Abstract
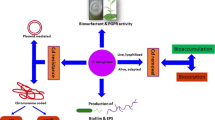
Biofilm-forming mercury-resistant marine bacterium Bacillus cereus BW-201B has been explored to evident that the bacterial biofilm-EPS (exopolymers) trap inorganic mercury but subsequently release EPS-bound mercury for induction of mer operon-mediated volatilization of inorganic mercury. The isolate was able to tolerate 50 ppm of mercury and forms biofilm in presence of mercury. mer operon-mediated volatilization was confirmed, and –SH was found to be the key functional group of bacterial EPS responsible for mercury binding. Biofilm-EPS-bound mercury was found to be internalized to the bacterial system as confirmed by reversible conformational change of –SH group and increased expression level of merA gene in a timescale experiment. Biofilm-EPS trapped Hg after 24 h of incubation, and by 96 h, the volatilization process reaches to its optimum confirming the internalization of EPS-bound mercury to the bacterial cells. Biofilm disintegration at the same time corroborates the results.






Similar content being viewed by others
References
Abdelatey LM, Khalil WK, Ali TH, Mahrous KF (2011) Heavy metal resistance and gene expression analysis of metal resistance genes in Gram-positive and Gram-negative bacteria present in Egyptian soils. J Appl Sci Environ Sanit 6:201–211
Allison DG (2003) The biofilm matrix. Biofouling 19:139–150
Asahara A, Nakajima M, Fukaya R, Tokoro H, Ohkoshi SI, Suemoto T (2012) Ultrafast dynamics of reversible photoinduced phase transitions in rubidium manganese hexacyanoferrate investigated by midinfrared CN vibration spectroscopy. Phys Rev B 86:195138
Barkay T, Miller SM, Summers AO (2003) Bacterial mercury resistance from atoms to ecosystems. FEMS Microbiol Rev 27:355–384
Boyer PD (1959) Sulfhydryl and disulfide groups of enzymes. Enzymes 1:511–588
Bura R, Cheung M, Liao B, Finlayson J, Lee BC, Droppo IG, Liss SN (1998) Composition of extracellular polymeric substances in the activated sludge floc matrix. Water Sci Technol 37:325–333
Cappuccino J, Sherman N (2013) Microbiology: a laboratory manual. Pearson/Benjamin Cummings, Dorling Kindersley, New Delhi
Chakraborty J, Das S (2014) Characterization and cadmium-resistant gene expression of biofilm-forming marine bacterium Pseudomonas aeruginosa JP-11. Environ Sci Pollut Res 21:14188–14201
Chakraborty J, Das S (2016) Characterization of the metabolic pathway and catabolic gene expression in biphenyl degrading marine bacterium Pseudomonas aeruginosa JP-11. Chemosphere 144:1706–1714
Clinical and Laboratory Standards (2006) Institute methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically, Approved Standard M7-A7, 7th edn. CLSI, Wayne
Dash HR, Das S (2012) Bioremediation of mercury and the importance of bacterial mer genes. Int Biodeterior Biodegrad 75:207–213
Dash HR, Das S (2014) Assessment of mercury pollution through mercury resistant marine bacteria in Bhitarkanika mangrove ecosystem, Odisha, India. Indian J Geo-Mar Sci 43:1103–1115
Dash HR, Mangwani N, Chakraborty J, Kumari S, Das S (2013) Marine bacteria: potential candidates for enhanced bioremediation. Appl Microbiol Biotechnol 97:561–571
Dash HR, Mangwani N, Das S (2014) Characterization and potential application in mercury bioremediation of highly mercury-resistant marine bacterium Bacillus thuringiensis PW-05. Environ Sci Pollut Res 21:2642–2653
Davies D (2003) Understanding biofilm resistance to antibacterial agents. Nat Rev Drug Discov 2:114–122
De Souza MJ, Nair S, Bharathi PL, Chandramohan D (2006) Metal and antibiotic-resistance in psychrotrophic bacteria from Antarctic Marine waters. Ecotoxicology 15:379–384
Deng X, Wang P (2012) Isolation of marine bacteria highly resistant to mercury and their bioaccumulation process. Bioresour Technol 121:342–347
Fang HHP, Xu LC, Chan KY (2002) Effects of toxic metals and chemicals on biofilm and biocorrosion. Water Res 36:4709–4716
Faust B (1997) Modern chemical techniques: An essential reference for students and teachers. Royal Society of Chemistry. ISBN 9780854042869
Flemming HC, Neu TR, Wozniak DJ (2007) The EPS matrix: the “house of biofilm cells”. J Bacteriol 189:7945–7947
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evol 39:783–791
Hancock V, Dahl M, Klemm P (2010) Abolition of biofilm formation in urinary tract Escherichia coli and Klebsiella isolates by metal interference through competition for Fur. Appl Environ Microbiol 76:3836–3841
Harrison JJ, Ceri H, Yerly J, Stremick CA, Hu Y, Martinuzzi R, Turner RJ (2006) The use of microscopy and three-dimensional visualization to evaluate the structure of microbial biofilms cultivated in the Calgary Biofilm Device. Biol Proced Online 8:194–215
Harrison JJ, Ceri H, Turner RJ (2007) Multimetal resistance and tolerance in microbial biofilms. Nat Rev Microbiol 5:928–938
Heydorn A, Nielsen AT, Hentzer M, Sternberg C, Givskov M, Ersbøll BK, Molin S (2000) Quantification of biofilm structures by the novel computer program COMSTAT. Microbiology 146:2395–2407
Høiby N, Bjarnsholt T, Givskov M, Molin S, Ciofu O (2010) Antibiotic resistance of bacterial biofilms. Int J Antimicrob Agents 35:322–332
Hong Z, Chen W, Rong X, Cai P, Dai K, Huang Q (2013) The effect of extracellular polymeric substances on the adhesion of bacteria to clay minerals and goethite. Chem Geol 360:118–125
Jain K, Parida S, Mangwani N, Dash HR, Das S (2013) Isolation and characterization of biofilm-forming bacteria and associated extracellular polymeric substances from oral cavity. Ann Microbiol 63:1553–1562
Jeffrey WH, Nazaret S, Barkay T (1996) Detection of the merA gene and its expression in the environment. Microb Ecol 32:293–303
Kristian SA, Birkenstock TA, Sauder U, Mack D, Götz F, Landmann R (2008) Biofilm formation induces C3a release and protects Staphylococcus epidermidis from IgG and complement deposition and from neutrophil-dependent killing. J Infect Dis 197:1028–1035
Leid JG, Willson CJ, Shirtliff ME, Hassett DJ, Parsek MR, Jeffers AK (2005) The exopolysaccharide alginate protects Pseudomonas aeruginosa biofilm bacteria from IFN-γ-mediated macrophage killing. J Immunol 175:7512–7518
Leonhäuser J, Wang W, Deckwer WD, Wagner-Döbler I (2007) Functioning of the mercury resistance operon at extremely high Hg(II) loads in a chemostat: a proteome analysis. J Biotechnol 132:469–480
Lisovskyy IP, Litovchenko VG, Mazunov DO, Kaschieva S, Koprinarova J, Dmitriev SN (2005) Infrared spectroscopy study of Si-SiO2 structures irradiated with high-energy electrons. J Optoelectron Adv Mater 7:325–328
Ma H, Bryers JD (2010) Non-invasive method to quantify local bacterial concentrations in a mixed culture biofilm. J Ind Microbiol Biotechnol 37:1081–1089
Mah TFC, O’Toole GA (2001) Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol 9:34–39
Mangwani N, Dash HR, Chauhan A, Das S (2012) Bacterial quorum sensing: functional features and potential applications in biotechnology. J Mol Microbiol Biotechnol 22:215–227
Nakamura K, Nakahara H (1988) Simplified X-ray film method for detection of bacterial volatilization of mercury chloride by Escherichia coli. Appl Environ Microbiol 54:2871–2873
Nascimento AM, Chartone-Souza E (2003) Operon mer: bacterial resistance to mercury and potential for bioremediation of contaminated environments. Genet Mol Res 2:92–101
Neurath H, Hill RL (1977) The proteins. Academic Press, New York
Nishikawa Y, Nakano T, Noda I (2012) Detection of reversible nonlinear dynamic responses of polymer films by using time-resolved soft-pulse compression attenuated total reflection step-scan fourier transform infrared spectroscopy. Appl Spectrosc 66:312–318
Oregaard G, Sorenson SJ (2007) High diversity of bacterial mercuric reductase genes from surface and sub-surface floodplain soil (Oak Ridge, USA). ISME J 1:453–467
O’Toole GA (2003) To build a biofilm. J Bacteriol 185:2687–2689
O’Toole G, Kaplan HB, Kolter R (2000) Biofilm formation as microbial development. Ann Rev Microbiol 54:49–79
Pavia D, Lampman G, Kriz G, Vyvyan J (2008) Introduction to spectroscopy. In: Cengage learning, pp 784
Pepi M, Gaggi C, Bernardini E, Focardi S, Lobianco A, Ruta M, Nicolardi V, Volterrani M, Gasperini S, Trinchera G, Renzi P, Gabellini M, Focardi SE (2011) Mercury-resistant bacterial strains Pseudomonas and Psychrobacter spp. isolated from sediments of Orbetello Lagoon (Italy) and their possible use in bioremediation processes. Int Biodeterior Biodegrad 65:85–91
Pepi M, Focardi S, Tarabelli A, Volterrani M, Focardi SE (2013) Bacterial strains resistant to inorganic and organic forms of mercury isolated from polluted sediments of the Orbetello Lagoon, Italy, and their possible use in bioremediation processes. E3S Web of Conferences. doi:10.1051/e3sconf/20130131002
Prakash B, Veeregowda BM, Krishnappa G (2003) Biofilms: a survival strategy of bacteria. Curr Sci 85:1299–1307
Ramaiah N, De J (2003) Unusual rise in mercury-resistant bacteria in coastal environs. Microb Ecol 45:444–454
Rapposch S, Zangerl P, Ginzinger W (2000) Influence of fluorescence of bacteria stained with acridine orange on the enumeration of microorganisms in raw milk. J Diary Sci 83:2753–2758
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Santos-Gandelman JF, Cruz K, Crane S, Muricy G, Giambiagi-deMarval M, Barkay T, Laport MS (2014) Potential application in mercury bioremediation of a marine sponge-isolated Bacillus cereus strain Pj1. Curr Microbiol 69:374–380
Schroeder WH, Munthe J (1998) Atmospheric mercury—an overview. Atmos Environ 32:809–822
Seigneur C, Vijayaraghavan K, Lohman K, Karamchandani P, Scott C (2004) Global source attribution for mercury deposition in the United States. Environ Sci Technol 38:555–569
Settle AF (1997) Handbook of instrumental techniques for analytical chemistry. Prentice Hall Inc., New Jersey
Sinha A, Pant KK, Khare SK (2012) Studies on mercury bioremediation by alginate immobilized mercury tolerant Bacillus cereus cells. Int Biodeterior Biodegrad 71:1–8
Sotero-Martins A, Jesus MSD, Lacerda M, Moreira JC, Filgueiras ALL, Barrocas PRG (2008) A conservative region of the mercuric reductase gene (merA) as a molecular marker of bacterial mercury resistance. Braz J Microbiol 39:307–310
Stewart PS (2002) Mechanisms of antibiotic resistance in bacterial biofilms. Int J Med Microbiol 292:107–113
Stewart PS, William Costerton J (2001) Antibiotic resistance of bacteria in biofilms. Lancet 358:135–138
Teitzel GM, Parsek MR (2003) Heavy metal resistance of biofilm and planktonic Pseudomonas aeruginosa. Appl Environ Microbiol 69:2313–2320
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) CLUSTAL X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Vuong C, Voyich JM, Fischer ER, Braughton KR, Whitney AR, DeLeo FR, Otto M (2004) Polysaccharide intercellular adhesin (PIA) protects Staphylococcus epidermidis against major components of the human innate immune system. Cell Microbiol 6:269–275
Wang X, Wood TK (2011) Toxin-antitoxin systems influence biofilm and persister cell formation and the general stress response. Appl Environ Microbiol 77:5577–5583
Wei X, Fang L, Cai P, Huang Q, Chen H, Liang W, Rong X (2011) Influence of extracellular polymeric substances (EPS) on Cd adsorption by bacteria. Environ Pollut 159:1369–1374
Whelan DR, Hiscox TJ, Rood JI, Bambery KR, McNaughton D, Wood BR (2014) Detection of an en masse and reversible B-to A-DNA conformational transition in prokaryotes in response to desiccation. J R Soc Interface. doi:10.1098/rsif.2014.0454
Zulaika E, Sembiring L (2014) Indigenous mercury resistant bacterial isolates belong to the genus Bacillus from Kalimas Surabaya as a potential mercury bioreducer. J Appl Environ Biol Sci 4:72–76
Acknowledgements
We would like to acknowledge the authorities of NIT, Rourkela, for providing facilities. S.D. thanks the Department of Biotechnology, Ministry of Science and Technology, Government of India for a research Grant (No. BT/PR7480/BCE/8/945/2012).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Shuang-Jiang Liu.
Hirak R. Dash and Subham Basu have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dash, H.R., Basu, S. & Das, S. Evidence of mercury trapping in biofilm-EPS and mer operon-based volatilization of inorganic mercury in a marine bacterium Bacillus cereus BW-201B. Arch Microbiol 199, 445–455 (2017). https://doi.org/10.1007/s00203-016-1317-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-016-1317-2