C3botQ172 was found to translocate from a surface exposed to a buried location. In the remodeled NAD-binding pocket C3botQ172 was then found to interact with the O-2′-hydroxyl ribose of the nicotinamide ribose (Fig. 4). The conformational change of C3botQ172 is thought to be important for correctly positioning Rho into the ternary Rho-NAD-C3 complex (Han et al. 2001; Ménétrey et al. 2002). Additionally, the catalytic glutamate C3botE174 was found to interact with O-2′-hydroxyl ribose of the nicotinamide ribose, which suggested the stabilization of an oxocarbenium transition state during the ADP-ribose transfer.
An analysis of overall conformational changes was reported by Evans et al. (2003). Comparing all available structural data, they concluded that the molecule possesses a rigid core formed largely from the β-strands and significant main-chain deviations are restricted to peripheral substructures, namely the ARTT loop, the PN loop, and the C-terminal end of the β1–α5 loop.
Recently, crystal structures of S. aureus C3stau2 were solved in the absence and presence of NAD (Evans et al. 2003). C3stau2 has a broadened substrate specificity compared with the other C3-like ADP-ribosyltransferases; in addition to RhoA, B, and C it also ADP-ribosylates the atypical Rho family member RhoE (Wilde et al. 2001).
The overall topology of C3stau2 is similar to that of C3bot despite a sequence similarity of only 35%. A deviation is found after strand β1, where C3stau2 has inserted two additional α-helices. A major difference between C3stau2 and C3bot1 was found in the conformation of the ARTT loop: in contrast to the cosubstrate-induced conformational changes observed for C3bot, the ARTT loop of C3stau, which contains two additional residues, folds in a fixed conformation in the absence and presence of NAD similar to that of the C3bot-NAD complex. Significant conformational changes after binding of NAD in C3stau were only found for the PN loop. A further difference in the structure of C3stau was the position of helix α3 from the helical bundle in comparison to C3bot. Together with the difference in length and conformation of the ARTT loop this could contribute to the additional modification of RhoE by C3stau2.
Structure–function analysis of the ADP-ribosylation of Rho by C3-like exoenzymes
Beside the crystallographic analysis a detailed structure–function analysis of C3-like exoenzymes (and RhoA) has been performed over the last few years. The available structural data in combination with a detailed biochemical analysis provides precise information about the binding of NAD, the substrate recognition and the mode of action of ADP-ribosylation of C3-like ADP-ribosyltransferases. Some important residues and their function in C3 toxins as well as in the substrate RhoA are discussed in the following paragraph.
The structural analysis showed that a conserved glutamate residue, the so-called catalytic glutamate, is involved in the binding of the nicotinamide ribose of NAD by hydrogen bonds with the O2-hydroxyl group (Fig. 4). Furthermore, this glutamate residue was proposed to stabilize an oxocarbenium transition state during the ADP-ribose transfer. The biochemical analysis supported the essential role of this residue for the enzymatic activity. Exchange of the catalytic glutamate in C3-like ADP-ribosyltransferases blocked the ADP-ribosylation of RhoA as well as the NAD hydrolysis in the absence of RhoA (Aktories et al. 1995; Böhmer et al. 1996; Wilde et al. 2002b, 2003; Han et al. 2001).
A conserved glutamine residue is located in turn 2 of the ARTT loop in all C3-like ADP-ribosyltransferases separated by a variable amino acid “X” from the catalytic glutamate. This sequence was termed QXE motif and is typical of non-arginine-modifying ADP-ribosyltransferases, e.g., the asparagine-modifying C3-like ADP-ribosyltransferases (Fig. 1). Exchange of this glutamine residue to alanine or glutamate blocked ADP-ribosylation of Rho; surprisingly, the binding of Rho and NAD and hydrolysis of NAD were hardly affected in these mutant proteins (Ménétrey et al. 2002; Wilde et al. 2002b, 2003). Building on the structural data a role of the glutamine residue in positioning of the acceptor RhoAN41 during formation of the Rho–C3 complex was proposed (Wilde et al. 2002b, 2003). The function of the glutamine residue was analyzed in more detail recently. Exchange to glutamate, which is at the respective position in several arginine-modifying ADP-ribosyltransferases blocked the asparagine modification of RhoA (Vogelsgesang and Aktories 2006). However, this glutamine–glutamate mutation in several C3-like transferases (e.g., C3botQ172E or EXE-C3bot) allowed ADP-ribosylation of arginine residues by the enzymes. Thus, the single mutation of the QXE motif into an EXE motif changes the specificity of C3-like ADP-ribosyltransferases from asparagine to arginine. The results indicate that the glutamine residue of the QXE motif stabilizes the acceptor asparagine in RhoA (RhoAN41) during the ADP-ribosylation reaction (Fig. 5).
Model for the interaction of asparagine or arginine with WT-C3 (QXE)- or EXE-C3 (e.g., C3limQ172E). A scheme of the catalytic core of C3, containing the nicotinamide ribose part of NAD is shown. C3limQ172, which is located in the ARTT loop, recognizes RhoAN41 via two hydrogen bonds. Due to the Q–N interaction RhoAN41 is in a favorable position for a nucleophilic attack on C1 of the ribose (asterisks). EXE-C3lim recognizes arginine residues. This R–E interaction is favorable because it takes advantage of the polar and electrostatic characteristics of both amino acids and allows ADP-ribosylation. H-bonds are not necessary, but support the recognition of arginine residues by EXE-C3lim
The structural prerequisites of RhoA for the recognition by C3bot were analyzed by (Wilde et al. 2000). The region for the interaction of RhoA with C3bot was restricted to the N-terminal half of RhoA by generating chimeric proteins between RhoA and the non-substrate Rac1. Within the N-terminal half, an alignment of Rho and the non-substrate Rac revealed remarkable amino acid differences in the two regions: RhoAR5, K6 and RhoAE40, E47, E54 respectively. Subsequent introduction of RhoAR5, K6, E40, E47, E54 at the equivalent positions in Rac1 allows its ADP-ribosylation by C3bot with kinetics comparable to the modification of RhoA. Interestingly, it has been speculated that RhoAR5, E40 may have a general impact for the enzyme–protein substrate interaction of ADP-ribosyltransferases, since functional homolog residues were identified at the interface in the crystal structure of eukaryotic elongation factor 2 (eEF2) in complex with the ADP-ribosylating toxin exotoxin A from Pseudomonas aeruginosa (Jorgensen et al. 2005). Beside their putative role in substrate recognition, RhoAE40 stabilizes the acceptor amino acid RhoAN41 via hydrogen bonds on the surface of RhoA-GTP (Ihara et al. 1998). This interaction brings the side chain of RhoAN41 into a close vicinity with the indole ring of RhoW58, which may enhance the nucleophilicity of RhoAN41 for the ADP-ribosylation.
Further important residues of Rho for the interaction with C3 are RhoAV38, F39 and RhoAV43. Their side chains form a hydrophobic ring around RhoAN41, which is suggested to interact with the conserved F/Y residue of C3 (Han et al. 2001). Thus, exchange of the respective residues inhibited ADP-ribosylation of the mutant proteins (Wilde et al. 2000).
Non-enzymatic interaction of C3-like exoenzymes with Ral
All C3-like exoenzymes ADP-ribosylate and thereby inactivate specifically RhoA, B, and C from more than 150 identified low molecular GTP-binding proteins. In addition, C3-like ADP-ribosyltransferases interact non-enzymatically with Ral without ADP-ribosylating the GTPase (Wilde et al. 2002a; Pautsch et al. 2005). Ral belongs to the Ras branch of low molecular GTPases and is involved in transcriptional activation (Hernandez-Munoz et al. 2000), Ras-mediated cell transformation (Feig et al. 1996; Urano et al. 1996), vesicle trafficking (Shen et al. 2001) and cytoskeletal rearrangements (Jullien-Flores et al. 1995; Ohta et al. 1999).
The interaction of Ral and C3bot was first identified in initial substrate scans of C3bot. In the presence of Ral, which is not a substrate for C3-like exoenzymes, the ADP-ribosylation of RhoA by C3bot was reduced.
More recently, it was shown that this inhibitory effect was specific for Ral and that Ral binds with high affinity to C3bot and C3lim. In contrast, Ras, which is the closest homolog to Ral, does not interact with C3 (Pautsch et al. 2005). However, ADP-ribosylation of RhoA by C3cer and C3stau2 was less or not influenced in the presence of Ral. Furthermore, C3bot decreased activation of the effector protein phospholipase D by RalA (Wilde et al. 2002a).
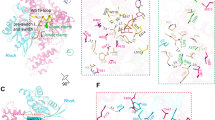
The molecular basis of the C3bot–RalA interaction was recently identified by two groups (Holbourn et al. 2005; Pautsch et al. 2005). The structure of a stochiometric RalA(GDP)-C3bot complex with an apparent KD value of ∼60 nM was determined by X-ray crystallography (Pautsch et al. 2005). In the structure of this complex C3bot inserts a previously unrecognized helix-loop-helix motif into a deep pocket formed predominantly by the switch II region and helix α3 of RalA (Fig. 6). The biological relevance of the interface deduced by Pautsch et al. was then verified by mutational analysis (Pautsch et al. 2005). The sequence of the C3 helix-loop-helix motif is less conserved in C3cer and the C3stau isoforms, explaining the specificity of Ral for the two other toxins. Notably, the Ral–C3 interface is located adjacent to regions that are proposed to be involved in Rho recognition (Fig. 4). On the other hand the C3 binding site in Ral does also not correspond with the ADP-ribosylation site in the switch I region of RhoA. In summary, the binding mode of Ral-C3 does not resemble the proposed substrate recognition mode of Rho by C3bot. It is highly likely that Ral inhibits the ADP-ribosylation by high affinity binding to an epitope of C3 in close proximity to the Rho-interaction site. Another possible explanation for the inhibitory effect is that Ral prevents conformational changes in C3bot, which occur during the ADP-ribosylation. It is noteworthy that the crystal structure of the RAL-C3 complex reported by Acharya and coworkers (Holbourn et al. 2005) predicts an interaction of the GTPase with the ARTT loop of C3. However, this model could not be supported by biochemical and mutational data (Pautsch et al. 2005).
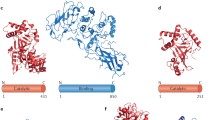
GTPase recognition of GDIs and a GDI-like acting toxin. The crystal structures of Ral in complex with C3bot (Pautsch et al. 2005), the Rab-like GTPase Ypt1 in complex with RabGDI (Rak et al. 2003), and Cdc42 with RhoGDI (Hoffman et al. 2000) are shown in an identical orientation relative to the GDP-bound GTPase. All three regulators bind to the GTPase effector binding region, in particular switch II, but differ in their detailed molecular mode of action. GDIs (RhoGDI and RabGDI) and GDI-like toxin C3bot are shown as blue ribbons; GTPases are shown in gray, with the switch I and switch II regions in red. GDP is shown as a stick model. NAD bound to C3bot and the C-terminal geranylgeranyl moiety of Cdc42 are shown as stick models. Residues of the GTPases that interact with the GDI are colored green. The molecular mode of the GDI-like stabilization of GDP by C3bot is shown in the insert. C3bot is displayed as a cyan worm. RalA is shown in gray except for the switch I (blue) and switch II (red) regions. Interacting residues of RalA-GDP and C3bot are shown in stick representation with atoms other than carbon colored by atom type ©, yellow; O, red; N, blue; P, magenta). Water molecules are shown as red spheres. The Mg2+ ion is presented as a black sphere, hydrogen bonds are shown as yellow dotted lines
The Ral–C3 complex described in Pautsch et al. (2005) suggested a functional consequence for Ral: binding of the helix-loop-helix motif stabilizes the conformation of the switch regions of Ral with respect to the native structures where the switch regions are found disordered (Nicely et al. 2004). Although C3bot does not directly interact with the guanine diphosphate, it stabilizes nucleotide binding indirectly through its interaction with the switch II region of Ral. A similar mode of nucleotide stabilization had been observed previously for two GDIs of the Rab family proteins (Rak et al. 2003, 2004) and for RhoGDI (Hoffman et al. 2000), suggesting that C3bot could act to Ral in a GDI-like manner. A structural comparison of known GDI complexes with Ral amd C3bot is shown in Fig. 6. All three regulators bind to the GTPase effector binding region, but differ in their detailed molecular mode of action.
This hypothesis that C3bot exerts a GDI-like effect was corroborated by the biochemical analysis of the nucleotide exchange of Ral with fluorescently labeled mant nucleotides in the presence of C3bot. It turned out that the nucleotide binding of Ral was altered in the Ral–C3bot complex. The addition of C3bot resulted in the concentration-dependent inhibition of the nucleotide release of Ral, which was preloaded with mant GDP. This inhibition of GDP exchange and the subsequent trapping of GDP was previously found with RhoGDI and RhoGTPases and termed GDI effect (Herrmann et al. 1995; Ahmadian et al. 2002). In contrast, C3bot had no influence on the exchange of mant GTP from Ral. The latter finding was in line with the low affinity for the interaction of Ral(GTP) and C3bot (KD value > 10 μM). Also the structural alignment of the Ral(GDP)–C3bot complex with Ral(GTP) in complex with the effector protein sec5 revealed that the conformation of the switch regions in Ral(GTP) precluded binding to C3bot. Taken together, these data suggest that C3bot is a bifunctional toxin targeting Rho by ADP-ribosylation and Ral by a GDI-like manner, which blocks nucleotide exchange and effector interaction. However, it remains to be shown that the effects of C3 in intact cells are not only caused by inactivation of Rho GTPases, but also by its interaction with Ral.
Other non enzymatic interaction of C3-like exoenzymes
Recently, another C3 effect, which is independent of the ADP-ribosyltransferase activity, has been reported (Ahnert-Hilger et al. 2004). It is well-known that Rho proteins regulate neurite outgrowth (Dergham et al. 2002). Several studies showed that C3 prevents neurite retraction induced by activated RhoA (Grunwald and Klein 2002; Wahl et al. 2000; Lehmann et al. 1999). Surprisingly, Ahnert-Hilger and coworkers found that C3bot, but not other C3 exoenzymes, promote the axonal growth and branching independent of the enzyme activity (Ahnert-Hilger et al. 2004). Moreover, this effect depended on the extracellular application of the exoenzyme. Intracellularly expressed C3bot did not induce axon growth. They propose a novel neurotrophic function of C3bot independent of its transferase activity.
Conclusions
The structural and biochemical analysis of C3-like ADP-ribosyltransferases over the few last years has provided significant progress in further understanding the enzymatic and non-enzymatic actions of these toxins. Despite their rather low sequence identity the overall folding of C3 toxins is very similar. The ARTT loop, together with the PN loop and helix α3, are important and conserved structural elements in the active site. From the analysis of the substrate recognition of the related ADP-ribosyltransferases ExoS and ExoT from Pseudomonas aeruginosa it was proposed that the recognition of Rho by C3 toxins is mediated by these conserved regions. However, to prove this model it is of major importance to solve the structure of C3 in complex with Rho. Novel properties of C3 toxins were identified deduced from the crystal structure of C3bot in complex with RalA and strengthened by biochemical data. The finding of the GDI-like non-enzymatical interaction of C3bot with Ral offered novel perspectives for the understanding of the mode of action and functional role of C3. Also important for the pathophysiological role of C3 was the finding that C3stau2 was secreted into the cytosol of eukaryotic cells after invasion of the bacteria. These data provide a potential explanation of how the C3-like exoenzymes, which lack a specific transportation unit, were able to enter their target cells.
References
Ahmadian MR, Wittinghofer A, Herrmann C (2002) Fluorescence methods in the study of small GTP-binding proteins. Methods Mol Biol 189:45–63
Ahnert-Hilger G, Holtje M, Grosse G, Pickert G, Mucke C, Nixdorf-Bergweiler B, Boquet P, Hofmann F, Just I (2004) Differential effects of Rho GTPases on axonal and dendritic development in hippocampal neurones. J Neurochem 90:9–18
Aktories K (2000) Bacterial protein toxins as tools in cell biology and pharmacology. In: Cossart P, Boquet P, Normark S, Rappuoli R (eds) Cellular microbiology. ASM Press, Washington, pp 221–237
Aktories K, Barbieri JT (2005) Bacterial cytotoxins: targeting eukaryotic switches. Nat Rev Microbiol 3:397–410
Aktories K, Just I (2005) Clostridial Rho-inhibiting protein toxins. Curr Top Microbiol Immunol 291:113–145
Aktories K, Wegner A (1989) ADP-ribosylation of actin by clostridial toxins. J Cell Biol 109:1385–1387
Aktories K, Bärmann M, Ohishi I, Tsuyama S, Jakobs KH, Habermann E (1986) Botulinum C2 toxin ADP-ribosylates actin. Nature 322:390–392
Aktories K, Weller U, Chhatwal GS (1987) Clostridium botulinum type C produces a novel ADP-ribosyltransferase distinct from botulinum C2 toxin. FEBS Lett 212:109–113
Aktories K, Rösener S, Blaschke U, Chhatwal GS (1988) Botulinum ADP-ribosyltransferase C3. Purification of the enzyme and characterization of the ADP-ribosylation reaction in platelet membranes. Eur J Biochem 172:445–450
Aktories K, Jung M, Böhmer J, Fritz G, Vandekerckhove J, Just I (1995) Studies on the active site structure of C3-like exoenzymes: involvement of glutamic acid in catalysis of ADP-ribosylation. Biochimie 77:326–332
Aktories K, Wilde C, Vogelsgesang M (2004) Rho-modifying C3-like ADP-ribosyltransferases. Rev Physiol Biochem Pharmacol 152:1–22
Aullo P, Giry M, Olsnes S, Popoff MR, Kocks C, Boquet P (1993) A chimeric toxin to study the role of the 21 kDa GTP binding protein rho in the control of actin microfilament assembly. EMBO J 12:921–931
Barbieri JT, Sun J (2004) Pseudomonas aeruginosa ExoS and ExoT. Rev Physiol Biochem Pharmacol 152:79–92
Barth H, Hofmann F, Olenik C, Just I, Aktories K (1998) The N-terminal part of the enzyme component (C2I) of the binary Clostridium botulinum C2 toxin interacts with the binding component C2II and functions as a carrier system for a Rho ADP-ribosylating C3-like fusion toxin. Infect Immun 66:1364–1369
Barth H, Olenik C, Sehr P, Schmidt G, Aktories K, Meyer DK (1999) Neosynthesis and activation of Rho by Escherichia coli cytotoxic necrotizing factor (CNF1) reverse cytopathic effects of ADP-ribosylated Rho. J Biol Chem 274:27407–27414
Bishop AL, Hall A (2000) Rho GTPases and their effector proteins. Biochem J 348:241–255
Böhmer J, Jung M, Sehr P, Fritz G, Popoff M, Just I, Aktories K (1996) Active site mutation of the C3-like ADP-ribosyltransferase from Clostridium limosum—analysis of glutamic acid 174. Biochemistry 35:282–289
Bokoch GM (2005) Regulation of innate immunity by Rho GTPases. Trends Cell Biol 15:163–171
Boyer L, Doye A, Rolando M, Flatau G, Munro P, Gounon P, Clement R, Pulcini C, Popoff MR, Mettouchi A, Landraud L, Dussurget O, Lemichez E (2006) Induction of transient macroapertures in endothelial cells through RhoA inhibition by Staphylococcus aureus factors. J Cell Biol 173:809–819
Burridge K, Wennerberg K (2004) Rho and Rac take center stage. Cell 116:167–179
Cantrell DA (2003) GTPases and T cell activation. Immunol Rev 192:122–130
Caron E, Hall A (1998) Identification of two distinct mechanisms of phagocytosis controlled by different Rho GTPases. Science 282:1717–1721
Chardin P, Boquet P, Madaule P, Popoff MR, Rubin EJ, Gill DM (1989) The mammalian G protein rho C is ADP-ribosylated by Clostridium botulinum exoenzyme C3 and affects actin microfilament in Vero cells. EMBO J 8:1087–1092
Chen LY, Zuraw BL, Liu FT, Huang S, Pan ZK (2002) IL-1 receptor-associated kinase and low molecular weight GTPase RhoA signal molecules are required for bacterial lipopolysaccharide-induced cytokine gene transcription. J Immunol 169:3934–3939
Dergham P, Ellezam B, Essagian C, Avedissian H, Lubell WD, McKerracher L (2002) Rho signaling pathway targeted to promote spinal cord repair. J Neurosci 22:6570–6577
DerMardirossian C, Bokoch GM (2005) GDIs: central regulatory molecules in Rho GTPase activation. Trends Cell Biol 15:356–363
Dreikhausen U, Varga G, Hofmann F, Barth H, Aktories K, Resch K, Szamel M (2001) Regulation by rho family GTPases of IL-1 receptor induced signaling: C3-like chimeric toxin and Clostridium difficile toxin B inhibit signaling pathways involved in IL-2 gene expression. Eur J Immmunol 31:1610–1619
Dvorsky R, Ahmadian MR (2004) Always look on the bright site of Rho: structural implications for a conserved intermolecular interface. EMBO Rep 5:1130–1136
Etienne-Manneville S, Hall A (2002) Rho GTPases in cell biology. Nature 420:629–635
Evans HR, Sutton JM, Holloway DE, Ayriss J, Shone CC, Acharya KR (2003) The crystal structure of C3stau2 from Staphylococcus aureus and its complex with NAD. J Biol Chem 278:45924–45930
Feig LA, Urano T, Cantor S (1996) Evidence for a Ras/Ral signaling cascade. Trends Biochem Sci 21:438–441
Flatau G, Lemichez E, Gauthier M, Chardin P, Paris S, Fiorentini C, Boquet P (1997) Toxin-induced activation of the G protein p21 Rho by deamidation of glutamine. Nature 387:729–733
Foster R, Hu K-Q, Lu Y, Nolan KM, Thissen J, Settleman J (1996) Identification of a novel human Rho protein with unusual properties: GTPase deficiency and in vivo farnesylation. Mol Cell Biol 16:2689–2699
Fu Y, Galan JE (1999) A Salmonella protein antagonizes Rac-1 and Cdc42 to mediate host-cell recovery after bacterial invasion. Nature 401:293–297
Fujihara H, Walker LA, Gong MC, Lemichez E, Boquet P, Somlyo AV, Somlyo AP (1997) Inhibition of RhoA translocation and calcium sensitization by in vivo ADP-ribosylation with the chimeric toxin DC3B. Mol Biol Cell 8:2437–2447
Fujisawa K, Madaule P, Ishizaki T, Watanabe G, Bito H, Saito Y, Hall A, Narumiya S (1998) Different regions of Rho determine Rho-selective binding of different classes of Rho target molecules. J Biol Chem 273:18943–18949
Genot E, Cleverley S, Henning S, Cantrell D (1996) Multiple p21ras effector pathways regulate nuclear factor of activated T cells. EMBO J 15:3923–3933
Genth H, Gerhard R, Maeda A, Amano M, Kaibuchi K, Aktories K, Just I (2003a) Entrapment of Rho ADP-ribosylated by Clostridium botulinum C3 exoenzyme in the Rho-guanine nucleotide dissociation inhibitor-1 complex. J Biol Chem 278:28523–28527
Genth H, Schmidt M, Gerhard R, Aktories K, Just I (2003b) Activation of phospholipase D1 by ADP-ribosylated RhoA. Biochem Biophys Res Commun 302:127–132
Goehring U-M, Schmidt G, Pederson KJ, Aktories K, Barbieri JT (1999) The N-terminal domain of Pseudomonas aeruginosa exoenzyme S is a GTPase-activating protein for Rho GTPases. J Biol Chem 274:36369–36372
Grunwald IC, Klein R (2002) Axon guidance: receptor complexes and signaling mechanisms. Curr Opin Neurobiol 12:250–259
Hall A (1998) Rho GTPases and the actin cytoskeleton. Science 279:509–514
Han S, Arvai AS, Clancy SB, Tainer JA (2001) Crystal structure and novel recognition motif of Rho ADP-ribosylating C3 exoenzyme from Clostridium botulinum: structural insights for recognition specificity and catalysis. J Mol Biol 305:95–107
Hardt W-D, Chen L-M, Schuebel KE, Bustelo XR, Galán JE (1998) S. typhimurium encodes an activator of Rho GTPases that induces membrane ruffling and nuclear responses in host cells. Cell 93:815–826
Haug G, Barth H, Aktories K (2006) Purification and activity of the Rho ADP-ribosylating binary C2/C3 toxin. Methods Enzymol 406:117–127
Henning SW, Galandrini R, Hall A, Cantrell DA (1997) The GTPase Rho has a critical regulatory role in thymus development. EMBO J 16:2397–2407
Hernandez-Munoz I, Malumbres M, Leonardi P, Pellicer A (2000) The Rgr oncogene (homologous to RalGDS) induces transformation and gene expression by activating Ras, Ral and Rho mediated pathways. Oncogene 19:2745–2757
Herrmann C, Martin GA, Wittinghofer A (1995) Quantitative analysis of the complex between p21ras and the Ras-binding domain of the human Raf-1 protein kinase. J Biol Chem 270:2901–2905
Hess DJ, Henry-Stanley MJ, Erickson EA, Wells CL (2003) Intracellular survival of Staphylococcus aureus within cultured enterocytes. J Surg Res 114:42–49
Hill CS, Wynne J, Treisman R (1995) The Rho family GTPases RhoA, Rac1, and CDC42Hs regulate transcriptional activation by SRF. Cell 81:1159–1170
Hoffman GR, Nassar N, Cerione RA (2000) Structure of the Rho family GTP-binding protein Cdc42 in complex with the multifunctional regulator RhoGDI. Cell 100:345–356
Holbourn KP, Sutton JM, Evans HR, Shone CC, Acharya KR (2005) Molecular recognition of an ADP-ribosylating Clostridium botulinum C3 exoenzyme by RalA GTPase. Proc Natl Acad Sci USA 102:5357–5362
Ihara K, Muraguchi S, Kato M, Shimizu T, Shirakawa M, Kuroda S, Kaibuchi K, Hakoshima T (1998) Crystal structure of human RhoA in a dominantly active form complexed with a GTP analogue. J Biol Chem 273:9656–9666
Inoue S, Sugai M, Murooka Y, Paik S-Y, Hong Y-M, Ohgai H, Suginaka H (1991) Molecular cloning and sequencing of the epidermal cell differentiation inhibitor gene from Staphylococcus aureus. Biochem Biophys Res Commun 174:459–464
Jaffe AB, Hall A (2002) Rho GTPases in transformation and metastasis. In: Jaffe AB, Hall A (eds) Advances in Cancer Research. Academic Press, New York, pp 57–80
Jaffe AB, Hall A (2005) Rho GTPases: biochemistry and biology. Annu Rev Cell Dev Biol 21:247–269
Jorgensen R, Merrill AR, Yates SP, Marquez VE, Schwan AL, Boesen T, Andersen GR (2005) Exotoxin A-eEF2 complex structure indicates ADP ribosylation by ribosome mimicry. Nature 436:979–984
Jullien-Flores V, Dorseuil O, Romero F, Letourneur F, Saragosti S, Berger R, Tavitian A, Gacon G, Camonis JH (1995) Bridging Ral GTPase to Rho pathways—RLIP76, a Ral effector with CDC42/Rac GTPase-activating protein activity. J Biol Chem 270:22473–22477
Just I, Gerhard R (2004) Large clostridial cytotoxins. Rev Physiol Biochem Pharmacol 152:23–47
Just I, Mohr C, Schallehn G, Menard L, Didsbury JR, Vandekerckhove J, van Damme J, Aktories K (1992) Purification and characterization of an ADP-ribosyltransferase produced by Clostridium limosum. J Biol Chem 267:10274–10280
Just I, Selzer J, Jung M, van Damme J, Vandekerckhove J, Aktories K (1995a) Rho-ADP-ribosylating exoenzyme from Bacillus cereus—purification, characterization and identification of the NAD-binding site. Biochemistry 34:334–340
Just I, Selzer J, Wilm M, Von Eichel-Streiber C, Mann M, Aktories K (1995b) Glucosylation of Rho proteins by Clostridium difficile toxin B. Nature 375:500–503
Laudanna C, Campbell JJ, Butcher EC (1996) Role of Rho in chemoattractant-activated leukocyte adhesion through integrins. Science 271:981–983
Lehmann M, Fournier A, Selles-Navarro I, Dergham P, Sebok A, Leclerc N, Tigyi G, McKerracher L (1999) Inactivation of Rho signaling pathway promotes CNS axon regeneration. J Neurosci 19:7537–7547
Liu L, Schwartz BR, Lin N, Winn RK, Harlan JM (2002)- Requirement for RhoA kinase activation in leukocyte de-adhesion. J Immunol 169:2330–2336
Lowy FD (2000) Is Staphylococcus aureus an intracellular pathogen? Trends Microbiol 8:341–344
Madden JC, Ruiz N, Caparon M (2001) Cytolysin-mediated translocation (CMT): a functional equivalent type III secretion in gram-positive bacteria. Cell 104:143–152
Maehama T, Sekine N, Nishina H, Takahashi K, Katada T (1994) Characterization of botulinum C3-catalyzed ADP-ribosylation of rho proteins and identification of mammalian C3-like ADP-ribosyltransferase. Mol Cell Biochem 138:135–140
Masuda M, Betancourt L, Matsuzawa T, Kashimoto T, Takao T, Shimonishi Y, Horiguchi Y (2000) Activation of Rho through a cross-link with polyamines catalyzed by Bordetella dermonecrotizing toxin. EMBO J 19:521–530
Meacci E, Vasta V, Moorman JP, Bobak DA, Bruni P, Moss J, Vaughan M (1999) Effect of Rho and ADP-ribosylation factor GTPases on phospholipase D activity in intact human adenocarcinoma A549 cells. J Biol Chem 274:18605–18612
Meehl MA, Caparon MG (2004) Specificity of streptolysin O in cytolysin-mediated translocation. Mol Microbiol 52:1665–1676
Ménétrey J, Flatau G, Stura EA, Charbonnier J-B, Gas F, Teulon J-M, Le Du M-H, Boquet P, Ménez A (2002) NAD binding induces conformational changes in Rho ADP-ribosylating Clostridium botulinum C3 exoenzyme. J Biol Chem 277:30950–30957
Meyer DK, Olenik C, Hofmann F, Barth H, Leemhuis J, Brünig I, Aktories K, Nörenberg W (2000) Regulation of somatodendritic GABAA receptor channels in rat hippocampal neurons: evidence for a role of the small GTPase Rac1. J Neurosci 20:6743–6751
Millan J, Ridley AJ (2005) Rho GTPases and leucocyte-induced endothelial remodelling. Biochem J 385:329–337
Molinari G, Rohde M, Wilde C, Just I, Aktories K, Chhatwal GS (2006) Localization of the C3-like ADP-ribosyltransferase from Staphylococcus aureus during bacterial invasion of mammalian cells. Infect Immun 3673–3677
Moon SY, Zheng Y (2003) Rho GTPase-activating proteins in cell regulation. Trends Cell Biol 13:13–22
Moss J, Vaughan M (1977) Mechanism of action of choleragen. Evidence for ADP-ribosyltransferase activity with arginine as an acceptor. J Biol Chem 252:2455–2457
Nemoto Y, Namba T, Kozaki S, Narumiya S (1991) Clostridium botulinum C3 ADP-ribosyltransferase gene. J Biol Chem 266:19312–19319
Nicely NI, Kosak J, de Serrano V, Mattos C (2004) Crystal structures of Ral-GppNHp and Ral-GDP reveal two binding sites that are also present in Ras and Rap. Structure (Camb) 12:2025–2036
Nusrat A, Giry M, Turner JR, Colgan SP, Parkos CA, Carnes D, Lemichez E, Boquet P, Madara JL (1995) Rho protein regulates tight junctions and perijunctional actin organization in polarized epithelia. Proc Natl Acad Sci USA 92:10629–10633
Ohishi I, Iwasaki M, Sakaguchi G (1980) Purification and characterization of two components of botulinum C2 toxin. Infect Immun 30:668–673
Ohta Y, Suzuki N, Nakamura S, Hartwig JH, Stossel TP (1999) The small RalA targets filamin to induce filopodia. Proc Natl Acad Sci USA 96:2122–2128
Park J, Kim JS, Jung KC, Lee HJ, Kim JI, Kim J, Lee JY, Park JB, Choi SY (2003) Exoenzyme Tat-C3 inhibits association of zymosan particles, phagocytosis, adhesion, and complement binding in macrophage cells. Mol Cells 16:216–223
Paterson HF, Self AJ, Garrett MD, Just I, Aktories K, Hall A (1990) Microinjection of recombinant p21rho induces rapid changes in cell morphology. J Cell Biol 111:1001–1007
Pautsch A, Vogelsgesang M, Trankle J, Herrmann C, Aktories K (2005) Crystal structure of the C3bot-RalA complex reveals a novel type of action of a bacterial exoenzyme. EMBO J 24:3670–3680
Popoff MR, Boquet P, Gill DM, Eklund MW (1990) DNA sequence of exoenzyme C3, an ADP-ribosyltransferase encoded by Clostridium botulinum C and D phages. Nucleic Acids Res 18:1291
Rak A, Pylypenko O, Durek T, Watzke A, Kushnir S, Brunsveld L, Waldmann H, Goody RS, Alexandrov K (2003) Structure of Rab GDP-dissociation inhibitor in complex with prenylated YPT1 GTPase. Science 302:646–650
Rak A, Pylypenko O, Niculae A, Pyatkov K, Goody RS, Alexandrov K (2004) Structure of the Rab7:REP-1 complex: insights into the mechanism of Rab prenylation and choroideremia disease. Cell 117:749–760
Ridley A (2000) Rho. In: Hall A (ed) GTPases. Oxford University Press, Oxford, pp 89–136
Riento K, Guasch RM, Garg R, Jin B, Ridley AJ (2003) RhoE binds to ROCK I and inhibits downstream signaling. Mol Cell Biol 23:4219–4229
Rossman KL, Der CJ, Sondek J (2005) GEF means go: turning on RHO GTPases with guanine nucleotide-exchange factors. Nat Rev Mol Cell Biol 6:167–180
Rubin EJ, Gill DM, Boquet P, Popoff MR (1988) Functional modification of a 21-Kilodalton G protein when ADP-ribosylated by exoenzyme C3 of Clostridium botulinum. Mol Cell Biol 8:418–426
Sahai E, Olson MF (2006) Purification of TAT-C3 exoenzyme. Methods Enzymol 406:128–140
Sakaguchi Y, Hayashi T, Kurokawa K, Nakayama K, Oshima K, Fujinaga Y, Ohnishi M, Ohtsubo E, Hattori M, Oguma K (2005) The genome sequence of Clostridium botulinum type C neurotoxin-converting phage and the molecular mechanisms of unstable lysogeny. Proc Natl Acad Sci USA 102:17472–17477
Sauzeau V, Le Mellionnec E, Bertoglio J, Scalbert E, Pacaud P, Loirand G (2001) Human urotensin II-induced contraction and arterial smooth muscle cell proliferation are mediated by RhoA and Rho-kinase. Circ Res 88:1102–1104
Schmidt G, Sehr P, Wilm M, Selzer J, Mann M, Aktories K (1997) Gln63 of Rho is deamidated by Escherichia coli cytotoxic necrotizing factor 1. Nature 387:725–729
Sehr P, Joseph G, Genth H, Just I, Pick E, Aktories K (1998) Glucosylation and ADP-ribosylation of Rho proteins—effects on nucleotide binding, GTPase activity, and effector-coupling. Biochemistry 37:5296–5304
Sekine A, Fujiwara M, Narumiya S (1989) Asparagine residue in the rho gene product is the modification site for botulinum ADP-ribosyltransferase. J Biol Chem 264:8602–8605
Shao F, Merritt PM, Bao Z, Innes RW, Dixon JE (2002) A Yersinia effector and a Pseudomonas avirulence protein define a family of cysteine proteases functioning in bacterial pathogenesis. Cell 109:575–588
Shao F, Vacratsis PO, Bao Z, Bowers KE, Fierke CA, Dixon JE (2003) Biochemical characterization of the Yersinia YopT protease: cleavage site and recognition elements in Rho GTPases. Proc Natl Acad Sci USA 100:904–909
Shen Y, Xu l, Foster DA (2001) Role for phospholipase D in receptor-mediated endocytosis. Mol Cell Biol 21:595–602
Stamatovic SM, Keep RF, Kunkel SL, Andjelkovic AV (2003) Potential role of MCP-1 in endothelial cell tight junction ‘opening’: signaling via Rho and Rho kinase. J Cell Sci 116:4615–4628
Urano T, Emkey R, Feig LA (1996) Ral-GTPases mediate a distinct downstream signaling pathway from Ras that facilitates cellular transformation. EMBO J 15:810–816
Valderrama F, Duran JM, Babia T, Barth H, Renau-Piqueras J, Egea G (2001) Actin microfilaments facilitate the retrograde transport from the Golgi complex to the endoplasmic reticulum in mammalian cells. Traffic 2:717–726
Vandekerckhove J, Schering B, Bärmann M, Aktories K (1988) Botulinum C2 toxin ADP-ribosylates cytoplasmic β/g-actin in arginine 177. J Biol Chem 263:696–700
Vetter IR, Wittinghofer A (2001) The guanine nucleotide-binding switch in three dimensions. Science 294:1299–1304
Vischer UM, Barth H, Wollheim CB (2000) Regulated von Willebrand factor secretion is associated with agonist-specific patterns of cytoskeletal remodeling in cultured endothelial cells. Arterioscler Thromb Vasc Biol 20:883–891
Vogelsgesang M, Aktories K (2006) Exchange of glutamine-217 to glutamate of Clostridium limosum exoenzyme C3 turns the asparagine-specific ADP-ribosyltransferase into an arginine-modifying enzyme. Biochemistry 45:1017–1025
Von Pawel-Rammingen U, Telepnev MV, Schmidt G, Aktories K, Wolf-Watz H, Rosqvist R (2000) GAP activity of the Yersinia YopE cytotoxin specifically targets the Rho pathway: a mechanism for disruption of actin microfilament structure. Mol Microbiol 36:737–748
Voth DE, Ballard JD (2005) Clostridium difficile toxins: mechanism of action and role in disease. Clin Microbiol Rev 18:247–263
Wahl S, Barth H, Ciossek T, Aktories K, Mueller BK (2000) Ephrin-A5 induces collapse of growth cones by activating Rho and Rho kinase. J Cell Biol 149:263–270
Watanabe N, Madaule P, Reid T, Ishizaki T, Watanabe G, Kakizuka A, Saito Y, Nakao K, Jockusch BM, Naumiya S (1997) p140mDia, a mammalian homolog of Drosophila diaphanous, is a target protein fro Rho small GTPase and is a ligand for profilin. EMBO J 16:3044–3056
Wennerberg K, Der CJ (2004) Rho-family GTPases: it’s not only Rac and Rho (and I like it). J Cell Sci 117:1301–1312
Wennerberg K, Forget MA, Ellerbroek SM, Arthur WT, Burridge K, Settleman J, Der CJ, Hansen SH (2003) Rnd proteins function as RhoA antagonists by activating p190 RhoGAP. Curr Biol 13:1106–1115
West RE, Moss J, Vaughan M, Liu T, Liu T-Y (1985) Pertussis toxin-catalyzed ADP-ribosylation of transducin. J Biol Chem 260:14428–14430
Wiegers W, Just I, Müller H, Hellwig A, Traub P, Aktories K (1991) Alteration of the cytoskeleton of mammalian cells cultured in vitro by Clostridium botulinum C2 toxin and C3 ADP-ribosyltransferase. Eur J Cell Biol 54:237–245
Wilde C, Genth H, Aktories K, Just I (2000) Recognition of RhoA by Clostridium botulinum C3 exoenzyme. J Biol Chem 275:16478–16483
Wilde C, Chhatwal GS, Schmalzing G, Aktories K, Just I (2001) A novel C3-like ADP-ribosyltransferase from Staphylococcus aureus modifying RhoE and Rnd3. J Biol Chem 276:9537–9542
Wilde C, Barth H, Sehr P, Han L, Schmidt M, Just I, Aktories K (2002a) Interaction of the Rho-ADP-ribosylating C3 exoenzyme with RalA. J Biol Chem 277:14771–14776
Wilde C, Just I, Aktories K (2002b) Structure-function analysis of the Rho-ADP-ribosylating exoenzyme C3stau2 from Staphylococcus aureus. Biochemistry 41:1539–1544
Wilde C, Vogelsgesang M, Aktories K (2003) Rho-specific Bacillus cereus ADP-ribosyltransferase C3cer cloning and characterization. Biochemistry 42:9694–9702
Winton MJ, Dubreuil CI, Lasko D, Leclerc N, McKerracher L (2002) Characterization of new cell permeable C3-like proteins that inactivate Rho and stimulate neurite outgrowth on inhibitory substrates. J Biol Chem 277:32820–32829
Worthylake RA, Lemoine S, Watson JM, Burridge K (2001) RhoA is required for monocyte tail retraction during transendothelial migration. J Cell Biol 154:147–160
Yamaguchi T, Hayashi T, Takami H, Ohnishi M, Murata T, Nakayama K, Asakawa K, Ohara M, Komatsuzawa H, Sugai M (2001) Complete nucleotide sequence of a Staphylococcus aureus exfoliative toxin B plasmid and identification of a novel ADP-ribsyltransferase, EDIN-C. Infect Immun 69:7760–7771
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is dedicated to Prof. Dr. Karl-Heinz Jakobs. Looking back to the 1980s, discussions with Karl-Heinz Jakobs during the discovery of C3 in the laboratory of K.A. were extremely helpful in establishing C3 as an important pharmacological tool.
Rights and permissions
About this article
Cite this article
Vogelsgesang, M., Pautsch, A. & Aktories, K. C3 exoenzymes, novel insights into structure and action of Rho-ADP-ribosylating toxins. Naunyn-Schmied Arch Pharmacol 374, 347–360 (2007). https://doi.org/10.1007/s00210-006-0113-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00210-006-0113-y