Abstract
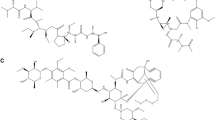
In the present study, we investigated the activity and modes of action of cajanin stilbene acid (CSA) and its derivatives in terms of cytotoxicity, gene expression profile, and transcription factor activity. XTT assays on MCF7 cells were performed upon treatment with CSA or derivatives. After the determination of IC50 values, gene expression profiling was performed with Agilent microarray experiments. Deregulated genes were determined with Chipster software, pathway and functional analyses were performed with Ingenuity pathway software. In order to identify the potential upstream regulators, MatInspector software was used to perform transcription factor binding motif search in the promoter regions of the deregulated genes. Molecular docking on MYC/MAX complex and reporter cell line experiments were performed to validate the MYC inhibitory activity of CSA and its derivatives. Two known MYC inhibitors: 10058-F4 and 10074-G5 were used as positive control. All compounds showed cytotoxicities in the micromolar range. Microarray analyses pointed to cell cycle, DNA damage, and DNA repair as mainly affected cellular functions. Promoter motif analysis of the deregulated genes further supported the microarray gene expression analysis results emphasizing the relevance of transcription factors regulating cell cycle and proliferation, with MYC as being the most pronounced one. Luciferase-based reporter cell line experiments and molecular docking studies yielded supportive results emphasizing the inhibitory activity of CSA and its derivatives on MYC. CSA and its derivatives are shown to be promising anticancer compounds with low toxicity. They inhibit MYC activity comparable to 10058-F4 and 10074-G5. Further studies are warranted to analyze the therapeutic applicability of these compounds in more detail.





Similar content being viewed by others
References
Adams JM, Harris AW, Pinkert CA et al (1985) The c-MYC oncogene driven by immunoglobulin enhancers induces lymphoid malignancy in transgenic mice. Nature 318(6046):533–538
Addou-Klouche L, Adelaide J, Cornen S et al (2012) Integrated genomic analysis of breast cancers. Balkan J Med Genet BJMG 15(Suppl):71–74. doi:10.2478/v10034-012-0023-x
Ali S, Coombes RC (2002) Endocrine-responsive breast cancer and strategies for combating resistance. Nat Rev Cancer 2(2):101–112. doi:10.1038/nrc721
Amundadottir LT, Johnson MD, Merlino G, Smith GH, Dickson RB (1995) Synergistic interaction of transforming growth factor alpha and c-MYC in mouse mammary and salivary gland tumorigenesis. Cell Growth Differ Mol Biol J Am Assoc Cancer Res 6(6):737–748
Arnold I, Watt FM (2001) c-MYC activation in transgenic mouse epidermis results in mobilization of stem cells and differentiation of their progeny. Curr Biol CB 11(8):558–568
Bhardwaj A, Rosen D, Liu M et al (2014) suppression of Akt-mTOR pathway—a novel component of oncogene induced DNA damage response barrier in breast tumorigenesis. PLoS ONE 9(5):e97076. doi:10.1371/journal.pone.0097076
Blackwell TK, Kretzner L, Blackwood EM, Eisenman RN, Weintraub H (1990) Sequence-specific DNA binding by the c-MYC protein. Science 250(4984):1149–1151
Butt AJ, McNeil CM, Musgrove EA, Sutherland RL (2005) Downstream targets of growth factor and oestrogen signalling and endocrine resistance: the potential roles of c-MYC, cyclin D1 and cyclin E. Endocr Relat Cancer 12(Suppl 1):S47–S59. doi:10.1677/erc.1.00993
Byler S, Goldgar S, Heerboth S et al (2014) Genetic and epigenetic aspects of breast cancer progression and therapy. Anticancer Res 34(3):1071–1077
Cartharius K, Frech K, Grote K et al (2005) MatInspector and beyond: promoter analysis based on transcription factor binding sites. Bioinformatics 21(13):2933–2942. doi:10.1093/bioinformatics/bti473
Castro F, Dirks WG, Fahnrich S, Hotz-Wagenblatt A, Pawlita M, Schmitt M (2013) High-throughput SNP-based authentication of human cell lines. Int J Cancer 132(2):308–314. doi:10.1002/ijc.27675
Chen Y, Olopade OI (2008) MYC in breast tumor progression. Expert Rev Anticancer Ther 8(10):1689–1698. doi:10.1586/14737140.8.10.1689
Cheng AS, Jin VX, Fan M et al (2006) Combinatorial analysis of transcription factor partners reveals recruitment of c-MYC to estrogen receptor-alpha responsive promoters. Mol Cell 21(3):393–404. doi:10.1016/j.molcel.2005.12.016
Clarke R, Liu MC, Bouker KB et al (2003) Antiestrogen resistance in breast cancer and the role of estrogen receptor signaling. Oncogene 22(47):7316–7339. doi:10.1038/sj.onc.1206937
Clausen DM, Guo J, Parise RA et al (2010) In vitro cytotoxicity and in vivo efficacy, pharmacokinetics, and metabolism of 10074-G5, a novel small-molecule inhibitor of c-MYC/MAX dimerization. J Pharmaco Exp Ther 335(3):715–727. doi:10.1124/jpet.110.170555
D’Cruz CM, Gunther EJ, Boxer RB et al (2001) c-MYC induces mammary tumorigenesis by means of a preferred pathway involving spontaneous Kras2 mutations. Nat Med 7(2):235–239. doi:10.1038/84691
Eichhorn T (2011) Molecular biology, pharmacogenomics and bioinformatics of natural compounds and synthetic compounds for cancer therapy. Ph.D. Thesis. Ruperto-Carola University of Heidelberg, Germany
Escot C, Theillet C, Lidereau R et al (1986) Genetic alteration of the c-MYC protooncogene (MYC) in human primary breast carcinomas. Proc Natl Acad Sci USA 83(13):4834–4838
Felsher DW, Bishop JM (1999) Reversible tumorigenesis by MYC in hematopoietic lineages. Mol Cell 4(2):199–207
Follis AV, Hammoudeh DI, Wang H, Prochownik EV, Metallo SJ (2008) Structural rationale for the coupled binding and unfolding of the c-MYC oncoprotein by small molecules. Chem Biol 15(11):1149–1155. doi:10.1016/j.chembiol.2008.09.011
Guerin M, Barrois M, Terrier MJ, Spielmann M, Riou G (1988) Overexpression of either c-MYC or c-erbB-2/neu proto-oncogenes in human breast carcinomas: correlation with poor prognosis. Oncog Res 3(1):21–31
Harrington EA, Bennett MR, Fanidi A, Evan GI (1994) c-MYC-induced apoptosis in fibroblasts is inhibited by specific cytokines. EMBO J 13(14):3286–3295
Jonkers J, Berns A (2004) Oncogene addiction: sometimes a temporary slavery. Cancer Cell 6(6):535–538. doi:10.1016/j.ccr.2004.12.002
Kallio MA, Tuimala JT, Hupponen T et al (2011) Chipster: user-friendly analysis software for microarray and other high-throughput data. BMC Genom 12:507. doi:10.1186/1471-2164-12-507
Kim J, Lee JH, Iyer VR (2008) Global identification of Myc target genes reveals its direct role in mitochondrial biogenesis and its E-box usage in vivo. PLoS ONE 3(3):e1798. doi:10.1371/journal.pone.0001798
Kim B, Fatayer H, Hanby AM et al (2013) Neoadjuvant chemotherapy induces expression levels of breast cancer resistance protein that predict disease-free survival in breast cancer. PLoS ONE 8(5):e62766. doi:10.1371/journal.pone.0062766
Kirkland JL, Murthy L, Stancel GM (1992) Progesterone inhibits the estrogen-induced expression of c-fos messenger ribonucleic acid in the uterus. Endocrinology 130(6):3223–3230. doi:10.1210/endo.130.6.1375896
Musgrove EA, Sergio CM, Loi S et al (2008) Identification of functional networks of estrogen- and c-MYC-responsive genes and their relationship to response to tamoxifen therapy in breast cancer. PLoS ONE 3(8):e2987. doi:10.1371/journal.pone.0002987
Nair SK, Burley SK (2003) X-ray structures of Myc-Max and Mad-Max recognizing DNA. Molecular bases of regulation by proto-oncogenic transcription factors. Cell 112(2):193–205
Nicholson RI, Hutcheson IR, Britton D et al (2005) Growth factor signalling networks in breast cancer and resistance to endocrine agents: new therapeutic strategies. J Steroid Biochem Mol Biol 93(2–5):257–262. doi:10.1016/j.jsbmb.2004.12.006
Parris TZ, Kovacs A, Hajizadeh S et al (2014) Frequent MYC coamplification and DNA hypomethylation of multiple genes on 8q in 8p11–p12-amplified breast carcinomas. Oncogenesis 3:e95. doi:10.1038/oncsis.2014.8
Pavesi G, Pesole G (2006) Using Weeder for the discovery of conserved transcription factor binding sites. Current protocols in bioinformatics/editoral board, Andreas D Baxevanis et al. Chapter 2: Unit 2 11. doi:10.1002/0471250953.bi0211s15
Pelengaris S, Khan M, Evan G (2002a) c-MYC: more than just a matter of life and death. Nat Rev Cancer 2(10):764–776. doi:10.1038/nrc904
Pelengaris S, Khan M, Evan GI (2002b) Suppression of Myc-induced apoptosis in beta cells exposes multiple oncogenic properties of Myc and triggers carcinogenic progression. Cell 109(3):321–334
Plourde KV, Labrie Y, Desjardins S et al (2013) Analysis of ZNF350/ZBRK1 promoter variants and breast cancer susceptibility in non-BRCA1/2 French Canadian breast cancer families. J Hum Genet 58(2):59–66. doi:10.1038/jhg.2012.127
Ponzielli R, Katz S, Barsyte-Lovejoy D, Penn LZ (2005) Cancer therapeutics: targeting the dark side of Myc. Eur J Cancer 41(16):2485–2501. doi:10.1016/j.ejca.2005.08.017
Prendergast GC, Ziff EB (1992) A new bind for Myc. Trends Genet TIG 8(3):91–96
Schuldiner O, Benvenisty N (2001) A DNA microarray screen for genes involved in c-MYC and N-MYC oncogenesis in human tumors. Oncogene 20(36):4984–4994. doi:10.1038/sj.onc.1204459
Shachaf CM, Kopelman AM, Arvanitis C et al (2004) MYC inactivation uncovers pluripotent differentiation and tumour dormancy in hepatocellular cancer. Nature 431(7012):1112–1117. doi:10.1038/nature03043
Shang Y, Baumrucker CR, Green MH (1998) c-MYC is a major mediator of the synergistic growth inhibitory effects of retinoic acid and interferon in breast cancer cells. J Biol Chem 273(46):30608–30613
Sousa SF, Fernandes PA, Ramos MJ (2006) Protein-ligand docking: current status and future challenges. Proteins 65(1):15–26. doi:10.1002/prot.21082
Venditti M, Iwasiow B, Orr FW, Shiu RP (2002) c-MYC gene expression alone is sufficient to confer resistance to antiestrogen in human breast cancer cells. Int J Cancer 99(1):35–42
Vennstrom B, Sheiness D, Zabielski J, Bishop JM (1982) Isolation and characterization of c-MYC, a cellular homolog of the oncogene (v-MYC) of avian myelocytomatosis virus strain 29. J Virol 42(3):773–779
Wang YH, Liu S, Zhang G et al (2005) Knockdown of c-MYC expression by RNAi inhibits MCF-7 breast tumor cells growth in vitro and in vivo. Breast Cancer Res BCR 7(2):R220–R228. doi:10.1186/bcr975
Wang H, Hammoudeh DI, Follis AV et al (2007) Improved low molecular weight Myc-Max inhibitors. Mol Cancer Ther 6(9):2399–2408. doi:10.1158/1535-7163.MCT-07-0005
Wu N, Kong Y, Fu Y et al (2011) In vitro antioxidant properties, DNA damage protective activity, and xanthine oxidase inhibitory effect of cajaninstilbene acid, a stilbene compound derived from pigeon pea [Cajanus cajan (L.) Millsp.] leaves. J Agric Food Chem 59(1):437–443. doi:10.1021/jf103970b
Yan Y, Zhang GX, Williams MS et al (2012) TCR stimulation upregulates MS4a4B expression through induction of AP-1 transcription factor during T cell activation. Mol Immunol 52(2):71–78. doi:10.1016/j.molimm.2012.04.011
Yap JL, Wang H, Hu A et al (2013) Pharmacophore identification of c-MYC inhibitor 10074-G5. Bioorg Med Chem Lett 23(1):370–374. doi:10.1016/j.bmcl.2012.10.013
Ziv-Av A, Taller D, Attia M et al (2011) RTVP-1 expression is regulated by SRF downstream of protein kinase C and contributes to the effect of SRF on glioma cell migration. Cell Signal 23(12):1936–1943. doi:10.1016/j.cellsig.2011.07.001
Zu YG, Liu XL, Fu YJ, Wu N, Kong Y, Wink M (2010) Chemical composition of the SFE-CO extracts from Cajanus cajan (L.) Huth and their antimicrobial activity in vitro and in vivo. Phytomedicine Int J Phytotherapy Phytopharm 17(14):1095–1101. doi:10.1016/j.phymed.2010.04.005
Conflict of interest
We declare that there is no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Onat Kadioglu and Yujie Fu have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kadioglu, O., Fu, Y., Wiench, B. et al. Synthetic cajanin stilbene acid derivatives inhibit c-MYC in breast cancer cells. Arch Toxicol 90, 575–588 (2016). https://doi.org/10.1007/s00204-015-1480-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-015-1480-2