Abstract
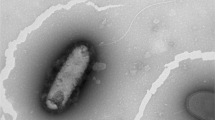
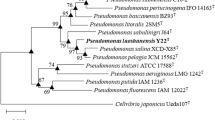
A Gram-staining-negative, rod-shaped and motile with several polar flagellums bacterium, designated WM-3T, was isolated from a rice paddy soil in South China. Growth occurred with 0–3.0 % (w/v) NaCl (optimum 2.0 %), at pH 5.5–9.0 (optimum pH 7.0) and at 25–42 °C (optimum 30–37 °C) in liquid Reasoner’s 2A medium. Analysis of the 16S rRNA gene and gyrB gene sequences revealed that strain WM-3T was most closely related to the type strains of the species Pseudomonas linyingensis and Pseudomonas sagittaria. Its sequence similarities with P. linyingensis CGMCC 1.10701T and P. sagittaria JCM 18195T were 97.4 and 97.3 %, respectively, for 16S rRNA gene, and were 94.1 and 94.2 %, respectively, for gyrB gene. DNA–DNA hybridization between strain WM-3T and these two type strains showed relatedness of 35.6 and 30.9 %, respectively. G+C content of genomic DNA was 69.4 mol%. The whole-cell fatty acids mainly consisted of C16:0 (30.0 %), C16:1 ω6c and/or C16:1 ω7c (19.3 %) and C18:1 ω6c and/or C18:1 ω7c (16.3 %). The results of phenotypic, chemotaxonomic and genotypic analyses clearly indicated that strain WM-3T belongs to genus Pseudomonas but represents a novel species, for which the name Pseudomonas oryzae sp. nov. is proposed. The type strain is WM-3T (=KCTC 32247T =CGMCC 1.12417T).


Similar content being viewed by others
References
Baker GC, Smith JJ, Cowan DA (2003) Review and reanalysis of domain-specific 16S primers. J Microbiol Methods 55:541–555
Chaudhary HJ, Peng G, Hu M, He Y, Yang L, Luo Y, Tan Z (2012) Genetic diversity of endophytic diazotrophs of the wild rice, Oryza alta and identification of the new diazotroph, Acinetobacter oryzae sp. nov. Microb Ecol 63:813–821
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Dong X, Cai M (2001) Determination of biochemical properties. Manual for the systematic identification of General Bacteria. Science Press, Beijing, pp 364–398
Ezaki T, Hashimoto Y, Yabuuchi E (1989) Fluorometric deoxyribonucleic acid deoxyribonucleic acid hybridization in microdilution wells as an alternative to membrane-filter hybridization in which radioisotopes are used to determine genetic relatedness among bacterial strains. Int J Syst Bacteriol 39:224–229
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
He WH, Wang YN, Du X, Zhou Y, Jia B, Bian J, Liu SJ, Chen GC (2012) Pseudomonas linyingensis sp. nov.: a novel bacterium isolated from wheat soil subjected to long-term herbicides application. Curr Microbiol 65:595–600
Jeon Y, Lee SS, Chung BS, Kim JM, Bar JW, Park SK, Jeon CO (2009) Mucilaginibacter oryzae sp. nov., isolated from soil of a rice paddy. Int J Syst Evol Microbiol 59:1451–1454
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeom YS, Lee JH, Yi H, Won S, Chun H (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
King EO, Ward MK, Raney DE (1954) Two simple media for the demonstration of pyocyanin and fluorescin. J Lab Clin Med 44:301–307
Kumar BV, Ramaprasad EVV, Sasikala C, Ramana CV (2013) Rhodopseudomonas pentothenatexigens sp. nov. and Rhodopseudomonas thermotolerans sp. nov., isolated from paddy soils. Int J Syst Evol Microbiol 63:200–207
Lin SY, Hameed A, Liu YC, Hsu YH, Lai WA, Chen WM, Shen FT, Young CC (2012) Pseudomonas sagittaria sp. nov., a novel siderophore-producing bacterium isolated from oil-contaminated soil. Int J Syst Evol Microbiol doi. doi:10.1099//ijs.0.045567-0
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Migula W (1894) Über ein neues System der Bakterien. Arb Bakteriol Inst Karlsruhe 1:235–238
Murray RGE, Doetsch RN, Robinow F (1994) Determinative and cytological light microscopy. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 21–41
Palleroni NJ (2005) Genus I. Pseudomonas migula, 1894. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s Manual of Systematic Bacteriology, 2nd edn. Springer, New York, pp 323–379
Pettersson B, Lembke F, Hammer P, Stackebrandt E, Priest FG (1996) Bacillus sporothermodurans, a new species producing highly heat-resistant endospores. Int J Syst Bacteriol 46:759–764
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Sohn JH, Kwon KY, Kang J-H, Jung HB, Kim S-J (2004) Novosphingobium pentaromativorans sp. nov., a high-molecular-mass polycyclic aromatic hydrocarbon-degrading bacterium isolated from estuarine sediment. Int J Syst Evol Microbiol 54:1483–1487
Tamaoka J, Katayama-Fujimura Y, Kuraishi H (1983) Analysis of bacterial menaquinone mixtures by high performance liquid chromatography. J Appl Bacteriol 54:31–36
Tchan YT, New PB (1984) Genus I. Azotobacter beijerinck. In: Krieg NR, Holt JG (eds) Bergey’s manual of systematic bacteriology. The Williams & Wilkins Co., Baltimore, pp 220–229
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL-X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tindall BJ, Rossello-Mora R, Busse H-J, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266
Wayne LG, Brenner DJ, Colwell RR, Grimont AD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murry RGE, Stackebrandt E, Starr MP (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Xie CH, Yokota A (2006) Zoogloea oryzae sp. nov., a nitrogen-fixing bacterium isolated from rice paddy soil, and reclassification of the strain ATCC 19623 as Crabtreella saccharophila gen. nov., sp. nov. Int J Syst Evol Microbiol 56:619–624
Xin YH, Zhang DC, Liu HC, Zhou HL, Zhou YG (2009) Pseudomonas tuomuerensis sp. nov., isolated from a bird’s nest. Int J of Syst Evol Microbiol 59:139–143
Yamamoto S, Harayama S (1995) PCR amplification and direct sequencing of gyrB genes with universal primers and their application to the detection and taxonomic analysis of Pseudomonas putida strains. Appl Environ Microbiol 61:1104–1109
Yamamoto S, Kasai H, Arnold DL, Jackson RW, Vivian A, Harayama S (2000) Phylogeny of the genus Pseudomonas: intrageneric structure reconstructed from the nucleotide sequences of gyrB and rpoD genes. Microbiology 146:2385–2394
Yang GQ, Chen M, Yu Z, Lu Q, Zhou SG (2013) Bacillus composti sp. nov. and Bacillus thermophilus sp. nov., two thermophilic Fe(III)-reducing bacteria isolated from a compost. Int J Syst Evol Microbiol doi. doi:10.1099/ijs.0.049106-0
Acknowledgments
This work was supported by the Team Project of Guangdong Natural Science Foundation (S2011030002882, S2012030006114 and S2011010002452) and the Science and Technology Planning Project of Guangdong Province, China (2012B010500035 and 2012B030800008).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Zhen Yu and Ming Chang have contributed equally to this work.
The GenBank accession numbers for the 16S rRNA and gyrB gene sequences of strain WM-3T are KF317694 and KF317695, respectively, and the GenBank accession number for gyrB gene sequence of Pseudomonas linyingensis CGMCC 1.10701T is KF056321.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, Z., Chang, M., Wu, M. et al. Pseudomonas oryzae sp. nov. isolated from a paddy soil in South China. Arch Microbiol 195, 815–822 (2013). https://doi.org/10.1007/s00203-013-0930-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-013-0930-6