Abstract
Aims/hypothesis
Abnormal gut microbiota and blood metabolome profiles have been reported both in children and adults with uncomplicated type 1 diabetes as well as in adults with type 1 diabetes and advanced stages of diabetic nephropathy. In this study we aimed to investigate the gut microbiota and a panel of targeted plasma metabolites in individuals with type 1 diabetes of long duration without and with different levels of albuminuria.
Methods
In a cross-sectional study we included 161 individuals with type 1 diabetes and 50 healthy control individuals. Individuals with type 1 diabetes were categorised into three groups according to historically measured albuminuria: (1) normoalbuminuria (<3.39 mg/mmol); (2) microalbuminuria (3.39–33.79 mg/mmol); and (3) macroalbuminuria (≥33.90 mg/mmol). From faecal samples, the gut microbiota composition at genus level was characterised by 16S rRNA gene amplicon sequencing and in plasma a targeted profile of 31 metabolites was analysed with ultra HPLC coupled to MS/MS.
Results
Study participants were aged 60 ± 11 years (mean ± SD) and 42% were women. The individuals with type 1 diabetes had had diabetes for a mean of 42 ± 15 years and had an eGFR of 75 ± 25 ml min−1 (1.73 m)−2. Measures of the gut microbial beta diversity differed significantly between healthy controls and individuals with type 1 diabetes, either with micro- or macroalbuminuria. Taxonomic analyses showed that 79 of 324 genera differed in relative abundance between individuals with type 1 diabetes and healthy controls and ten genera differed significantly among the three albuminuria groups with type 1 diabetes. For the measured plasma metabolites, 11 of 31 metabolites differed significantly between individuals with type 1 diabetes and healthy controls. When individuals with type 1 diabetes were stratified by the level of albuminuria, individuals with macroalbuminuria had higher plasma concentrations of indoxyl sulphate and l-citrulline than those with normo- or microalbuminuria and higher plasma levels of homocitrulline and l-kynurenine compared with individuals with normoalbuminuria. Whereas plasma concentrations of tryptophan were lower in individuals with macroalbuminuria compared with those with normoalbuminuria.
Conclusions/interpretation
We demonstrate that individuals with type 1 diabetes of long duration are characterised by aberrant profiles of gut microbiota and plasma metabolites. Moreover, individuals with type 1 diabetes with initial stages of diabetic nephropathy show different gut microbiota and plasma metabolite profiles depending on the level of albuminuria.

Graphical abstract
Similar content being viewed by others

Introduction
A large community of diverse bacteria live in the human intestinal tract in symbiosis with each other and their human host. Evidence suggests that aberrations in bacterial composition and function, also known as dysbiosis, are linked to a variety of chronic non-communicable diseases, including inflammatory bowel disease [1], type 1 and 2 diabetes [2, 3] and end-stage kidney disease (ESKD) [4, 5]. Dysbiosis, together with intestinal barrier dysfunction, are hypothesised to exacerbate abnormalities in the blood metabolome profile in individuals with type 1 diabetes [6, 7].
Diabetic nephropathy is a common microvascular complication of type 1 diabetes and a leading cause of ESKD [8, 9]. The aetiology of diabetic nephropathy is complex, including major risk factors such as hyperglycaemia and hypertension [9]. Moreover, changes in the gut microbiota and plasma metabolome occur at the various stages of diabetic nephropathy and an aberrant gut microbiota has been suggested as a co-triggering factor of kidney disease [10].
Therefore, using an explorative approach, the objective of this cross-sectional study was to add to this nascent research field by: (1) comparing intestinal microbiome profiles and plasma metabolites in individuals with type 1 diabetes of long duration with healthy control individuals of similar sex and age distribution; and (2) elucidating whether differences in gut microbiota composition and a panel of plasma metabolites levels can be detected in individuals with type 1 diabetes at initial stages of diabetic nephropathy compared with individuals with type 1 diabetes without diabetic nephropathy.
Methods
Study population
A sample of 161 individuals with type 1 diabetes who were followed at the outpatient clinic at Steno Diabetes Center Copenhagen and 50 non-diabetic healthy control individuals were recruited in a cross-sectional study during 2016–2017. Included type 1 diabetes individuals were >18 years of age and diagnosed with type 1 diabetes according to the WHO-criteria. Exclusion criteria were: (1) non-diabetic kidney disease; (2) renal failure (eGFR <15 ml min−1[1.73 m]−2), dialysis or kidney transplantation; (3) change in renin–angiotensin–aldosterone system (RAAS)-blocking treatment during the month prior to study inclusion; (4) treatment with systemic antibiotics in the 3 months prior to recruitment; and (5) treatment with systemic immunosuppressive agents. The individuals with type 1 diabetes were stratified into three groups of albuminuria based on the highest urine albumin/creatinine ratio (UACR) level measured at the study visit or documented previously in two out of three consecutive urine samples within 1 year (as albumin content in 24 h urine samples [UAER] or UACR). The three albuminuria groups consisted of 50 with normoalbuminuria (<3.39 mg/mmol corresponding to <30 mg/24 h or mg/g), 50 with microalbuminuria (3.39–33.79 mg/mmol corresponding to 30–299 mg/24 h or mg/g) and 61 with macroalbuminuria (≥33.90 mg/mmol corresponding to ≥300 mg/24 h or mg/g). Participants classified as having normoalbuminuria did not have any recorded history of micro- or macroalbuminuria. For the macroalbuminuria group, at least 30 individuals were selected based on concurrent eGFR <60 ml min−1[1.73 m]−2. The study design is illustrated in electronic supplementary material (ESM) Fig. 1.
The control group was healthy volunteers by self-report, who were recruited by newspaper advertisement within the greater Copenhagen area. None of the healthy control participants took any prescribed medication at the time of study inclusion or had taken antibiotics 3 months prior to study initiation. Study participants were recruited to ensure equal distribution of sex and similar mean age in the four study groups. The study was conducted in accordance with the Declaration of Helsinki and approved by the Ethics Committee of the Danish Capital Region (protocol H-15018107). All participants gave written informed consent.
Bioclinical measures
Body weight (to nearest 0.05 kg) and height (to nearest 0.5 cm) of each participant were measured and used to calculate BMI (body weight/height2 [kg/m2]). Cholesterol was measured in plasma by an enzymatic method (Vitros 5600, Ortho Clinical Diagnostics, USA). High-sensitivity C-reactive protein (hsCRP) in serum was measured by a latex-enhanced turbidimetric immunoassay method on a Cobas 8000 modular c502 system. HbA1c was measured by HPLC calibrated against the IFCC standard. Plasma creatinine was measured by an enzymatic method (Vitros 5600, Ortho Clinical Diagnostics, USA) and eGFR was calculated using the Chronic Kidney Disease Epidemiology Collaboration (CKD-EPI) equation [11]. UACR was measured in three consecutive morning urine samples by an enzyme immunoassay and the geometric mean was calculated for each participant. Twenty-four hour blood pressure was recorded with a validated device programmed to measure blood pressure every 15 min between 07:00 hours and 22:00 hours and every 30 min between 22:00 hours and 07:00 hours (TM2430, Takeda, Japan) [12]. Detailed medical history was obtained, including current medication and history of diabetes complications, by interview and cross-referenced with electronic patient records. Participants completed a questionnaire on health and lifestyle, including information on smoking, alcohol consumption, defaecation habits and stool consistency on the Bristol stool scale [13]. Information on diet was collected based on a food frequency questionnaire (FFQ) used to obtain frequency and portion sizes of food and beverage consumption over the previous month and originally designed and validated for use in a larger population-based study (the Inter99 study) [14].
Faecal sampling, microbial DNA extraction, library preparation and 16S gene amplicon sequencing
A faecal sample was collected for each participant at home following a recommended standard protocol [15], with immediate freezing of samples at home at −18°C. No later than 72 h after collection, the frozen samples were transferred to the laboratory on dry ice and stored at −80°C until DNA extraction. A total of 210 samples were collected (one participant from the macroalbuminuria group failed to collect a stool sample). All samples contained sufficient material for DNA extraction, and DNA was extracted using the NucleoSpin Soil kit (Macherey-Nagle, Düren, Germany), according to the manufacturer’s instructions.
Details on methods used for library preparation and 16S gene amplicon sequencing are provided in the ESM Methods. Processing of the raw sequencing DNA data to amplicon sequence variants (ASVs) and quality control resulted in a total of 4616 unique ASVs which underwent taxonomic annotation and were merged at genus rank into a total of 324 genera. All annotated ASVs are listed in ESM Table 1.
Targeted ultra HPLC MS/MS analysis of selected plasma metabolites
A previously validated panel of 34 metabolites was used for targeted metabolomics analysis by mass spectrometry, as previously reported [16], except that in this study leucine was distinguished from isoleucine by difference in retention time. The selection of plasma metabolites was based on clinical relevance to diabetes and association with kidney function. A total of 31 metabolites passed the data quality checks, of which 23 were quantified absolutely and eight relatively. The 31 metabolites included in the analyses are presented in ESM Table 2. A more detailed description of the targeted metabolomics analysis is provided in ESM Methods.
Statistics
Descriptive analyses
Unpaired Student’s t test or ANOVA for continuous variables and χ2 test for categorical variables were applied for comparison between groups. For non-normally distributed variables, comparisons between groups were performed by Kruskal–Wallis test. Results with a two-tailed p value of <0.05 were considered significant.
Microbiome analyses
For alpha diversity the phyloseq R package was used to calculate richness, Shannon’s diversity index and Simpson’s diversity index based on rarefied ASV abundance [17]. The difference in alpha diversity between groups was tested using ANOVA. Community structure (beta diversity) was analysed using the vegan R package; multivariate homogeneity of group dispersions was assessed using the betadisper function, and difference in multidimensional centroid assessed by permutational analysis of variance (PERMANOVA) of Bray–Curtis dissimilarity and Jaccard distances implemented in the adonis R function.
To assess the differences between study groups at genus level, differential abundance analyses on unrarefied abundance tables were performed using a negative binomial Wald test as implemented in the DESeq2 R package [18]. Genera present in at least 10% of samples were analysed, leaving 210 of 325 for analyses. In addition, the differential abundance analyses were adjusted for medication, HbA1c and diabetes duration, adding these as covariates to the Wald test in DESeq2. A separate adjusted analysis was performed with diet as a covariate. Relevant medication was chosen to include use of RAAS-blocking treatment, other blood pressure-lowering agents, diuretics, aspirin, statins and proton pump inhibitors.
Adjustment for diet was performed using macronutrients and energy intake obtained from a FFQ completed by the participants [14]. To avoid multicollinearity between covariates, principal component (PC) analysis was performed by a singular value decomposition of the centred and scaled nutrient values. The first three PCs, all with eigenvalues >1 and capturing 95.2% of the variance combined, were selected as covariates. To avoid the biologically implausible assumption of constant fold change in relative abundance for each unit change of the independent variable, each PC was divided into quintiles and included as categorical covariates.
Differential relative abundance analyses on rarefied data included 204 of the 325 genera. For all differential abundance results, p values were corrected for multiple testing by applying the Benjamini–Hochberg procedure. A false discovery rate (FDR) of <0.1 was considered significant.
Metabolome analyses
Transformed and standardised metabolomics data were used for one-way ANOVA and correlation analyses, while transformed, standardised and imputed data were used for ANCOVA and for partial correlation analyses. A detailed description of the statistical analyses is given in ESM Methods. In brief, for ANOVA, correlation analyses, and partial correlation analyses with eGFR adjustment, data assumptions were checked prior to testing and then parametric or nonparametric tests were performed in accordance with met assumptions. For ANOVA, differences between means were tested among all individuals with type 1 diabetes and healthy control individuals, and, in addition, among the three albuminuria groups for the individuals with type 1 diabetes (normo-, micro- and macroalbuminuria). In each ANCOVA, differences in mean values for the metabolites between the three albuminuria groups were adjusted for medication, HbA1c and diabetes duration. Adjustment for diet was performed in a separate analysis. For all tests, p values were adjusted for multiple testing using Benjamini–Hochberg procedure. An FDR of <0.1 was considered significant.
All statistical analyses were performed using R (http://www.r-project.org/). Version 3.5.2 was used to analyse gut microbiota data while version 3.6.1 was used for metabolite data and for analyses of combined data.
Results
Clinical characteristics of the study sample
Our study included 211 participants (89 women) with an age of 60 ± 11 years (mean ± SD). Individuals with type 1 diabetes had a mean diabetes duration of 42 ± 15 years and an eGFR of 75 ± 25 ml min−1 (1.73 m)−2. The clinical characteristics of the study groups are presented in Table 1 and ESM Table 3. The individuals with type 1 diabetes and micro- or macroalbuminuria were more often treated with antihypertensive drugs, diuretics and proton pump inhibitors than the normoalbuminuric individuals with type 1 diabetes. Among the individuals with type 1 diabetes, 24 h systolic blood pressure, HbA1c and serum concentration of high-sensitivity C-reactive protein were, as expected, higher with increasing albuminuria grouping. The participants in the three albuminuria groups were comparable in the Bristol stool scale score and in an estimate of bowel movement frequency. The daily intake of macronutrients was significantly different between albuminuria groups in protein, fat and fibre (ESM Table 3). Overall, 44 individuals with diabetes had a concurrent eGFR of <60 ml min−1 [1.73 m]−2, of which 30 individuals were in the macroalbuminuria group (data not shown).
Gut microbiota diversity and community structure
At a rarefied sequencing depth of 90% of the lowest read count (17,714 reads), assessment of alpha diversity did not differ between the four study groups (ESM Fig. 2a–c; p ≥ 0.64). Analyses of Bray–Curtis dissimilarity showed a difference in community structure dispersion between the four groups (pomnibus = 0.008) (ESM Fig. 2d), with pairwise comparisons showing larger dispersion in individuals with type 1 diabetes with microalbuminuria (Tukey’s honest significance test p = 0.013) and macroalbuminuria (p = 0.017). Community membership dispersion, as assessed by Jaccard dissimilarity, was similar in the four study groups (pomnibus = 0.17, ESM Fig. 2e). We found, however, an overall difference in centroid position (PERMANOVA p = 0.002) between healthy controls and individuals with type 1 diabetes and microalbuminuria (p = 0.006) and macroalbuminuria (p = 0.006) when the groups were compared pairwise. Analyses of diversity and community structure and membership at ASV level showed similar results.
Taxonomic profiles of type 1 diabetes and diabetic nephropathy
We identified 79 genera that were differentially abundant in individuals with type 1 diabetes and healthy control individuals (Fig. 1 and ESM Table 4). Of these, Firmicutes was the most dominant phylum (60 out of 79). Seventy genera were enriched in individuals with type 1 diabetes compared with healthy control individuals, of which the majority pertained to the families Ruminococcaceae (22 genera) and Lachnospiraceae (11 genera). In contrast, nine genera were depleted in individuals with type 1 diabetes. The most frequently depleted ones were within the Lachnospiraceae family (four genera from this family, including one of which was genus Lachnospira) and the Ruminococcaceae family (two genera belonged to this family, one of which was genus Faecalibacterium).
ASVs at genus level that are differentially abundant in individuals with type 1 diabetes and healthy control individuals. Volcano plot of estimated log2-fold change in genera abundance between individuals with type 1 diabetes (n = 160) and healthy controls (n = 50). Q represents multiple testing adjusted p values (FDR) from negative binomial Wald tests implemented in the R package DESeq2. Prevalence indicates the percentage of participants in which a given genus is present. Abundance indicates mean relative abundance (ppm) of a given genus. Horizontal dotted line indicates an FDR of 0.1. Names of genera differentially abundant at FDR <0.001 are given at the genus level. T1D, type 1 diabetes
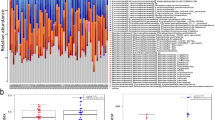
Among the individuals with type 1 diabetes, the abundance of ten genera, predominately from the phylum Firmicutes (nine out of ten), differed among the albuminuria groups (Fig. 2 and ESM Table 5). The relative abundance of genus Solobacterium from the Erysipelotrichaceae family was significantly enriched in the macroalbuminuria group compared with the microalbuminuria group, but significantly lower in the microalbuminuria group than in the normoalbuminuria group. In addition, several genera from the order Clostridiales were enriched in individuals with macroalbuminuria compared with the normoalbuminuria group, mainly from the Lachnospiraceae and Ruminococcaceae families (including ASVs assigned to genera [Eubacterium] fissicatena group and Anaerotruncus), and [Eubacterium] nodatum group from the family XI_2II. In individuals with type 1 diabetes with microalbuminuria, genus Merdibacter, UBA1819 and [Eubacterium] nodatum group were enriched compared with individuals with normoalbuminuria (the two latter also assigned to Clostridiales). Even though statistical significance was not reached, we found that several genera had a higher relative abundance with increasing albuminuria group (Fig. 2 and ESM Fig. 3).
ASVs at genus level associated with level of albuminuria in individuals with type 1 diabetes. Heatmap of associations (log2-fold change) between bacterial genera and level of albuminuria in individuals with type 1 diabetes, without and with adjustment for medication, HbA1c, diabetes duration and all three (medication, HbA1c and diabetes duration). FDR represents adjusted p values from negative binomial Wald test implemented in the R package DESeq2. Only genera with a significant association are shown. Results for all 210 genera are presented in ESM Table 5 (unadjusted) and ESM Table 6 (fully adjusted). The colour key indicates effect size as measured using Cohen’s d, with the histogram summarising frequencies of effects in the plot. Dur, diabetes duration; Macro, type 1 diabetes with macroalbuminuria; Med, medication; Micro, type 1 diabetes with microalbuminuria; Normo, type 1 diabetes with normoalbuminuria
The influence of potential confounding was assessed by adjustment for multiple medications (use of RAAS-blocking agents, other antihypertensive drugs, diuretics, aspirin, statins and proton pump inhibitors), HbA1c and diabetes duration (Fig. 2 and ESM Table 6). After adjustment for medication, 14 genera differed among the three albuminuria groups; when further adding HbA1c and diabetes duration in the model, 13 genera differed among the three albuminuria groups at an FDR of <0.1. Overall, the associations with albuminuria groups remained for some genera after adjustment (a genus from the order Selenomonadales, genus UBA1819 and the previous mentioned genera assigned to the Pasteurellaceae family), while other genera showed stronger significant associations. These included genus CHKC1002 and especially genus Prevotellaceae NK3B31 group and genus CAG-352, which were significantly higher in the macroalbuminuria group compared with the microalbuminuria group in all adjusted analyses. Other associations lost significance after adjustment, including [Eubacterium] fissicatena group and genus Tyzerella, however, the overall pattern remained similar with the different adjustments.
Adjustment for diet was added in a separate adjusted analysis, where 25 genera differed among the three albuminuria groups with an FDR of <0.1 (Fig. 3a and ESM Table 7). Information on diet was missing for seven individuals with type 1 diabetes, leaving n = 153 individuals for this analysis. Of the 25 genera, a genus from the order Selenomonadales, a genus assigned to the Pasteurellaceae family and genus Anaerotruncus had, as for the other unadjusted and adjusted analyses, significantly lower relative abundance in the macroalbuminuria group compared with the normoalbuminuria group. In addition, the abundance of the genus assigned to the order of Selenomonadales was, together with a genus of the family Corynebacteriaceae, lower in the macroalbuminuria group compared with the microalbuminuria group which was also observed in Fig. 2. As for the other adjusted analyses (Fig. 2), the genus Prevotellaceae NK3B31 was also significant when adjusting for diet with a higher relative abundance in the macroalbuminuric group compared with the microalbuminuric group.
ASVs at genus level and plasma metabolite levels associated with level of albuminuria in individuals with type 1 diabetes, adjusted for diet. (a) Associations (log2-fold change) between bacterial genera and level of albuminuria in individuals with type 1 diabetes with adjustment for diet. FDR represents adjusted p values from negative binomial Wald test implemented in the R package DESeq2. Only genera with a significant association are shown. Results for all 210 genera are presented in ESM Table 7. (b) Differences in plasma metabolites among the three groups of albuminuria tested with adjustment for diet. Only results from those with one or more test at FDR <0.1 are included. The colour key indicates effect size as measured using Cohen’s d with the histogram summarising frequencies of effects in the plot. Cit, l-citrulline; Crea, creatinine; HCit, homocitrulline; IndS, indoxyl sulphate; Kynu, l-kynurenine; Macro, type 1 diabetes with macroalbuminuria; Micro, type 1 diabetes with microalbuminuria; Normo, type 1 diabetes with normoalbuminuria; Trp, tryptophan
The abundance of 17 genera, predominately assigned to the Ruminococcaceae family, correlated inversely with eGFR, while five genera (three from the order Clostridiales and two from the order Bacteroidales) correlated positively with eGFR. Moreover, the genera Ruminiclostridium 1, Anaerostipes and Candidatus Soleaferrea were positively associated with UACR (ESM Fig. 4 and ESM Table 8).
Targeted metabolites profiles of type 1 diabetes and albuminuria
In total, 11 of the 31 measured plasma metabolites differed significantly in concentration between individuals with type 1 diabetes and healthy control individuals (Fig. 4). Dimethylarginine (asymmetric [ADMA] and symmetric [SDMA]), α- and β-hydroxybutyrate and N-methylnicotinamide were higher, and five amino acids, including tryptophan, were lower in individuals with type 1 diabetes compared with control individuals. Among the individuals with type 1 diabetes, differences between the three albuminuria groups were predominately seen between macro- and normoalbuminuria. Here, levels of indoxyl sulphate and l-citrulline were higher in the macroalbuminuria group compared with both the micro- and normoalbuminuria groups. Furthermore, plasma concentrations of homocitrulline and l-kynurenine were higher in individuals with macroalbuminuria compared with individuals with normoalbuminuria. The level of tryptophan was lower in macroalbuminuric compared with normoalbuminuric participants. We did not find any difference in the measured metabolites between individuals with micro- or normoalbuminuria.
Differences in plasma metabolite levels among the four study groups. Differences in plasma metabolite levels were tested between all individuals with type 1 diabetes and healthy control individuals, and among the albuminuria groups in type 1 diabetes. Additional adjustments for medication, HbA1c and diabetes duration were tested among the three albuminuria groups with diabetes. All 31 plasma metabolites were tested but only results from those with one or more test at FDR <0.1 are included. The colour key indicates effect size as measured using Cohen’s d with the histogram summarising frequencies of effects in the plot. Ala, alanine; a-OHB, α-hydroxybutyrate; b-OHB, β-hydroxybutyrate; Cit, l-citrulline; Crea, creatinine; DM, type 1 diabetes; HCit, homocitrulline; IndS, indoxyl sulphate; Kynu, l-kynurenine; Leu, leucine; Macro, type 1 diabetes with macroalbuminuria; Med, medication; Micro, type 1 diabetes with microalbuminuria; N-NMA, N-methylnicotinamide; Normo, type 1 diabetes with normoalbuminuria; Phe, phenylalanine; Trp, tryptophan
After adjustment for use of medication, HbA1c and diabetes duration, the differences between the three albuminuria groups remained significant for all metabolites, except for dimethylarginine and tryptophan (Fig. 4). For homocitrulline and indoxyl sulphate the associations were more pronounced (FDR <0.01), and the difference between the macro- and microalbuminuria groups became significant. Adjustment for diet showed similar results as for the other adjusted analyses (Fig. 3b).
Seven metabolites, including homocitrulline, ADMA and SDMA, indoxyl sulphate and l-kynurenine were associated with eGFR and six of these were also associated with UACR (ESM Fig. 5). There was a negative association between eGFR and the metabolites found to be significant, except for tryptophan, which correlated positively. Of the six metabolites significantly associated with UACR; five metabolites were positively associated and tryptophan was negatively associated.
To examine whether the gut microbiota taxonomy was associated with the measured plasma concentrations of metabolites, correlation analysis was performed between genera associated with albuminuria strata, eGFR or UACR, applying a threshold of FDR <0.1 (ESM Fig. 6) and, in addition, adjusting for eGFR (ESM Fig. 7). Circulating concentrations of tryptophan, leucine and homocitrulline were inversely associated with annotated bacteria at the genus level and cholic acid, homocitrulline, citrulline, ADMA and SDMA were positively associated with annotated bacteria. These associated bacteria included both ASVs assigned to the Ruminococaceae and the Lachnospiracae families. Correction for eGFR did not overall change the results, although some correlations were less pronounced (ESM Fig. 7). Partial correlations analyses including metabolites, bacterial taxa and renal variables in the same analyses showed similar results (ESM Fig. 8).
Discussion
In the present study including 161 individuals with type 1 diabetes and 50 healthy control individuals with equal sex and age distribution, we examined the composition and taxonomic profiles of the gut microbiota and a panel of measured plasma metabolites to identify potential associations of the gut microbiome and the plasma metabolites with type 1 diabetes without and with early renal complications. Compared with healthy control individuals, in participants with type 1 diabetes we observed differences in the gut microbiota composition and taxonomy as well as in levels of several circulating metabolites. Among individuals with type 1 diabetes, we found that relative abundance of certain gut bacteria genera and plasma concentrations of some metabolites were associated with the level of albuminuria, both in unadjusted and adjusted analyses.
Small studies in humans and rodents have examined the microbial composition in CKD at advanced stages [19]. Although different sequencing methods have been used and different bacterial taxonomic resolution levels have been achieved, alterations in the gut microbiota of individuals with ESKD have been observed compared with control individuals [4, 5, 19]. In one study, individuals with ESKD receiving haemodialysis had a gut microbiota composition that was distinct from that of healthy controls, characterised by significantly higher relative abundances of 190 microbial operational taxonomic units (OTUs) [4]. To examine the effect of renal failure per se, the gut microbiota in nephrectomised rats were examined and different bacterial taxa were present compared with control rats [4]. A study of 30 individuals with ESKD who were not on dialysis showed translocation of bacterial DNA to the blood, which was associated with low-grade inflammation, and the authors suggested that the finding might potentially have contributed to progression of CKD [5].
In our study, focusing on the initial stages of diabetic nephropathy with preserved renal function but increasing albuminuria between the study groups, we found that ten genera of gut bacteria differed significantly between individuals with type 1 diabetes stratified by levels of albuminuria. Differences remained after adjustment for several relevant potential covariates, with 13 significant differences in genera when adjusting for medication, HbA1c and diabetes duration and 25 genera with adjustment for diet. Compared with previous studies in more advanced CKD stages, our study only included 11 participants with an eGFR between 15–30 ml min−1 [1.73 m]−2 and none with ESKD. In addition, the included individuals with type 1 diabetes with micro- and macroalbuminuria were well-treated according to the relatively low measured levels of albuminuria, with median (IQR) micro- and macroalbuminuria levels of 1.36 mg/mmol (0.57–3.73 mg/mmol) and 19.66 mg/mmol (5.99–56.61 mg/mmol), respectively, which may be reflected in a less pronounced bacterial taxonomy difference among the albuminuria groups.
A general challenge in observational studies of the association between aberrant intestinal microbiota and a given pathology is to control for the impact of confounding factors, particularly diet. In the present study, the question is whether the altered gut microbiota is directly linked to diabetic nephropathy per se and not just altered due to the confounding or contributing effect of factors such as diet, medication, HbA1c or diabetes duration. Our adjusted results support the existence of significant associations between gut bacterial genera and the subgroups of albuminuria; associations that are not explained by these factors. Since the FFQ for diet information was developed for a population-based study [14], the diet data at an individual level in our relatively small study should be interpreted with caution. However, overall the diet adjusted analyses demonstrated more pronounced differences between the albuminuria groups compared with the unadjusted analyses (25 genera vs ten genera).
Whether features of the gut microbiota will eventually add to the value of albuminuria as a marker of diabetic nephropathy remain to be explored in future longitudinal studies with analyses of the intestinal microbiota at deep taxonomic and functional levels.
The colonic microbiota is involved in protein fermentation of amino acids, including tryptophan, which is broken down to indole and other derivates [20,21,22]. Indole is metabolised in the liver to generate indoxyl sulphate. Circulating concentrations of indoxyl sulphate correlate inversely with the level of kidney function [23] and an elevated circulating concentration of indoxyl sulphate is a risk marker for cardiovascular disease, progression of CKD and all-cause mortality [24, 25]. In animal studies, it has been demonstrated that indoxyl sulphate damages renal tubular cells and podocytes [26,27,28,29]. In line with these observations, we observed higher plasma concentrations of indoxyl sulphate in individuals with type 1 diabetes with macroalbuminuria compared with micro- and normoalbuminuria and an inverse correlation with eGFR.
An aberrant gut microbiota in individuals with CKD may contribute to higher uraemic toxin levels, which promotes progression of the disease. Uremic toxins produced by the gut microbiota increase intestinal epithelial inflammation, resulting in higher gut permeability. Accumulation of uraemic toxins in the circulation such as indoxyl sulphate and p-cresyl sulphate leads to systemic inflammation in capillaries, endothelial dysfunction and activation of the RASS, which are all common features of hypertension and CKD [30, 31]. In our study we found several genera, including Ruminiclostridium 1, a genus from the Ruminococcaceae family, and genus Anaerostipes from the Lachnospiraceae family inversely associated with the plasma concentration of tryptophan in individuals with type 1 diabetes. Other studies have demonstrated associations between circulating indoxyl sulphate and the gut microbiota [32,33,34,35]. A study by Barrios et al reported associations (both negative and positive) between plasma indoxyl sulphate and the relative abundance of three OTUs from the Ruminococcaceae and Lachnospiraceae families. These were from the genus Ruminococcus, Lachnospira or Oscillospira, respectively [33]. Interestingly, genus Oscillospira was significantly and positively associated with the albuminuria groups in our study when the analyses were performed with diet adjustment.
By the design of our study we can only identify associations. However, the interaction between the human gut microbiota and kidney disease does not seem unidirectional [31]. Individuals with ESKD accumulate uraemic toxins, which secondarily may lead to affection of the gut microbiota composition promoting uraemic toxins producing bacteria resulting in further imbalance of the gut microbiota and production of uraemic toxins. Triggering of such bidirectional interaction between the gut microbiota and CKD may require advanced stages of the disease or ESKD as the taxonomic associations with diabetic nephropathy disease groups (level of albuminuria) in our study were not as pronounced compared with studies in individuals with ESKD. Thus, the taxonomic and metabolite differences observed in the present study of initial stages of diabetic nephropathy may reflect an early phase of such an interaction. Future longitudinal and shotgun-based sequencing studies of the gut microbiota allowing for detailed functional microbiomics may facilitate an understanding of which gut bacterial taxa and microbial pathways are causing higher circulating indoxyl sulphate concentrations in people with CKD.
Our study has certain limitations. A limitation is the cross-sectional design, which does not allow for any conclusions on directionality of the identified microbiota and metabolite features nor causality. For the analyses of the gut microbiota, we applied the 16S RNA gene sequencing approach that allows determination of bacterial taxa at a modest resolution (genus) level. Therefore, the lack of deep profiling of the whole microbiome prohibited us from studying species and subspecies taxonomy and their functional potentials. The strengths of our study include the well-characterised study samples and the targeted quantification of measured plasma metabolites.
In conclusion, we identified aberrant profiles of the intestinal gut microbiota and plasma concentration of metabolites in individuals with type 1 diabetes of long duration. When stratifying individuals with type 1 diabetes by presence of normo-, micro- and macroalbuminuria reflecting absence or presence of initial stages of diabetic nephropathy, we identified additional differences in both the gut microbiota and the plasma metabolome.
Data availability
The datasets generated and/or analysed is available from the corresponding authors on reasonable requests.
Abbreviations
- ADMA:
-
Asymmetric dimethylarginine
- ASV:
-
Amplicon sequence variant
- CKD:
-
Chronic kidney disease
- ESKD:
-
End-stage kidney disease
- FDR:
-
False discovery rate
- FFQ:
-
Food frequency questionnaire
- OTU:
-
Operational taxonomic unit
- PC:
-
Principal component
- RAAS:
-
Renin–angiotensin–aldosterone system
- SDMA:
-
Symmetric dimethylarginine
- UACR:
-
Urine albumin/creatinine ratio
References
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci U S A 104(34):13780–13785. https://doi.org/10.1073/pnas.0706625104
Durazzo M, Ferro A, Gruden G (2019) Gastrointestinal microbiota and type 1 diabetes mellitus: The state of art. J Clin Med 8(11):1843. https://doi.org/10.3390/jcm8111843
Qin J, Li Y, Cai Z et al (2012) A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490(7418):55–60. https://doi.org/10.1038/nature11450
Vaziri ND, Wong J, Pahl M et al (2013) Chronic kidney disease alters intestinal microbial flora. Kidney Int 83(2):308–315. https://doi.org/10.1038/ki.2012.345
Wang F, Jiang H, Shi K, Ren Y, Zhang P, Cheng S (2012) Gut bacterial translocation is associated with microinflammation in end-stage renal disease patients. Nephrology (Carlton) 17(8):733–738. https://doi.org/10.1111/j.1440-1797.2012.01647.x
Arneth B, Arneth R, Shams M (2019) Metabolomics of type 1 and type 2 diabetes. Int J Mol Sci 20(10):2467. https://doi.org/10.3390/ijms20102467
Bervoets L, Massa G, Guedens W, Louis E, Noben JP, Adriaensens P (2017) Metabolic profiling of type 1 diabetes mellitus in children and adolescents: a case-control study. Diabetol Metab Syndr 9:48. https://doi.org/10.1186/s13098-017-0246-9
Jha V, Garcia-Garcia G, Iseki K et al (2013) Chronic kidney disease: global dimension and perspectives. Lancet 382(9888):260–272. https://doi.org/10.1016/s0140-6736(13)60687-x
Thomas MC, Brownlee M, Susztak K et al (2015) Diabetic kidney disease. Nat Rev Dis Primers 1:15018. https://doi.org/10.1038/nrdp.2015.18
Anders HJ, Andersen K, Stecher B (2013) The intestinal microbiota, a leaky gut, and abnormal immunity in kidney disease. Kidney Int 83(6):1010–1016. https://doi.org/10.1038/ki.2012.440
Levey AS, Stevens LA, Schmid CH et al (2009) A new equation to estimate glomerular filtration rate. Ann Intern Med 150(9):604–612. https://doi.org/10.7326/0003-4819-150-9-200905050-00006
Palatini P, Frigo G, Bertolo O, Roman E, Da Cortà R, Winnicki M (1998) Validation of the A&D TM-2430 device for ambulatory blood pressure monitoring and evaluation of performance according to subjects' characteristics. Blood Press Monit 3(4):255–260
Lewis SJ, Heaton KW (1997) Stool form scale as a useful guide to intestinal transit time. Scand J Gastroenterol 32(9):920–924. https://doi.org/10.3109/00365529709011203
Toft U, Kristoffersen L, Ladelund S et al (2008) Relative validity of a food frequency questionnaire used in the Inter99 study. Eur J Clin Nutr 62(8):1038–1046. https://doi.org/10.1038/sj.ejcn.1602815
Dore J, Ehrlich SD, Levene F et al (2015-04-14) IHMS_SOP 04 V1: Standard operating procedure for fecal samples frozen preserved self-collection, laboratory analysis handled within 24 hours to 7 days (24 hours ˂ x ≤ 7 days). International Human Microbiome Standards. Available from http://www.microbiome-standards.org/index.php?id=Sop&num=004. Accessed 10 Aug 2020
Ahonen L, Jantti S, Suvitaival T et al (2019) Targeted clinical metabolite profiling platform for the stratification of diabetic patients. Metabolites 9(9):184. https://doi.org/10.3390/metabo9090184
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8(4):e61217. https://doi.org/10.1371/journal.pone.0061217
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Yang T, Richards EM, Pepine CJ, Raizada MK (2018) The gut microbiota and the brain-gut-kidney axis in hypertension and chronic kidney disease. Nat Rev Nephrol 14(7):442–456. https://doi.org/10.1038/s41581-018-0018-2
Roager HM, Licht TR (2018) Microbial tryptophan catabolites in health and disease. Nat Commun 9(1):3294. https://doi.org/10.1038/s41467-018-05470-4
Aronov PA, Luo FJ, Plummer NS et al (2011) Colonic contribution to uremic solutes. J Am Soc Nephrol 22(9):1769–1776. https://doi.org/10.1681/asn.2010121220
Ramezani A, Raj DS (2014) The gut microbiome, kidney disease, and targeted interventions. J Am Soc Nephrol 25(4):657–670. https://doi.org/10.1681/asn.2013080905
Lin CJ, Chen HH, Pan CF et al (2011) p-Cresylsulfate and indoxyl sulfate level at different stages of chronic kidney disease. J Clin Lab Anal 25(3):191–197. https://doi.org/10.1002/jcla.20456
Barreto FC, Barreto DV, Liabeuf S et al (2009) Serum indoxyl sulfate is associated with vascular disease and mortality in chronic kidney disease patients. Clin J Am Soc Nephrol 4(10):1551–1558. https://doi.org/10.2215/cjn.03980609
Wu IW, Hsu KH, Lee CC et al (2011) p-Cresyl sulphate and indoxyl sulphate predict progression of chronic kidney disease. Nephrol Dial Transplant 26(3):938–947. https://doi.org/10.1093/ndt/gfq580
Satoh M, Hayashi H, Watanabe M et al (2003) Uremic toxins overload accelerates renal damage in a rat model of chronic renal failure. Nephron Exp Nephrol 95(3):e111–e118. https://doi.org/10.1159/000074327
Tumur Z, Shimizu H, Enomoto A, Miyazaki H, Niwa T (2010) Indoxyl sulfate upregulates expression of ICAM-1 and MCP-1 by oxidative stress-induced NF-kappaB activation. Am J Nephrol 31(5):435–441. https://doi.org/10.1159/000299798
Enomoto A, Takeda M, Tojo A et al (2002) Role of organic anion transporters in the tubular transport of indoxyl sulfate and the induction of its nephrotoxicity. J Am Soc Nephrol 13(7):1711–1720. https://doi.org/10.1097/01.asn.0000022017.96399.b2
Ichii O, Otsuka-Kanazawa S, Nakamura T et al (2014) Podocyte injury caused by indoxyl sulfate, a uremic toxin and aryl-hydrocarbon receptor ligand. PLoS One 9(9):e108448. https://doi.org/10.1371/journal.pone.0108448
Al Khodor S, Shatat IF (2017) Gut microbiome and kidney disease: a bidirectional relationship. Pediatr Nephrol 32(6):921–931. https://doi.org/10.1007/s00467-016-3392-7
Felizardo RJ, Castoldi A, Andrade-Oliveira V, Câmara NO (2016) The microbiota and chronic kidney diseases: a double-edged sword. Clin Transl Immunol 5(6):e86. https://doi.org/10.1038/cti.2016.36
Mishima E, Fukuda S, Mukawa C et al (2017) Evaluation of the impact of gut microbiota on uremic solute accumulation by a CE-TOFMS-based metabolomics approach. Kidney Int 92(3):634–645. https://doi.org/10.1016/j.kint.2017.02.011
Barrios C, Beaumont M, Pallister T et al (2015) Gut-microbiota-metabolite axis in early renal function decline. PLoS One 10(8):e0134311. https://doi.org/10.1371/journal.pone.0134311
Kikuchi M, Ueno M, Itoh Y, Suda W, Hattori M (2017) Uremic toxin-producing gut microbiota in rats with chronic kidney disease. nephron 135(1):51–60. https://doi.org/10.1159/000450619
Wong J, Piceno YM, DeSantis TZ, Pahl M, Andersen GL, Vaziri ND (2014) Expansion of urease- and uricase-containing, indole- and p-cresol-forming and contraction of short-chain fatty acid-producing intestinal microbiota in ESRD. Am J Nephrol 39(3):230–237. https://doi.org/10.1159/000360010
Acknowledgements
The authors would like to thank all participants. We thank T. Ragnholm Juhl, B. Ruud Jensen, J.Hermann, A. Lundgaard, L. Aabo, L. Jelstrup and N. Christiansen from Steno Diabetes Center Copenhagen for technical assistance. We also thank A. Forman, K. Kaadtmann for managerial assistance and P. Sandbeck for technical assistance (all from the Novo Nordisk Foundation Center for Basic Metabolic Research).
Authors’ relationships and activities
The authors declare no competing interest associated with this manuscript. Outside this study, PR has received research grants from AstraZeneca and Novo Nordisk. He has received lecture and/or consultancy fees (to his institution) from Astellas, AstraZeneca, Bayer, Boehringer Ingelheim, Eli Lilly, Gilead, Merck, Mundipharma, Novo Nordisk and Sanofi Aventis.
Funding
The project is funded by the Novo Nordisk Foundation (grant number NNF14OC0013659; ‘PROTON: PeRsOnalising Treatment Of diabetic Nephropathy’. SAW was funded by Innovation Fund Denmark (grant number 5016-00150B). The Novo Nordisk Foundation Center for Basic Metabolic Research is an independent Research Center at the University of Copenhagen and is partially funded by an unrestricted donation from the Novo Nordisk Foundation (www.cbmr.ku.dk) (grant number NNF18CC0034900). The funding sources were not involved in the design of the study; the collection, analysis, and interpretation of data; writing the report; and did not impose any restrictions regarding the publication of the report.
Author information
Authors and Affiliations
Contributions
SAW, TWH, MFM, TH, HHP, PR and OP conceived the study. Acquisition of data was performed by SAW and EHZ. SAW drafted the manuscript. SAW and PH performed the statistical analyses. JKV performed quality control of the 16S microbial data. JKV, THH and OP supervised the microbiota analyses. JKV, THH and TS contributed to the statistical analyses. CLQ supervised the metabolomics analyses. SAW, PH, JKV, THH, MFM, TWH, HHP, PR and OP interpreted the results and PH, JKV, THH, MFM, TWH, HHP, PR and OP critically revised the manuscript for important intellectual content. LA performed laboratory analyses of the metabolomics data. LA, TS, EHZ, TH and CLQ edited the manuscript and contributed to interpretation of the data. All authors have read and approved the final version of the manuscript. SAW, PR and OP are responsible for the integrity of the manuscript as a whole.
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM
(PDF 1.67 mb)
ESM Tables 1 and 4–8
(XLSX 0.98 mb)
Rights and permissions
About this article
Cite this article
Winther, S.A., Henriksen, P., Vogt, J.K. et al. Gut microbiota profile and selected plasma metabolites in type 1 diabetes without and with stratification by albuminuria. Diabetologia 63, 2713–2724 (2020). https://doi.org/10.1007/s00125-020-05260-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-020-05260-y