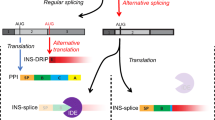
Abstract
Aims/hypothesis
The inflammatory milieu characteristic of insulitis affects translation fidelity and generates defective ribosomal products (DRiPs) that participate in autoimmune beta cell destruction in type 1 diabetes. Here, we studied the role of early innate cytokines (IFNα) and late immune adaptive events (IFNɣ) in insulin DRiP-derived peptide presentation to diabetogenic CD8+ T cells.
Methods
Single-cell transcriptomics of human pancreatic islets was used to study the composition of the (immuno)proteasome. Specific inhibition of the immunoproteasome catalytic subunits was achieved using siRNA, and antigenic peptide presentation at the cell surface of the human beta cell line EndoC-βH1 was monitored using peptide-specific CD8 T cells.
Results
We found that IFNγ induces the expression of the PSMB10 transcript encoding the β2i catalytic subunit of the immunoproteasome in endocrine beta cells, revealing a critical role in insulin DRiP-derived peptide presentation to T cells. Moreover, we showed that PSMB10 is upregulated in a beta cell subset that is preferentially destroyed in the pancreases of individuals with type 1 diabetes.
Conclusions/interpretation
Our data highlight the role of the degradation machinery in beta cell immunogenicity and emphasise the need for evaluation of targeted immunoproteasome inhibitors to limit beta cell destruction in type 1 diabetes.
Data availability
The single-cell RNA-seq dataset is available from the Gene Expression Omnibus (GEO) using the accession number GSE218316 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE218316).
Graphical Abstract

Similar content being viewed by others

Introduction
Type 1 diabetes is characterised by the progressive destruction of insulin-producing beta cells by CD8+ T cells [1]. While impaired central and peripheral immunological tolerance in combination with low-affinity T cells have been suspected to play a role in the immune attack [2], the demonstration that naive autoreactive T cells are part of the normal T cell repertoire supports the participation of beta cells and the islet microenvironment in triggering or driving disease progression [3, 4]. We have shown that local inflammation and the unfolded protein response to stress disturb the cellular equilibrium and affect translation fidelity, generating defective ribosomal products (DRiPs) and alternative reading frame-encoded peptides that the immune system is trained to act on [5]. Although the molecular mechanisms leading to processing and presentation of these ‘junk’ polypeptides remain unclear, the presence of insulin-derived defective ribosomal product (INS-DRiP)-specific CD8 T cells, with an effector phenotype in donors with type 1 diabetes, supports their potential relevance as autoimmune T cell targets [6, 7].
Upon inflammation or during cellular stress, misfolded proteins that have accumulated in the ER are redirected to the ubiquitin–proteasome system pathway for degradation to restore cellular homeostasis. Dysregulation of this system and inappropriate processing may lead to cytoplasmic aggregates and the manifestation of neurodegenerative diseases [8] and type 2 diabetes [9], as well as to the development of autoimmunity. In proteasome-associated autoinflammatory syndromes, mutations within the genes encoding the immunoproteasome catalytic subunits are associated with an excessive IFN response [10]. The proteasome is the main component of the antigen presentation machinery. In homeostatic conditions the proteasome consists of a 20S catalytic barrel and two regulatory 19S complexes, forming the 26S holo structure. Within the 20S core reside the β subunits—β1 (encoded by PSMB6), β2 (PSMB7) and β5 (PSMB5)—which display caspase, trypsin and chymotrypsin cleavage specificities, respectively. Upon inflammation, these subunits are partially or completely substituted by the induced counter forms β1i/low-molecular mass polypeptide (LMP)2 (encoded by PSMB9), β2i/multicatalytic endopeptidase complex-like-1 (MECL1; PSMB10) and β5i/LMP7 (PSMB8), respectively, to generate intermediate or full immunoproteasomes [11], affecting proteolytic activity and producing qualitative or quantitative differences in the peptide ligandome presented by MHC class I molecules [12, 13].
In this study, we investigated the composition of the (immuno)proteasome in human beta cells under inflammatory conditions using single-cell RNA-seq analysis to determine the molecular mechanisms underlying the immunogenicity of INS-DRiP.
Methods
Cells and reagents
The human beta cell line EndoC-βH1 (Human Cell Design, Paris, France), mycoplasma free, kindly provided by R. Scharfmann (Paris Descartes University, France) [14], was maintained in low-glucose DMEM supplemented with 5.5 μg/ml human transferrin, 10 mmol/l nicotinamide, 6.7 ng/ml selenite, 50 μmol/l β-mercaptoethanol, 2% human albumin (wt/vol), 100 units/ml penicillin and 100 μg/ml streptomycin. Cells were seeded in extracellular matrix (fibronectin) pre-coated culture plates. Inflammatory stress was induced with IFNα (PBL Bioscience, USA), IFNγ (Bio-Techne, USA) or IL1β (Sigma-Aldrich, USA) at the concentrations and for the times indicated. INS-DRiP-directed cytotoxic T lymphocytes (CTLs) were isolated from freshly isolated peripheral blood mononuclear cells (PBMC) from a long-term HLA-A2+ individual with type 1 diabetes. As described previously [7], 150,000 PBMC/well were seeded with 10 μg/ml DRiP1–9 peptide in Iscove's modified Dulbecco's medium (IMDM; Life Technologies, USA) supplemented with 10% human serum, 0.5% LeucoA, 0.1 ng/ml IL-12, 10 ng/ml IL-7, 25 U/ml IL-2 and 5 ng/ml IL-15. After 14 days of culture, cells were restimulated specifically with irradiated DRiP1–9 peptide-pulsed JY cells (2 μg/ml peptide with 10 × 106 cells in AIM-V medium (Life Technologies, USA) for 2 h at 37°C and 100,000 cells/well irradiated allogeneic PBMCs in IMDM supplemented with human serum and cytokines as described above. JY (ATCC 77441) cells, mycoplasma free, were maintained in IMDM supplemented with 8% FCS, 100 U/ml penicillin and 100 μg/ml streptomycin.
Lentivirus production and transduction
The lentiviral vector containing HLA-A*02:01 under the elongation factor 1α (EF1α) promotor was obtained from R. J. Lebbink (Medical Microbiology, University Medical Center Utrecht, Utrecht, the Netherlands) [16] and produced as described previously [15]. Briefly, the EF1α-HLA-A*02:01 containing lentiviral vector and the three ‘helper’ plasmids (encoding HIV-1 gag–pol, HIV-1 rev and VSV-G envelope) were co-transfected overnight using polyethylenimine into 293T cells. The medium was refreshed and viruses were harvested after 48 and 72 h, passed through 0.45 μm filters and stored at −80°C. Viral supernatants (multiplicity of infection [MOI]=2) were added to EndoC-βH1 cells in fresh medium supplemented with 8 μg/ml Polybrene (Sigma-Aldrich, USA) and the cells were incubated overnight.
siRNA transfection
Transfection of siRNAs (SMARTpool) was performed using the Dharmacon transfection reagent (Dharmafect1) according to the manufacturer’s instructions (Horizon Discovery, USA). EndoC-βH1 cells were transfected in 12-well plates for 72 h, using a final concentration of 5 nmol/l total siRNA pool mix.
RT-PCR
Total RNA was extracted from EndoC-βH1 cells using the NucleoSpin Kit (740609.50S, Bioke, the Netherlands). Approximately 0.5 µg of RNA was used for reverse transcription. Oligo (dT) primers were used in the reactions. Expression of the transcript of interest was detected using primers listed in ESM Table 1.
Western blot analyses
EndoC-βH1 cells were lysed in RIPA buffer supplemented with a protease inhibitor cocktail (Roche Applied Science, Germany). Protein quantification was performed using the BCA protein assay kit (Thermo Fisher Scientific, USA). A total of 25 µg of protein was subjected to electrophoresis on 12% acrylamide/bis acrylamide SDS page gels. After electrophoresis, proteins were transferred onto nitrocellulose membranes (GE Healthcare, USA). Membranes were stained with primary antibodies overnight at 4°C and secondary HRP-conjugated antibodies (Santa Cruz Biotechnology, USA) for 1h at room temperature. Primary antibodies were from Enzo Life Sciences (Switzerland) and were used at a dilution of 1:1000 (β1i: BML-PW8345; β5i: BML-PW8355; β2i: BML-PW8350). The loading control was β-actin (MAB1501, EMD Millipore, USA) and was used at 1:1000 dilution. Secondary antibodies were anti-mouse (#G21040) or anti-rabbit (#sc-2004) antibodies from Santa Cruz Biotechnology and were used at a dilution of 1:5000. Western ECL substrate was used for imaging (1705062, BioRad, USA).
Pancreatic islet treatment
Pancreatic islets used in the T cell co-culture assays were obtained from a human organ donor. Research consent was obtained according to national law and regulations. The islets were isolated in the GMP (good manufacturing practice) facility of Leiden University Medical Center according to a previously reported protocol [17]. For experimental use, human islets were maintained in ultra-low attachment plates (Corning, USA) in low-glucose DMEM supplemented with 10% FBS, 100 U/ml penicillin and 100 μg/ml streptomycin. Dispersed islet cells were treated with 1000 U/ml IFNγ or 2000 U/ml IFNα for 24 h. All methods were performed in accordance with relevant guidelines and regulations.
Human islets used in single-cell RNA-seq were provided through the Integrated Islet Distribution Program (IIDP). Pancreatic islets, isolated from three healthy donors, with a purity of at least 90%, were cultured on receipt in regular CMRL 1066 medium (5.5 mmol/l glucose) supplemented with 10% FCS, 20 mg/ml ciprofloxacin, 50 mg/ml gentamycin, 2 mmol/l l-glutamine, 10 mmol/l HEPES and 1.2 mg/ml nicotinamide. Islets were maintained in culture at 37°C in a 5% CO2 humidified atmosphere and medium was refreshed on receipt and every 2 days thereafter. Intact islets were treated with the following cytokines: 2000 U/ml IFNα and a combination of 1 ng/ml IL1β and 1000 U/ml IFNγ (R&D Systems, USA). After treatments, islets were dispersed into single cells using 0.025% trypsin (Gibco, USA) and 10 mg/ml Dnase (Pulmozyme, Genentech, USA). Single cells were then processed for single-cell RNA-seq following the standard 10x Genomics 3’ V3 chemistry protocol (10x Genomics, USA).
RNA-seq data processing and analysis
Single-cell RNA-seq output was processed and analysed following the Seurat pipeline (version 4.0; https://cran.r-project.org/package=Seurat). Clustered cells were labelled and sorted into subsets based on the expression of canonical cell markers: insulin (beta cells), glucagon (alpha cells), somatostatin (delta cells), pancreatic polypeptide (pancreatic polypeptide/gamma cells), human cationic trypsinogen (acinar cells) and keratin 19 (duct cells). Differential gene expression analysis was performed using the Wilcoxon rank sum test to analyse gene expression differences between treated and untreated cell groups. The output of this analysis shows genes that are expressed in at least 25% of the cells in any group (treated or untreated). Bonferroni correction was applied to adjust the p values. Up-/downregulated genes (log fold change [FC] >0.5 and <–0.5 respectively) with an adjusted p value of <0.05 were considered to be significantly altered by the treatment.
Analyses of the pseudobulk differential expression of the endocrine cells was performed by clusterProfiler using the enrichPathway classification (https://bioconductor.org/packages/release/bioc/html/ReactomePA.html) [18].
T cell activation assays
Target cells were harvested and co-cultured with CTLs specific for INS-DRiP1–9 (MLYQHLLPL) [7] or preproinsulin (PPI)15–24 (ALWGPDPAAA) [19] at an effector/target ratio of 2:1. Co-cultures were incubated at 37°C for 4 h in IMDM supplemented with 10% human albumin and 40 U/ml IL-2 (Novartis, Switzerland). The supernatant was used for detection of macrophage inflammatory protein-1 beta (MIP-1β) production by T cells, using the MIP-1β ELISA kit (88–7034–22, Thermo Fisher Scientific), according to the manufacturer’s protocol.
Statistical analysis
Data are presented as means ± SEM. Calculations were performed using GraphPad Prism 7 (GraphPad Software, USA). Unpaired t tests were carried out for all comparisons.
Results
Type II but not type I IFN enhances INS-DRiP-derived peptide presentation to specific CTLs
Although we previously provided evidence for a role of INS-DRiP-directed T cells in beta cell destruction in type 1 diabetes [7], the contribution of the islet microenvironment that is characteristic of the early disease phase (the presence of type I IFN as a possible consequence of exposure to microorganisms, disturbed metabolism and tissue stress) or later disease phases (the presence of type II IFN resulting from activation of the adaptive immune response) remains unknown [20].
To dissect whether and when INS-DRiP-derived peptide recognition is altered in the course of disease progression, we co-cultured dispersed HLA-A2+ human islets treated with IFNα (type I) or IFNɣ (type II) with an HLA-A2-specific CD8+ T cell clone, isolated from an individual with type 1 diabetes and directed against the N-terminal part of the INS-DRiP polypeptide (INS-DRiP1–9) [7] (Fig. 1a). Under these standardised conditions, previously described to mimic the islet inflammatory milieu [21,22,23,24,25], although IFNα did not trigger additional T cell activation compared with untreated islets, IFNɣ treatment caused an increase in MIP-1β secretion, indicating enhanced INS-DRiP peptide presentation (Fig. 1b). To validate the assay and confirm the correlation between the amount of peptide presented at the cell surface and the increase in MIP-1β secretion by T cells, we pulsed JY cells (HLA-A2+) with increasing titres of cognate peptide derived from INS-DRiP1–9 or HLA-A2 peptide derived from native PPI15–24. As expected, INS-DRiP CTLs were highly specific and the level of activation was proportional to the peptide concentration (ESM Fig. 1).
IFNα and IFNγ differentially regulate INS-DRiP presentation in human islets. (a) Schematic of the islet cells/INS-DRiP-specific CTLs co-culture experiment in the presence of 2000 U/ml IFNα or 1000 U/ml IFNγ. Created with BioRender.com. (b) MIP-1β secretion by INS-DRiP-specific CTLs after co-culture with HLA-A2+ primary human islet cells treated with IFNα or IFNγ for 24 h (n=3). NT, non-treated
Considering that HLA class I is the most important variable involved in antigen presentation to CD8 T cells, we used a single-cell RNA-seq dataset of human islets exposed to IFNα or IFNγ/IL1β (ESM Fig. 2) and evaluated the effect of both treatments on HLA-A expression in endocrine cells. As expected, both cytokines upregulated HLA-A expression to similar levels (Fig. 2a), suggesting that additional mechanisms are involved during protein processing after IFNɣ treatment to increase INS-DRiP-derived peptide presentation.
Cytokines differentially regulate immunoproteasome catalytic subunit expression in primary human endocrine cells. (a) Violin plots showing the expression levels of HLA-A after IFNα or IFNγ/IL1β treatment of endocrine cells. (b) Schematic representation of the composition of the proteasome and immunoproteasome. (c) Violin plots showing the expression levels of the mRNAs encoding the constitutive (PSMB5, PSMB6 and PSMB7) and induced (PSMB8, PSMB9 and PSMB10) catalytic subunits of the (immuno)proteasome. Expression levels correspond to log2 normalised counts/cell, as obtained in single-cell RNA-seq of human islets treated with IFNα (2000 U/ml) or IFNγ (1000 U/ml)+IL1β (2 ng/ml) for 24 h. NT, non-treated
In our previous proteogenomic study [5], we demonstrated that IFNγ/IL1β treatment did not increase INS transcript expression nor favour ribosome docking to the DRiP start codon (as determined by long RNA-seq and ribosome profiling of EndoC-βH1 cells), suggesting that DRiP is produced equally in normal and inflamed conditions. Therefore, we reasoned that a difference in the protein degradation machinery may provide a rational explanation for the observed increase in INS-DRiP presentation after IFNɣ treatment. Using single-cell RNA-seq, we analysed the effect of IFNα and IFNγ/IL1β on the expression of the mRNAs encoding the constitutive catalytic subunits of the proteasome (β1 [encoded by PSMB6], β2 [PSMB7], β5 [PSMB5]) and the induced catalytic subunits of the immunoproteasome (β1i [encoded by PSMB9], β2i [PSMB10], β5i [PSMB8]) (Fig. 2b). While the constitutive subunits were not significantly affected by treatment, PSMB8 and PSMB9 mRNAs were upregulated by both IFNα and IFNɣ/IL1β in all endocrine and exocrine cell types (Fig. 2c and ESM Fig. 3). In contrast, the expression of PSMB10 was significantly increased in endocrine alpha and beta cells after IFNɣ/IL1β treatment but not in delta or exocrine cells (ESM Table 2).
To confirm these results and determine the main driver for the upregulation of PSMB10 observed after IFNɣ/IL1β treatment in primary human beta cells, we exposed EndoC-βH1cells to recombinant IFNα, IFNɣ or IL1β for increasing amounts of time. As expected, we noted a rapid induction of β1i (encoded by PSMB9) and β5i (PSMB8) after IFNα and IFNɣ treatment and the absence of expression of β2i (PSMB10) in IFNα-treated cells (Fig. 3a). The absence of effect of IL1β on both HLA class I surface expression and immunoproteasome components (ESM Fig. 4) in these assays suggests that the expression of PSMB10 observed in primary human islets on IFNɣ/IL1β treatment is mainly driven by type II IFN. To dissect the impact of cytokines on EndoC-βH1 cell immunogenicity, we generated a stable EndoC-βΗ1 cell line expressing HLA-A*02:01 by lentiviral transduction and exposed the cells for 4, 16 or 24 h to IFNα or IFNγ prior to co-culture with HLA-A2-restricted INS-DRiP-specific CD8+ T cells. We measured HLA class I expression in EndoC-βΗ1 cells and T cell activation by MIP-1β secretion. While IFNα and IFNγ differentially affected HLA class I surface expression, both treatments equally upregulated the expression of the HLA-A2 transgene (Fig. 3b). Under these conditions, and despite similar HLA-A2 expression levels, we observed an increase in T cell activation after IFNγ treatment, illustrating increased presentation of the DRiP-derived peptide to the specific CTLs, as observed for human islets (Fig. 3c).
Proteasome composition in EndoC-βH1 cells after exposure to cytokines and sensitivity to INS-DRiP-specific CTLs. (a) β5i (encoded by PSMB8), β1i (PSMB9) and β2i (PSMB10) protein expression after incubation with 2000 U/ml IFNα or 1000 U/ml IFNγ determined by Western blot analysis. Actin was used as an internal control. Treatment duration was 4, 16 or 24 h, as indicated. (b) Surface expression of HLA-A/B/C (dashed lines) and HLA-A2 (solid lines) on EndoC-βΗ1-HLA-A2 cells treated with IFNα (orange lines) or IFNγ (red lines) for 4, 16 and 24 h. Values are presented as mean fluorescence intensity (MFI). (c) INS-DRiP-specific CTL activation determined by MIP-1β secretion after co-culture with EndoC-βΗ1-HLA-A2 cells treated with IFNα (orange line) or IFNγ (red line) for 4, 16 and 24 h. n=3 independent experiments. ***p≤ 0.001. NT, non-treated
The β2i subunit enhances INS-DRiP presentation
To determine the effect of the different immunoproteasome subunits on EndoC-βH1-HLA-A2 cell recognition by the INS-DRiP CD8+ T cell clone, siRNAs specific for PSMB8, PSMB9 and PSMB10 were used to selectively interfere with gene expression prior to IFNα and IFNγ treatment (Fig. 4a–c). We found that downregulation of the immunoproteasome catalytic subunits had no impact on HLA-A2 surface expression (Fig. 5a). To test for their immunogenicity, the modified beta cells were co-cultured with autoreactive T cells directed against INS-DRiP1–9 or PPI15–24 [26]. While activation of the PPI-specific T cell clone was unaltered after IFN treatment and immunoproteasome subunit modulation, PSMB10 inhibition annihilated the IFNγ effect and reduced INS-DRiP-specific T cell reactivity to the levels seen after treatment with IFNα (Fig. 5b,c). Of note, the increased epitope presentation in the presence of IFNα after PSMB9 knockdown is peculiar but may be related to a higher amount of intact epitope, as suggested by the presence in the epitope of hydrophobic residues targeted for chymotrypsin-like cleavage by β1i [27] (ESM Fig. 5). Accordingly, in silico analysis of the INS-DRiP sequence on the Proteasome Cleavage Prediction Server (PCPS) [28] shows that the INS-DRiP1–9 (MLYQHLLPL) 9-mer has a higher propensity to be processed by the immunoproteasome complex than by the constitutive proteasome. In addition, the localisation of trypsin digestion sites outside the 9-mer epitope region suggests that the β2i subunit may be involved in the processing of the INS-DRiP polypeptide but should not destroy the MLYQHLLPL epitope, meaning that its integrity is maintained.
Immunoproteasome silencing in the EndoC-βΗ1 cell line. (a) PSMB8, (b) PSMB9 and (c) PSMB10 mRNA expression in EndoC-βΗ1-HLA-A2 cells transfected with non-targeted siRNA (siCTRL) and siRNAs specific for PSMB8, PSMB9 and PSMB10. Cells were treated with 2000 U/ml IFNα (orange bars), 1000 U/ml IFNγ (red bars) or control medium (blue bars) 72 h post transfection for 24 h. Gene expression levels were corrected for levels of the housekeeping gene GAPDH and are presented as the induction ratio (relative to levels for siCTRL(1) in IFNα samples and siCTRL(2) for IFNγ samples) (n=3). *p≤ 0.05, **p≤ 0.01, ***p≤ 0.001
PSMB10 silencing exclusively reduces INS-DRiP-specific CTL activation. (a) HLA-A2 (genetically introduced) surface expression of EndoC-βΗ1-HLA-A2 cells transfected with non-targeted siRNA (siCTRL) and siRNAs specific for PSMB8, PSMB9 and PSMB10. Cells were left untreated (blue bars) or treated with 2000 U/ml IFNα (orange bars) or 1000 U/ml IFNγ (red bars) 72 h post transfection for 24 h. Values are presented as mean fluorescence (FITC) intensity (MFI) (n=3). (b, c) MIP-1β secretion by (b) INS-DRiP1–9-specific CTLs and (c) PPI15–24 after co-culture with EndoC-βΗ1-HLA-A2 cells. Prior to co-culture, target cells were transfected with non-targeted siRNA and siRNAs specific for PSMB8, PSMB9 and PSMB10. Cells were left untreated (blue circles) or treated with IFNα (orange circles) or IFNγ (red circles) 72 h post transfection for 24 h (n=3). Statistical analyses are comparing IFNγ to IFNα treatments. ***p≤0.001. NT, non-treated
PSMB10, encoding the β2i subunit, is specifically expressed in a beta cell subset that is preferentially destroyed in pancreases of individuals with type 1 diabetes
We next assessed the potential clinical relevance of our findings in the context of the immunopathogenesis of type 1 diabetes using the single-cell multi-omics analysis launched by the Human Pancreas Analysis Program (HPAP) consortium [29]. This dataset identified two distinct beta cell clusters (beta-1 and beta-2) when comparing the transcriptomic profile of endocrine cells from donors with type 1 diabetes and endocrine cells from autoantibody-positive donors (AAb+). Those clusters are differentiated by the specific upregulation of apoptotic and adaptive immune system signalling in the beta-2 subset in AAb+, indicating that this cluster is undergoing cell death. Further examination of the pseudobulk differential expression analysis of the endocrine cells showed upregulation of the apoptosis pathway specifically in the beta-2 (‘minor’) beta cell subset (Fig. 6a, ESM Fig. 6 and ESM Table 3) and pointed to differences in the composition of the catalytic subunits of the immunoproteasome and to the selective expression of PSMB10 in this subset (Fig. 6b,c). Altogether our data demonstrate the differential immune visibility of beta cells in the early and late phases of disease progression in type 1 diabetes and illustrate how IFNɣ can accelerate beta cell destruction by changing the composition of the immunoproteasome.
Immunoproteasome composition of beta cells from individuals with type 1 diabetes. (a) Gene ontology plot of transcripts upregulated in two distinct beta cell subsets (p<0.05 and logFC>1) in individuals with type 1 diabetes compared with control individuals (donors without diabetes). The plot was generated with clusterProfiler using the enrichPathway classification. (b) Heatmap showing the logFC (p<0.05) of PSMB8, PSMB9 and PSMB10 in the beta-1 and beta-2 populations from individuals with type 1 diabetes compared with control donors. (c) Heatmap showing the logFC (p<0.05) of PSMB8, PSMB9 and PSMB10 in the beta-1 and beta-2 populations from individuals with type 1 diabetes patients compared with AAb+ individuals. This figure has been generated using data described in Fasolino et al, 2022 [29]
Discussion
In this study we show that IFNα and IFNγ differentially regulate the proteasomal composition of beta cells and demonstrate that INS-DRiP peptide recognition may be involved in later phases of type 1 diabetes pathogenesis as an amplificatory phenomenon of beta cell destruction. Previous studies have investigated the effect of IFNα and IFNγ/IL1β on human islets using bulk RNA-seq and shown increased expression of the immunoproteasome catalytic subunits [30, 31]. In this study we found that PSMB10 expression, responsible for INS-DRiP processing, represents a unique feature of beta cells during the late phase of type 1 diabetes, enhancing beta cell immunogenicity and discriminating beta cells that are targeted by the immune system from those that are protected. This observation adds to the accumulating evidence that the islet microenvironment acting in concert with beta cells controls immunoreactivity by altering antigen presentation pathways [26, 32]. The composition of the proteasome complex in beta cells has been studied in primary human islets and rodent insulin-producing cell lines after cytokine stimulation [33, 34]. However, few studies have connected an altered catalytic core composition with functional differences [35]. To our knowledge, our data are the first showing direct evidence for the participation of the immunoproteasome in beta cell immunogenicity and INS-DRiP-derived peptide presentation to CTLs.
Potential limitations of this study include the use of recombinant cytokines that simplify and may only partially reflect the local inflammation seen during early and late events of type 1 diabetes progression [36]. In addition, the use of a transformed human beta cell line and the limited number of autoreactive T cell clones tested mean that further validation in primary human islets in combination with other DRiP-specific T cells, when available, is required to draw broader conclusions about β2i participation in the processing of peptides derived from defective ribosomal products (currently the MLYQHLLPL peptide, studied here, remains the only alternative translational product-derived peptide identified in beta cells). Of note, while HLA-A2 expression in the EndoC-βH1 transductants [5] may not mirror normal physiological behaviour, its stable expression under different conditions allows an unbiased comparison of INS-DRiP peptide processing after IFNα or IFNɣ treatment. In addition, the selective upregulation of PSMB10 in a subset of vulnerable beta cells reinforces the relevance of our findings and points to a role of β2i in the processing and presentation of islet neoantigens to diabetogenic T cells in type 1 diabetes.
Inhibitors targeting both the standard proteasome and the immunoproteasome, such as bortezomib, are clinically approved for the treatment of multiple myeloma and have been shown to have some beneficial effects in autoimmunity and transplantation [37,38,39]. However, the lack of specificity results in severe side effects, impairing their therapeutic value [40,41,42]. Similarly, in our analysis we observed that PSMB9 inhibition sensitised beta cells to CTL-mediated destruction, illustrating that general interference comes at a risk and highlighting the need for specific inhibitors rather than general proteasome suppression. Immunoproteasome inhibitors have been shown to ameliorate symptoms in preclinical models of different autoimmune diseases [43, 44], and KZR-616, a drug targeting both PSMB8 and PSMB9, has been tested in a Phase II trial of systemic lupus erythematosus [45]. In each study, inhibitors were used as immunosuppressants through their capacity to target immune cells that constitutively express the immunoproteasome. Some of the outcomes observed were decreased cytokine secretion, inhibition of T and B cell activation and modified macrophage polarisation [40]. While these effects may be beneficial in type 1 diabetes, a recent study showed that the use of ONX-914, a PSMB8 inhibitor, exacerbated beta cell apoptosis during inflammation [34]. ONX-914 was later shown to have some off-target effects on PSMB9 [46], but whether this occurred in the inflamed human islets has not yet been determined. In fact, at least two immunoproteasome subunits should be targeted to obtain successful immunosuppression [40, 46]. The use of immunoproteasome inhibitors to alleviate non-immune tissue inflammation has not yet been explored. Our study suggests that the selective modulation of MECL1 (encoded by PSMB10) using selective cell-permeable inhibitors of the trypsin-like site [47] may be beneficial to reduce beta cell immunogenicity and the T cell response mounted against the INS-DRiP peptide to limit type 1 diabetes progression, shedding new light on the role of the immunoproteasome, rather than the constitutive proteasome, as an important player in beta cell immunogenicity.
Abbreviations
- AAb+:
-
Autoantibody-positive
- CTL:
-
Cytotoxic T lymphocyte
- DRiP:
-
Defective ribosomal product
- EF1α:
-
Elongation factor 1α
- FC:
-
Fold change
- IMDM:
-
Iscove's modified Dulbecco's medium
- INS-DRiP:
-
Insulin-derived defective ribosomal product
- LMP2:
-
Low-molecular mass polypeptide
- MECL-1:
-
Multicatalytic endopeptidase complex-like-1
- MIP-1β:
-
Macrophage inflammatory protein-1 beta
- PBMC:
-
Peripheral blood mononuclear cells
- PPI:
-
Preproinsulin
References
Pugliese A (2017) Autoreactive T cells in type 1 diabetes. J Clin Invest 127(8):2881–2891. https://doi.org/10.1172/JCI94549
Abreu JR, Martina S, Verrijn Stuart AA et al (2012) CD8 T cell autoreactivity to preproinsulin epitopes with very low human leucocyte antigen class I binding affinity. Clin Exp Immunol 170(1):57–65. https://doi.org/10.1111/j.1365-2249.2012.04635.x
Bender C, Rodriguez-Calvo T, Amirian N, Coppieters KT, von Herrath MG (2020) The healthy exocrine pancreas contains preproinsulin-specific CD8 T cells that attack islets in type 1 diabetes. Sci Adv 6(42):eabc5586. https://doi.org/10.1126/sciadv.abc5586
Culina S, Lalanne AI, Afonso G et al (2018) Islet-reactive CD8(+) T cell frequencies in the pancreas, but not in blood, distinguish type 1 diabetic patients from healthy donors. Sci Immunol 3(20):eaao4013. https://doi.org/10.1126/sciimmunol.aao4013
Thomaidou S, Slieker RC, van der Slik AR et al (2021) Long RNA sequencing and ribosome profiling of inflamed beta-cells reveal an extensive translatome landscape. Diabetes 70(10):2299–2312. https://doi.org/10.2337/db20-1122
Anderson AM, Landry LG, Alkanani AA et al (2021) Human islet T cells are highly reactive to preproinsulin in type 1 diabetes. Proc Natl Acad Sci U S A 118(41):e2107208118. https://doi.org/10.1073/pnas.2107208118
Kracht MJ, van Lummel M, Nikolic T et al (2017) Autoimmunity against a defective ribosomal insulin gene product in type 1 diabetes. Nat Med 23(4):501–507. https://doi.org/10.1038/nm.4289
Zheng Q, Huang T, Zhang L et al (2016) Dysregulation of ubiquitin-proteasome system in neurodegenerative diseases. Front Aging Neurosci 8:303. https://doi.org/10.3389/fnagi.2016.00303
Press M, Jung T, Konig J, Grune T, Hohn A (2019) Protein aggregates and proteostasis in aging: amylin and beta-cell function. Mech Ageing Dev 177:46–54. https://doi.org/10.1016/j.mad.2018.03.010
Sarrabay G, Mechin D, Salhi A et al (2020) PSMB10, the last immunoproteasome gene missing for PRAAS. J Allergy Clin Immunol 145(3):1015–1017. https://doi.org/10.1016/j.jaci.2019.11.024. (e1016)
Thomaidou S, Zaldumbide A, Roep BO (2018) Islet stress, degradation and autoimmunity. Diabetes Obes Metab 20(Suppl 2):88–94. https://doi.org/10.1111/dom.13387
Mishto M, Liepe J, Textoris-Taube K et al (2014) Proteasome isoforms exhibit only quantitative differences in cleavage and epitope generation. Eur J Immunol 44(12):3508–3521. https://doi.org/10.1002/eji.201444902
Vigneron N, Van den Eynde BJ (2014) Proteasome subtypes and regulators in the processing of antigenic peptides presented by class I molecules of the major histocompatibility complex. Biomolecules 4(4):994–1025. https://doi.org/10.3390/biom4040994
Ravassard P, Hazhouz Y, Pechberty S et al (2011) A genetically engineered human pancreatic beta cell line exhibiting glucose-inducible insulin secretion. J Clin Invest 121(9):3589–3597. https://doi.org/10.1172/JCI58447
Carlotti F, Bazuine M, Kekarainen T et al (2004) Lentiviral vectors efficiently transduce quiescent mature 3T3-L1 adipocytes. Mol Ther 9(2):209–217. https://doi.org/10.1016/j.ymthe.2003.11.021
van der Torren CR, Zaldumbide A, Roelen DL et al (2016) Innate and adaptive immunity to human beta cell lines: implications for beta cell therapy. Diabetologia 59(1):170–175. https://doi.org/10.1007/s00125-015-3779-1
Ricordi C, Lacy PE, Finke EH, Olack BJ, Scharp DW (1988) Automated method for isolation of human pancreatic islets. Diabetes 37(4):413–420. https://doi.org/10.2337/diab.37.4.413
Yu G, He QY (2016) ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol Biosyst 12(2):477–479. https://doi.org/10.1039/c5mb00663e
Skowera A, Ellis RJ, Varela-Calvino R et al (2008) CTLs are targeted to kill beta cells in patients with type 1 diabetes through recognition of a glucose-regulated preproinsulin epitope. J Clin Invest 118(10):3390–3402. https://doi.org/10.1172/JCI35449
Colli ML, Szymczak F, Eizirik DL (2020) Molecular footprints of the immune assault on pancreatic beta cells in type 1 diabetes. Front Endocrinol (Lausanne) 11:568446. https://doi.org/10.3389/fendo.2020.568446
Colli ML, Hill JLE, Marroqui L et al (2018) PDL1 is expressed in the islets of people with type 1 diabetes and is up-regulated by interferons-alpha and-gamma via IRF1 induction. EBioMedicine 36:367–375. https://doi.org/10.1016/j.ebiom.2018.09.040
Demine S, Schiavo AA, Marin-Canas S, Marchetti P, Cnop M, Eizirik DL (2020) Pro-inflammatory cytokines induce cell death, inflammatory responses, and endoplasmic reticulum stress in human iPSC-derived beta cells. Stem Cell Res Ther 11(1):7. https://doi.org/10.1186/s13287-019-1523-3
Nakayasu ES, Syed F, Tersey SA et al (2020) Comprehensive proteomics analysis of stressed human islets identifies GDF15 as a target for type 1 diabetes intervention. Cell Metab 31(2):363–374. https://doi.org/10.1016/j.cmet.2019.12.005. (e366)
Ortis F, Cardozo AK, Crispim D, Storling J, Mandrup-Poulsen T, Eizirik DL (2006) Cytokine-induced proapoptotic gene expression in insulin-producing cells is related to rapid, sustained, and nonoscillatory nuclear factor-kappaB activation. Mol Endocrinol 20(8):1867–1879. https://doi.org/10.1210/me.2005-0268
Ramos-Rodriguez M, Raurell-Vila H, Colli ML et al (2019) The impact of proinflammatory cytokines on the beta-cell regulatory landscape provides insights into the genetics of type 1 diabetes. Nat Genet 51(11):1588–1595. https://doi.org/10.1038/s41588-019-0524-6
Thomaidou S, Kracht MJL, van der Slik A et al (2020) beta-cell stress shapes CTL immune recognition of preproinsulin signal peptide by posttranscriptional regulation of endoplasmic reticulum aminopeptidase 1. Diabetes 69(4):670–680. https://doi.org/10.2337/db19-0984
Ferrington DA, Gregerson DS (2012) Immunoproteasomes: structure, function, and antigen presentation. Prog Mol Biol Transl Sci 109:75–112. https://doi.org/10.1016/B978-0-12-397863-9.00003-1
Gomez-Perosanz M, Ras-Carmona A, Reche PA (2020) PCPS: a web server to predict proteasomal cleavage sites. Methods Mol Biol 2131:399–406. https://doi.org/10.1007/978-1-0716-0389-5_23
Fasolino M, Schwartz GW, Patil AR et al (2022) Single-cell multi-omics analysis of human pancreatic islets reveals novel cellular states in type 1 diabetes. Nat Metab 4(2):284–299. https://doi.org/10.1038/s42255-022-00531-x
Colli ML, Ramos-Rodriguez M, Nakayasu ES et al (2020) An integrated multi-omics approach identifies the landscape of interferon-alpha-mediated responses of human pancreatic beta cells. Nat Commun 11(1):2584. https://doi.org/10.1038/s41467-020-16327-0
Eizirik DL, Sammeth M, Bouckenooghe T et al (2012) The human pancreatic islet transcriptome: expression of candidate genes for type 1 diabetes and the impact of pro-inflammatory cytokines. PLoS Genet 8(3):e1002552. https://doi.org/10.1371/journal.pgen.1002552
Gonzalez-Duque S, Azoury ME, Colli ML et al (2018) Conventional and neo-antigenic peptides presented by beta cells are targeted by circulating naive CD8+ T cells in type 1 diabetic and healthy donors. Cell Metab 28(6):946–960. https://doi.org/10.1016/j.cmet.2018.07.007. (e946)
Khilji MS, Verstappen D, Dahlby T et al (2020) The intermediate proteasome is constitutively expressed in pancreatic beta cells and upregulated by stimulatory, low concentrations of interleukin 1 beta. PLoS One 15(2):e0222432. https://doi.org/10.1371/journal.pone.0222432
Lundh M, Bugliani M, Dahlby T et al (2017) The immunoproteasome is induced by cytokines and regulates apoptosis in human islets. J Endocrinol 233(3):369–379. https://doi.org/10.1530/JOE-17-0110
Freudenburg W, Gautam M, Chakraborty P et al (2013) Immunoproteasome activation during early antiviral response in mouse pancreatic beta-cells: new insights into auto-antigen generation in type i diabetes? J Clin Cell Immunol 4(2):141. https://doi.org/10.4172/2155-9899.1000141
Lu J, Liu J, Li L, Lan Y, Liang Y (2020) Cytokines in type 1 diabetes: mechanisms of action and immunotherapeutic targets. Clin Transl Immunology 9(3):e1122. https://doi.org/10.1002/cti2.1122
Walhelm T, Gunnarsson I, Heijke R et al (2021) Clinical experience of proteasome inhibitor bortezomib regarding efficacy and safety in severe systemic lupus erythematosus: a nationwide study. Front Immunol 12:756941. https://doi.org/10.3389/fimmu.2021.756941
Mondanelli G, Albini E, Pallotta MT et al (2017) The proteasome inhibitor bortezomib controls indoleamine 2,3-dioxygenase 1 breakdown and restores immune regulation in autoimmune diabetes. Front Immunol 8:428. https://doi.org/10.3389/fimmu.2017.00428
Khalesi N, Korani S, Korani M, Johnston TP, Sahebkar A (2021) Bortezomib: a proteasome inhibitor for the treatment of autoimmune diseases. Inflammopharmacology 29(5):1291–1306. https://doi.org/10.1007/s10787-021-00863-2
Basler M, Groettrup M (2020) Recent insights how combined inhibition of immuno/proteasome subunits enables therapeutic efficacy. Genes Immun 21(5):273–287. https://doi.org/10.1038/s41435-020-00109-1
Huber EM, Groll M (2021) A nut for every bolt: subunit-selective inhibitors of the immunoproteasome and their therapeutic potential. Cells 10(8):1929. https://doi.org/10.3390/cells10081929
von Brzezinski L, Saring P, Landgraf P, Cammann C, Seifert U, Dieterich DC (2017) Low neurotoxicity of ONX-0914 supports the idea of specific immunoproteasome inhibition as a side-effect-limiting, therapeutic strategy. Eur J Microbiol Immunol (Bp) 7(3):234–245. https://doi.org/10.1556/1886.2017.00025
Basler M, Maurits E, de Bruin G, Koerner J, Overkleeft HS, Groettrup M (2018) Amelioration of autoimmunity with an inhibitor selectively targeting all active centres of the immunoproteasome. Br J Pharmacol 175(1):38–52. https://doi.org/10.1111/bph.14069
Goetzke CC, Ebstein F, Kallinich T (2021) Role of proteasomes in inflammation. J Clin Med 10(8):1783. https://doi.org/10.3390/jcm10081783
Kirk CJ, Muchamuel T, Wang J, Fan RA (2021) Discovery and early clinical development of selective immunoproteasome inhibitors. Cells 11(1):9. https://doi.org/10.3390/cells11010009
Basler M, Lindstrom MM, LaStant JJ et al (2018) Co-inhibition of immunoproteasome subunits LMP2 and LMP7 is required to block autoimmunity. EMBO Rep 19(12):e46512. https://doi.org/10.15252/embr.201846512
Geurink PP, van der Linden WA, Mirabella AC et al (2013) Incorporation of non-natural amino acids improves cell permeability and potency of specific inhibitors of proteasome trypsin-like sites. J Med Chem 56(3):1262–1275. https://doi.org/10.1021/jm3016987
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Acknowledgements
The authors would like to thank S. Cramer and M. Rabelink (both Cell and Chemical Biology Department, Leiden University Medical Center, the Netherlands) for technical help.
Data availability
The single-cell RNA-seq dataset is available from the Gene Expression Omnibus (GEO) using the accession number GSE218316 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE218316). All data are available in the main text or the electronic supplementary material.
Funding
This work is supported by DON and the Dutch Diabetes Research Foundation and by the Innovative Medicines Initiative 2 Joint Undertaking (IMI2-JU) under grant agreement no. 115797 (INNODIA). This Joint Undertaking receives support from the EU’s Horizon 2020 research and innovation programme and the European Federation of Pharmaceutical Industries and Associations (EFPIA), JDRF and Leona M. and Harry B. Helmsley Charitable Trust. BOR was supported by the Wanek Family Project for Type 1 Diabetes.
Authors’ relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Contribution statement
ST designed and performed the experiments and wrote the manuscript. AMG analysed the single-cell RNA-seq dataset and wrote the manuscript. ARvdS, JG and SdL performed the experiments and analysed the data. FC, RCH and BOR interpreted the data and critically revised the manuscript. AZ supervised the project, designed the experiments and wrote the manuscript. All co-authors revised and approved the final manuscript. AZ is the guarantor of this work and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thomaidou, S., Munoz Garcia, A., de Lange, S. et al. IFNɣ but not IFNα increases recognition of insulin defective ribosomal product-derived antigen to amplify islet autoimmunity. Diabetologia 66, 2075–2086 (2023). https://doi.org/10.1007/s00125-023-05991-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-023-05991-8