Abstract
Aims/hypothesis
An increase in PAI-1 activity is thought to be a key factor underlying myocardial infarction. Mouse Pai-1 (mPai-1) activity shows a daily rhythm in vivo, and its transcription seems to be controlled not only by clock genes but also by humoral factors such as insulin and triglycerides. Thus, we investigated daily clock genes and mPai-1 mRNA expression in the liver of db/db mice exhibiting high levels of glucose, insulin and triglycerides.
Methods
Locomotor activity was measured using an infrared detection system. RT-PCR or in situ hybridisation methods were applied to measure gene expression. Humoral factors were measured using measurement kits.
Results
The db/db mice showed attenuated locomotor activity rhythms. The rhythmic expression of mPer2 mRNA was severely diminished and the phase of mBmal1 oscillation was advanced in the db/db mouse liver, whereas mPai-1 mRNA was highly and constitutively expressed. Night-time restricted feeding led to a recovery not only from the diminished locomotor activity, but also from the diminished Per2 and advanced mBmal1 mRNA rhythms. Expression of mPai-1 mRNA in db/db mice was reduced to levels far below normal. Pioglitazone treatment slightly normalised glucose and insulin levels, with a slight reduction in mPai-1 gene expression.
Conclusions/interpretation
We demonstrated that Type 2 diabetes impairs the oscillation of the peripheral oscillator. Night-time restricted feeding rather than pioglitazone injection led to a recovery from the diminished locomotor activity, and altered oscillation of the peripheral clock and mPai-1 mRNA rhythm. Thus, we conclude that scheduled restricted food intake may be a useful form of treatment for diabetes.
Similar content being viewed by others
Introduction
Various physiological and behavioural phenomena are under the control of an endogenous circadian rhythm [1]. The molecular circadian clock system is thought to be based on transcriptional/translational feedback loops consisting of several ‘clock genes’ and their products [2]. For example, in mammals, CLOCK, a member of the basic helix-loop-helix-PAS domain that contains transcription factors, heterodimerises with BMAL1 and activates transcription of the Per1/2 and Cry1/2 genes. On the other hand, CRY1/2 proteins negatively regulate the E-box binding ability of CLOCK: BMAL1 [2]. Clock genes are expressed not only in the suprachiasmatic nucleus (SCN) of the hypothalamus where the central clock exists, but also in other brain regions and various peripheral tissues. Recent study shows that mammalian peripheral tissues contain an oscillator that couples to a central oscillator under normal conditions [3], and the clock genes in these peripheral tissues show clear circadian rhythms [4]. Plasminogen activator inhibitor type 1 (PAI-1), a primary regulator of the fibrinolytic cascade, has two functional E-box motifs in the promoter [5], and its gene expression is regulated by the circadian clock [6].
In vitro investigations have found that both glucose and insulin stimulate the production and subsequent release of PAI-1 in human vascular endothelial and smooth muscle cells [7, 8]. In addition, it was suggested that hyperglycaemia is responsible for stimulating transcription of the Pai-1 gene in vascular smooth muscle cells [9]. Hyperglycaemia, which results from the absolute or relative impairment of insulin secretion and/or insulin action is a characteristic of diabetes mellitus, of which there is Type 1 and Type 2 [10].
The db/db mouse is a recognised animal model of Type 2 diabetes patients. These mice, which lack functional leptin receptors [11, 12], become identifiably obese around 3 to 4 weeks of age and show symptoms of insulin resistance. Diabetes in db/db mice can be divided into two stages [13, 14]: the hyperinsulinaemic phase (phase I) and the hypoinsulinaemic phase (phase II) after the age of 2 to 3 months. Increased plasma insulin levels seen at the age of 10 days continue to increase until the age of 2 to 3 months (phase I), at which time there is an abrupt decrease followed by a return to normal levels. Blood glucose content is relatively normal or slightly elevated during phase I. However, after the age of 2 to 3 months (phase II), the blood glucose level markedly increases and remains at a high level until death.
Application of glucose to rat-1 cell culture reduces Per1 and Per2 gene expression and resets the circadian rhythm of clock gene expression [15]. In addition, acute administration of insulin increases Per1 gene expression in rat-1 cell culture [16]. These findings provided the impetus for our present investigation of clock gene and Pai-1 gene expression in the liver of Type 1 diabetic db/db mice, since these mice exhibit hyperglycaemia and hyperinsulinaemia.
Indeed, strategies that have been shown to prevent diabetes, such as diet and exercise or administration of thiazolidinediones, also decrease plasma PAI-1 concentrations [17, 18, 19]. Thus, in this study, we allowed diabetic db/db mice daily access to food for only a limited period of time and examined whether this restricted feeding or pioglitazone administration led to a recovery from abnormal clock gene and mPai-1 gene expression in the liver.
Materials and methods
Animals
Male C57BL/KsJ-db/db Jcl (db/db) and C57BL/KsJ-db/+m Jcl (db/+) mice were purchased from Clear Japan (Tokyo, Japan). Male ddY mice were obtained from Takasugi Experimental Animals (Saitama, Japan). Animals were maintained on a light–dark cycle (12 h light, 12 h dark, with lights on at 08.30 hours) at room temperature (23 °C), and were given free access to food and water except for during daily restricted feeding (RF) experiments. All animals were treated in accordance with the Principles of Laboratory Animal Care, and with the Law (no. 105) and Notification (no. 6) of the Japanese Government.
RNA isolation and RT-PCR
Mice (n=3–6 mice per time point) were deeply anaesthetised with ether and intracardially perfused with ice-cold saline. After perfusion, the liver was isolated, frozen in liquid nitrogen and stored at −80 °C until RNA isolation. Total RNA was extracted using Isogen reagent (Nippon Gene, Tokyo, Japan). After RNase-free DNase treatment, 100 ng of total RNA was reverse transcribed and amplified using the Superscript One-Step RT-PCR System (Invitrogen, Calif., USA) in a GeneAmp PCR System 9700 (Applied Biosystems, Calif., USA). A more detailed description of the specific primer pairs of clock (mPer1, mPer2 and mBmal1), mPai-1 and β-actin genes, and the semi-quantitative RT-PCR method used for measuring the expression level of mRNA can be found in Minami et al. [20]. Briefly, each target clock gene cDNA was co-amplified with β-actin cDNA in a single PCR tube. The PCR products were electrophoresed on a 3% agarose gel, stained with ethidium bromide and analysed by an EDAS-290 system (Kodak, N.Y., USA). The intensity of PCR product of the target gene was normalised to the intensity of β-actin. The amplitude (ratio of peak : trough) and phase determined by this method were reproducible and found to be comparable to those determined by Northern blot [21]. Thus, the present experimental conditions are considered capable of detecting a daily change of clock gene expression in the mouse liver.
In situ hybridisation
The quantity of mPer1 and mPer2 mRNA expression in the SCN was assessed by in situ hybridisation as previously described [22]. On the day of the experiment, mice were deeply anaesthetised by ether and were intracardially perfused with 0.1 mol/l phosphate buffer (PB; pH 7.4) containing 4% paraformaldehyde at the following Zeitgeber times (ZT; ZT 0 is defined as lights on, and ZT 12 as lights off): 0, 6, 12 and 18. Six and four mice were used for each time point in db/+ mice and db/db mice respectively. After animals were killed, brains were removed, post-fixed in 0.1 mol/l PB containing 4% paraformaldehyde for 24 h at 4 °C, and transferred into 20% sucrose in 0.1 mol/l PB for 72 h at 4 °C. Brain slices (40 µm thick) containing the temporal cortex and the SCN were made using cryostat (Microm, HM505E; Heidelberg, Germany) then placed in 2× standard saline citrate until processing for hybridisation. Slices were first treated with 1 µg/ml proteinase K in 10 mmol/l Tris-HCl buffer (pH 7.5) containing 10 mmol/l EDTA for 10 min at 37 °C, followed by 0.25% sodium dodecyl sulphate) containing 33P-labelled cRNA probes for 16 h at 60 °C. Radioisotope (α-[33P] UTP; PerkinElmer Life Sciences, Boston, Mass., USA)-labelled antisense cRNA probes were made from restriction-enzyme-linearised cDNA templates. After high-stringency post-hybridisation wash in 2× standard saline citrate/50% formamide, slices were treated with RNaseA (10 µg/ml) for 30 min at 37 °C. Slices were washed in 2× standard saline citrate/50% formamide and air-dried. Images were visualised by autoradiogram using BioMax MR film (Eastman Kodak, Rochester, N.Y., USA) and analysed with an image analysing system (MCID; Imaging Research, St. Catherines, Ont, Canada) after conversion into optical density by 14C-autoradiographic microscales (Amersham Pharmacia Biotech, Little Chalfont, UK).
We subtracted the intensity of the optical density in the corpus callosum from that in the SCN and regarded this value as the net intensity in the SCN. The intensity values of sections taken from rostral to caudal areas of the SCN (four sections per mouse) were then summed; the sum was considered to be a measure of the amount of clock gene data analysis. Four to six animals made up each ZT data point, and the values were expressed as means ± SEM.
RF experimentation
RF experimentation was performed as previously described [23] with the exception that food could only be accessed for 4 h during the night. In brief, after 1 day of fasting (termed day 0), mice were allowed access to food for 4 h from ZT 18 to ZT 22 for 6 consecutive days (day 1 to day 6) or 12 consecutive days (day 1 to day 12). In RF groups, animals were killed at ZT 0, 6, 12 and 18 on day 6 or day 12 under RF conditions. In groups with free access to food and water, animals were killed under these conditions.
Locomotor activity rhythm
Mice were housed individually in transparent plastic cages (31×20×13 cm) under light–dark (12 h, 12 h) conditions and their locomotor activity was measured using an area sensor (F5B; Omron, Tokyo, Japan) with an infrared detector. Locomotor activity was continuously recorded in 6-min epochs by personal computer. Chi square periodogram analysis [24] was applied to examine the periodicity ranging from 20- to 30-h periods. In general, Qp value calculated by chi square periodogram is related to the robustness of circadian rhythm [25]; a high Qp value indicates strong robustness of rhythm. Daily locomotor activity counts were also calculated.
Measurement of blood glucose, insulin and triglycerides
Immediately prior to liver isolation, 0.5 ml of blood was withdrawn from the mouse heart, and blood glucose was measured by Novo Assist Plus (Novo Nordic Pharma, Tokyo, Japan) according to manufacturer’s instructions. Insulin and triglycerides were measured in the plasma using the Insulin Measurement Kit (Morinaga, Tokyo, Japan) and Triglyceride E-test (Wako, Tokyo, Japan) according to manufacturer’s instructions.
Drugs
Streptozotocin (STZ) was purchased from Wako Chemical (Osaka, Japan). STZ was dissolved in 0.1 mol/l sodium citrate buffer (pH 5.2) and intraperitoneally injected (200 mg/kg) into 13-week-old ddY mice.
Pioglitazone (10 mg/kg, p.o.; Wako Chemical, Osaka, Japan) suspended in 0.5% carboxymethylcellulose was administered to db/db mice (10 weeks old) at ZT 12 for 12 days. On day 13, mice were killed at ZT 6 or ZT 18, with no drug administration. Control db/db and db/+ mice were administered with 0.5% carboxymethylcellulose at a dose of 10 ml/kg.
Statistical analysis
The values are expressed as means ± SEM. Rhythmicity of each group was tested by one-way ANOVA. The significance of differences between groups at each time point was determined by the Student’s t test. Differences were considered significant when the associated probability value was less than 0.05.
Results
Locomotor activity and daily expression of clock genes in the db/db mouse liver
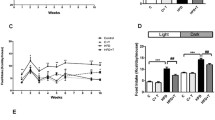
Using a semi-quantitative RT-PCR method, we examined clock (mPer1, mPer2, mBmal1) and mPai-1 gene expression at four time points in the liver of db/+ and db/db mice placed in light–dark conditions. We used both phase I (6–8 weeks old) and phase II (13–14 weeks old) db/db mice to study the effect of diabetic progress on clock gene mRNA expression. The db/+ mice showed clear daily locomotor activity rhythms under light–dark conditions (Fig. 1a). The mRNA levels of all clock genes examined showed a clear daily rhythm for both age groups of db/+ mice (Fig. 1c). One-way ANOVA revealed a significant rhythmicity in all clock gene expression in db/+ mice (phase I: mPer1, F [3, 20]=8.3, p<0.01; mPer2, F [3, 20]=12.9, p<0.01; mBmal1, F [3, 19]=8.4, p<0.01; phase II: mPer1, F [3, 19]=11.9, p<0.01; mPer2, F [3, 19]=12,218, p<0.01; mBmal1, F [3, 19]=22.9, p<0.01). Levels of mPer1 and mPer2 mRNA peaked during the night (ZT 12 and ZT 18 respectively), while mBmal1 mRNA levels peaked in the morning (ZT 0).
Daily locomotor activity rhythm and fluctuation of clock genes in the liver of db/+ and db/db mice. Left panels: phase I (6- to 8-week-old) mice; right panels: phase II (13- to 14-week-old) mice. a. Representative double-plotted actograms showing locomotor activity records of db/+ (upper) and db/db (lower) mice under light–dark (12 h, 12 h) conditions. The uppermost light and dark bars indicate light and dark periods respectively. Right panel shows result of chi square periodogram analysis. Horizontal axis shows the period and vertical axis shows the amplitude (Qp). The ascending straight line in the periodograms represents p<0.01. b. Representative electrophoresis photographs of PCR products from the db/+ and db/db mouse liver at ZT 0, 6, 12 and 18. c. Daily mRNA abundance of each clock (mPer1, mPer2, mBmal1) and mPai-1 gene was plotted as a relative mRNA level that was normalised to β-actin mRNA. The lowest value was set to 1. Each value represents the mean ± SEM (n=3–6 mice at each time point). **p<0.01, *p<0.05 vs db/+ at each time point (Student’s t test). Open circles, db/+; closed circles, db/db
The daily locomotor activity rhythm of phase I db/db mice was moderately attenuated (Fig. 1a). All phase I db/db mice (n=8) showed significant rhythmicity as evaluated by chi square periodogram analysis. All clock gene mRNA was expressed in a daily fashion under these experimental conditions (mPer1, F [3, 8]=7.7, p<0.01; mPer2, F [3, 8]=4.3, p<0.01; mBmal1, F [3, 8]=37.4, p<0.01, one-way ANOVA) in phase I db/db mice, but the peak time of mPer2 mRNA was advanced from ZT 18 to ZT 12. While mPer1 and mPer2 mRNA expression were low at ZT 6 in db/+ mice, moderately expressed mPer1 and mPer2 could be observed at ZT 6 in the phase I db/db mouse liver (Fig. 1b, c; mPer1: df=9, p<0.05; mPer2: df=11, p<0.01, Student’s t test vs db/+). On the other hand, mBmal1 expression in phase I db/db mice was similar to that in db/+ mice.
The daily locomotor activity rhythm of phase II db/db mice was severely diminished (Fig. 1a). About 75% of the phase II db/db mice (6/8) did not show significant rhythmicity as evaluated by chi square periodogram analysis. Daily counts of locomotor activity were significantly lower in phase II db/db mice (1824±659) than in db/+ mice (6078±1190) (df=8, p<0.01, Student’s t test). The rhythmicity of mPer2 mRNA was attenuated (F [3, 8]=2.2, p=0.17, one-way ANOVA). There was elevated mPer1 mRNA expression (Fig. 1, right column; ZT 0: df=7, p<0.05; ZT 6: df=8, p<0.01; ZT 12: df=8, p<0.01, Student’s t test vs db/+). However, the rhythmicity of mPer1 and mBmal1 mRNA expression was still significant (mPer1: F [3, 8]=12.8, p<0.01; mBmal1, F [3, 8]=20.7, p<0.01, one-way ANOVA). The mBmal1 mRNA rhythm was diminished and advanced in phase II db/db mice but not in db/+ mice (Fig. 1b, c).
Daily expression of mPai-1 gene in the db/db mouse liver
In the liver of phase I db/+ mice, mPai-1 expression was very weak at all sampling times (Fig. 1c), and not significant (F [3, 8]=2.3, p=0.14, one-way ANOVA), whereas phase I db/db mice exhibited a clear daily rhythm of mPai-1 mRNA with a peak at ZT 18 (Fig. 1c; F [3, 8]=15.0, p<0.01, one-way ANOVA). Phase II db/+ mice exhibited weakly expressed mPai-1 during the day (F [3, 8]=17.9, p<0.01, one-way ANOVA), whereas db/db mice exhibited strongly expressed mPai-1 throughout the entire day with a peak at ZT 12 (Fig. 1c; F [3, 8]=3.7, p=0.06, one-way ANOVA).
Daily expression of mPer genes in the SCN of db/db mice
We examined mPer1 and mPer2 expression in the SCN of phase II db/db mice at four time points. In contrast to mPer mRNA expression in the liver, mPer1 mRNA expression in the SCN was not affected (Fig. 2; F [1, 26]=1.5, p=0.22, two-way ANOVA) and mPer2 mRNA expression in the SCN was slightly affected (Fig. 2; F [1, 30]=4.9, p<0.05, two-way ANOVA).
Daily mPer1 and mPer2 mRNA expression in the suprachiasmatic nucleus (SCN) of db/+ and phase II db/db mice. Upper: representative autoradiograms showing mPer1 (a) and mPer2 (b) mRNA of db/+ (upper) and db/db (lower) mice at each ZT. Lower: expression of mPer1 (a) and mPer2 (b) mRNA levels of each genotype (13–14 weeks of age) at ZT 0, 6, 12 and 18. Each value represents the mean ± SEM (n=4–6 mice per time point). Open circles, db/+; closed circles, db/db
Locomotor activity and daily expression of clock genes in the STZ-induced diabetic mouse liver
The phases of clock and clock-controlled gene mRNA rhythms were previously shown to have advanced for approximately 3 hours in the STZ-induced diabetic rat heart [26]. We examined locomotor activity rhythm and clock gene mRNA expression in the liver of STZ-injected mice. The blood glucose level of STZ-injected mice at ZT 12, 5 days after injection (29.4±2.4 mmol/l), was significantly higher than that of control mice (7.6±0.69 mmol/l; p<0.01, Student’s t test) and comparable to that of phase II db/db mice (24.2±1.3 mmol/l). The averaged daily activity counts were slightly reduced from 9488±891 to 7309±2062 by STZ injection (df=9, p=0.36, Student’s t test; Fig. 3a). The vehicle injection had no effect on locomotor activity counts (11 469±649 for before injection, 10 956±1964 for after injection). Although the robustness indicated by Qp values was also slightly reduced by STZ injection, 24-h rhythmicity was maintained in all STZ-treated mice.
Effect of streptozotocin (STZ) on locomotor activity, and clock and mPai-1 gene expression. a. Representative double-plotted actograms showing daily locomotor activity before and after vehicle (upper) or STZ (lower) injection (arrows). The uppermost light and dark bars indicate light and dark periods respectively. Right panel shows result of chi square periodogram analysis. Horizontal axis shows the period and vertical axis shows the amplitude (Qp). The ascending straight line in the periodograms represents p<0.01. b. Representative electrophoresis photographs of PCR products from 5 days after vehicle injection (ctrl) and STZ injection (STZ) at ZT 0, 6, 12 and 18. c. Daily mRNA abundance of each clock and mPai-1 gene from control (ctrl) and STZ-treated mice was plotted as a relative mRNA level that was normalised to β-actin mRNA. The lowest value was set to 1. Each value represents the mean ± SEM (n=3 mice per time point). **p<0.01, *p<0.05 vs control (Student’s t test). Open circles, control; closed circles, STZ
In STZ-treated mice, the daily rhythm of mPer2 mRNA expression was diminished and the peak of the mBmal1 mRNA rhythm advanced from ZT 0 to ZT 18 (Fig. 3). The peak of the mPer1 mRNA rhythm was not affected by STZ treatment. However, as seen in db/db mice, moderate mPer1 and mPer2 mRNA expression was observed at ZT 6 in the STZ-injected mouse liver (Fig. 3b, c; mPer1: df=8, p<0.01; mPer2: df=8, p<0.01, Student’s t test vs control). The expression of mPai-1 in both control and STZ-treated mice was very low at all ZT points, and there was no significant difference between the two groups (F [1, 16]=0.07, p=0.78, two-way ANOVA).
RF-induced recovery from abnormal expression of clock and mPai-1 genes in the db/db mouse liver
Phase II db/+ and db/db mice were placed on a night-time RF (ZT 18–22) schedule for 6 or 12 days under light–dark conditions. As shown in Figure 4a, db/+ mice placed under RF for 12 days maintained clear daily locomotor activity rhythms similar to those seen before RF. The db/db mice, which normally have no or a severely diminished locomotor activity rhythm when given free access to food and water, showed an RF-dependent, clear daily locomotor activity rhythm under RF conditions (Fig. 4a). Chi square periodogram analysis demonstrated the recovery of significant rhythmicity. Daily counts of locomotor activity were also increased by RF treatment (1.7 times in db/+ mice and 4.5 times in db/db mice). Thus, there was no significant difference in locomotor activity count between the two groups during the RF period (df=8, p=0.36, Student’s t test).
Effect of night-time restricted feeding (RF) on daily clock and mPai-1 gene expression in the liver of db/+ (left) and phase II db/db (right) mice. a. Representative double-plotted actograms showing daily locomotor activity before and after RF (dotted box). The uppermost light and dark bars indicate light and dark periods respectively. Lower panel shows result of chi square periodogram analysis. Horizontal axis shows the period and vertical axis shows the amplitude (Qp). The ascending straight line in the periodograms represents p<0.01. b. Representative electrophoresis photographs of PCR products from free access (ad lib) and RF groups of db/+ and db/db mice at ZT 0, 6, 12 and 18 on day 6 (RF 6d) or day 12 (RF 12d). c. Expression of clock and Pai-1 genes from the db/+ and db/db mouse liver at each ZT was plotted as a relative mRNA level that was normalised to β-actin expression. The lowest value was set to 1. Each value represents the mean ± SEM (n=3–4 mice per time point). **p<0.01, *p<0.05 (RF 6d vs ad lib), ##p<0.01, #p<0.05 (RF 12d vs ad lib) (Student’s t test). Open circles, ad lib; closed circles, RF 6d; closed triangles, RF 12d
When given free access to food and water, db/+ mice showed a clear daily expression of clock gene mRNA (Fig. 4). Although 6 days of RF did not affect clock gene expression in db/+ mice, 12 days of RF reduced the amplitude of the rhythm of mPer2 in these mice. When db/db mice were shifted from free access feeding conditions to RF for 6 or 12 days, mPer2 mRNA rhythmicity changed from severely diminished to robust rhythmic expression in the liver (Fig. 4). In particular, the mPer2 expression at ZT 6 was reduced to the level seen in db/+ mice. Furthermore, the peak expression of mBmal1 mRNA returned to ZT 0 from ZT 18, and the amplitude of the rhythm also returned to the amplitude seen in db/+ mice. Although mPai-1 mRNA expression was low in db/+ mice with free access to food and water, RF treatment lowered the levels of mPai-1 mRNA in a manner dependent on the RF treatment period (6 or 12 days). Under free access feeding conditions, db/db mice exhibited arrhythmic (F [3, 8]=1.06, p=0.41, one-way ANOVA) high expression of mPai-1 in the liver. RF treatment for 6 days lowered the mPai-1 expression in db/db mice, and RF for 12 days eliminated the mPai-1 mRNA expression (Fig. 4c). During RF, the phase of the mRNA expression rhythm of all clock genes in db/db mice started to resemble that of control (db/+) mice (Fig. 4).
We examined the blood insulin, glucose and triglyceride levels before and after RF treatment in db/db mice. RF for 12 days lowered the insulin level at ZT 6 (df=5, p<0.05, Student’s t test), blood glucose level at all sampling points (ZT 0: df=5, p<0.05; ZT 6: df=5, p<0.05; ZT 12: df=5, p<0.01; ZT 18: df=5, p<0.01, Student’s t test), and triglyceride level (ZT 0: df=8, p<0.05; ZT 6: df=8, p<0.01; ZT 12: df=8, p<0.01; ZT 18: df=8, p<0.01, Student’s t test) of db/db mice, and RF for 12 days, lowered the insulin level at ZT 18 (df=5, p<0.01, Student’s t test), blood glucose level (ZT 0: df=5, p<0.01; ZT 6: df=5, p<0.01; ZT 18: df=5, p<0.05, Student’s t test), and triglyceride level (ZT 0: df=5, p<0.05; ZT 12: df=5, p<0.01, Student’s t test) in db/+ mice (Fig. 5).
Effect of night-time restricted feeding (RF) for 12 days on daily pattern of blood glucose level (a), insulin level (b), and triglycerides level (c) in phase II db/+ (upper) and db/db (lower) mice. Each value represents the mean ± SEM (n=3–4). **p<0.01, *p<0.05 (ad lib vs RF 12d) (Student’s t test). Open circles, ad lib; closed circles, RF 12d
RF for 12 days in db/+ mice led to reduced food intake (Fig. 6; df=8, p<0.01, Student’s t test). However, RF caused no significant effect on body weight in db/+ mice (day 6: df=8, p=0.12; day 12: df=8, p=0.18, Student’s t test). RF treatment significantly reduced food intake (day 6: df=11, p<0.05; day 12: df=11, p<0.01, Student’s t test) and body weight (day 6: df=11, p<0.05; day 12: df=11, p<0.01, Student’s t test) in db/db mice depending on the duration of RF (6 or 12 days).
Effect of night-time restricted feeding (RF) for 12 days on food intake (a) and body weight (b) of db/db mice. Each value represents the mean ± SEM (n=12–24). **p<0.01, *p<0.05 (ad lib vs RF 6 or RF 12) (Student’s t test). Open circles, db/db ad lib; closed circles, db/db RF; open triangles, db/+ ad lib; closed triangles, db/+ RF
Alterations in daily clock genes expressed in the livers isolated from db/db mice were normalised by RF (Fig. 4). Therefore, alterations of daily clock genes observed in livers isolated from STZ-treated mice were examined under RF conditions for 10 days. Interestingly, RF treatment recovered a night (ZT 18)–day (ZT 6) difference of mPer2 gene expression in STZ-treated mice (15.6 times higher expression for RF, 1.3 times higher with free access feeding, 11 times higher for non-STZ-treated group).
Pioglitazone-induced recovery from abnormal expression of clock and mPai-1 genes in the db/db mouse liver
Treatment of db/db mice with pioglitazone may normalise mPai-1 gene expression through normalisation of humoral influences such as glucose and insulin in the liver without affecting peripheral clock gene expression, as pioglitazone treatment has been reported to decrease the plasma glucose and insulin levels in db/db mice [27]. Daily administration of pioglitazone slightly lowered the mPai-1 gene expression at both ZT 6 and ZT 18 (Fig. 7), and increased mPer2 gene expression, especially at ZT 6 (Fig. 7). Pioglitazone slightly lowered the plasma glucose and insulin level (Fig. 7) and significantly increased the body weight, especially at ZT 18 (df=7, p<0.01, Student’s t test).
Effect of pioglitazone (10 mg/kg) administration for 12 days on body weight (a), blood glucose level (b), insulin level (c) and mPai-1 and mPer2 gene expression (d–f). On day 13, mice were killed at ZT 6 or ZT 18 under withdrawal of drug. f. Representative electrophoresis photographs of PCR products from the db/+ and db/db mouse liver at ZT 6 and 18. d, e. mRNA abundance of mPer2 and mPai-1 gene was plotted as a relative mRNA level that was normalised to β-actin mRNA. The lowest value was set to 1. Each value represents the mean ± SEM (n=3–4 mice at each time point). **p<0.01, *p<0.05 (Student’s t test)
Discussion
The initial hypothesis of the present study appears to be that altered humoral factors in Type 2 diabetes will alter peripheral circadian clocks, which in turn will result in abnormal mPai-1 expression. In the present experiment, we used young (phase I) and middle-aged (phase II) db/db mice as an animal model to study the effects of Type 2 diabetes on clock genes and the clock-controlled gene expression of mPai-1. We found that the daily oscillation of mPer2 gene expression was severely attenuated (or blunted in some cases) and that the peak time of mPer2 mRNA was advanced from ZT 18 to ZT 12 in the liver of phase I and phase II db/db mice. In addition, the peak time of mBmal1 gene mRNA oscillation had also advanced from ZT 0 to ZT 18. While the peak time of mPer1 was not altered, mPer1 mRNA was highly expressed at ZT 6 in both phase I and phase II db/db mice. These results suggest that oscillation of the peripheral oscillator is advanced and attenuated specifically in phase II db/db mice.
The phase of the clock gene mRNA oscillation rhythm advances by about 3 h in the STZ-treated rat heart [26]. Similar results were obtained from our present study of the STZ-treated liver of mice. STZ disrupts pancreatic beta cells and induces Type 1 diabetes [28]. Our results suggest that Type 2 diabetes also induces phase advances of clock gene mRNA in the peripheral tissues. Although the amplitudes of clock gene oscillation were reportedly not altered in the STZ-induced diabetic rat heart [26], we observed a diminished oscillation of mPer2 mRNA in both the STZ-injected and db/db mouse liver. The different species and/or tissues used in these studies may explain this discrepancy. Furthermore, we injected higher doses of STZ than Young et al. [26] (200 mg/kg vs 65 mg/kg respectively), which may have caused more severe diabetes in the mice used in this study, resulting in diminished mPer2 mRNA oscillation.
Expression of mPai-1 mRNA in the liver of young wild (data not shown) and db/+ mice was very low. The db/+ mice at 13 to 14 weeks old exhibited weak mPai-1 expression that peaked at night, and older mice (6 months to 1 year old) also exhibited weak mPai-1 expression in the liver (data not shown). Comparatively, mPai-1 mRNA in the 13- to 14-week-old db/+ mice was higher than that in the 6- to 8-week-old db/+ mice, possibly due to an ageing effect. Interestingly, 6- to 8-week-old db/db mice strongly exhibited expression of mPai-1 in the liver at ZT 18. This may be explained by the hypothesis that some factors [29] possessing promoter activity on mPai-1 gene expression are increased at this age in db/db mice under the control of clock regulation. The constant high level of mPai-1 mRNA expression observed in 13- to 14-week-old db/db mice may be related to the increased factors [29] for elevation of mPai-1 expression, and also to the reduced rhythmicity of clock genes such as mPer1, mPer2, mBmal1 and Bmal2 for arrhythmicity of mPai-1 expression. An increase in blood glucose, insulin and triglyceride levels produces Pai-1 mRNA expression in the liver [30], and insulin and triglycerides synergistically induce Pai-1 biosynthesis in hepatocytes [30]. Previously published data suggest that the CLOCK:BMAL2 (or BMAL1) heterodimer may be responsible for regulating mPai-1 circadian expression [6]. All of these factors together may cause a constant high level of mPai-1 gene expression in the liver of db/db mice.
Thus, an elevation of blood insulin and triglycerides may strongly induce mPai-1 mRNA expression in the liver of db/db mice, but not in STZ-treated mice. Surprisingly, we found that mPai-1 mRNA was not elevated in STZ-treated mice. In fact, STZ-treated mice exhibited an elevation of glucose, but not insulin or triglycerides. Such an observation might suggest that alterations in the circadian clock in the db/db and STZ-treated mice are independent of the changes in mPai-1 expression.
Clock gene mRNA oscillation in the SCN did not differ between db/db and control (db/+) mice. This result leads us to believe that the central oscillator in the SCN is not altered in Type 1 diabetic db/db mice and that the diabetes-induced advanced and attenuated oscillation of the peripheral clock does not result from change in the central SCN clock oscillation. Phase II db/db mice showed an extremely attenuated locomotor activity rhythm under free access conditions. As clock gene oscillation in the SCN was normal in phase II db/db mice, attenuation and arrhythmicity of the locomotor activity and clock gene mRNA rhythm in the liver did not result from a diminishing of the core SCN oscillator but rather from abnormalities stemming from the output mechanism mediating locomotor activity and peripheral clock resetting. Similar results were reported using old animals, in which clock gene expression in the SCN was maintained but locomotor activity was severely diminished [31]. Thus, obesity may be responsible for the reduction of locomotor activity, and hyperphagia may be responsible for the arrhythmicity of locomotor activity in db/db mice.
A circadian rhythm in obese Zucker (fa/fa) rats [32, 33] that have a functionally defective leptin receptor [12, 34] showed significant deficits in both the daily mean and circadian amplitude of body temperature and activity, but both rhythms persisted irrespectively of lighting conditions with a period similar to that of lean control mice [32]. The db/db mice, on the other hand, had an extremely attenuated locomotor activity rhythm, and about 75% of phase II db/db mice lost rhythmicity. Although both fa/fa rats and db/db mice have functionally deficient leptin receptors [11, 12], unlike db/db mice, fa/fa rats become severely hyperglycaemic because of their different genetic background [33]. Obese fa/fa rats are relatively normoglycaemic or only slightly hyperglycaemic [35, 36, 37]. Thus, the severe circadian deficits seen in db/db mice may be related to their more severe diabetic symptoms in comparison with fa/fa rats.
Our next aim was to elucidate the recovery effect of RF on the abnormal expression of clock and mPai-1 genes in db/db and STZ-treated mice. To do this, we conducted RF at night when animals normally eat chow. Night-time RF not only led to the reappearance of locomotor activity, but also to the recovery of the robust clock gene mRNA oscillation in db/db mice. We demonstrated that the advanced expression of mBmal1 mRNA under free access conditions was normalised under night-time RF conditions in db/db mice. A diminished oscillation of mPer2 and mBmal1 was also recovered with night-time RF. Interestingly, RF treatment reduced the mPai-1 mRNA expression in the liver of db/db as well as db/+ mice. Indeed, RF treatment lowered the plasma glucose, insulin and triglyceride levels in db/db mice, and also glucose and triglycerides in db/+ mice. It has been revealed that RF is one of the strong entraining factors of peripheral circadian clocks not only in normal but also in clock/clock [20, 38] and STZ-treated mice (present result), as well as a modulator of circulating factors such as glucose, insulin and triglycerides. RF treatment reduced the glucose, insulin and triglyceride levels in db/db mice. However, these levels are still high in comparison with those in db/+ mice. Thus, changing the level of circulation factors may not be directly responsible for expression of mPai-1, because RF reduced mPai-1 expression in the db/db mice to levels far below normal. We should consider the role of other humoral factors such as angiotensin II, NEFA and glucocorticoids [29] in the regulation of Pai-1 gene expression.
RF increased the activity counts and reproduced 24-h rhythmicity in the locomotor activity of db/db mice. Therefore, we could not rule out the possibility that the RF produced recovery of gene expression through “exercise” or through food-related signals in db/db mice. Further experimentation is required for dissociation of these two factors.
As pioglitazone was reported to reduce plasma glucose and insulin levels under fasting by increasing the body weights [27], this drug was expected to normalise the mPai-1 expression in db/db mice without affecting clock gene expression. Similarly to in a previous paper [27], pioglitazone slightly reduced glucose and insulin levels as well as mPai-1 expression. When pioglitazone was co-applied with insulin, pioglitazone strongly produced the expression of Pai-1 gene in HepG2 cells [39] and 3T3-L1 cells [40]. Such a facilitatory effect of pioglitazone might cancel out the reduction of mPai-1 gene caused by lowering glucose and insulin in pioglitazone-treated db/db mice. Abnormal expression of mPer2 in the liver of db/db mice was facilitated by the treatment with pioglitazone. Although at present we do not know the mechanism of the augmentation of mPer2, the increase in body weight caused by pioglitazone at ZT 6 may facilitate the mPer2 gene expression. Indeed, food intake increased the mPer2 gene expression in the liver of normal fasted mice (unpublished observations).
Thus, RF treatment rather than pioglitazone administration leads to not only the successful recovery of peripheral clock gene oscillation, but also to the normalisation of Pai-1 gene expression, which is evidence for the usefulness of RF treatment for diabetes mellitus patients.
In summary, we demonstrated impaired clock gene mRNA rhythm in the db/db mouse liver. A restricted feeding schedule was sufficient to recover peripheral clock gene expression and also reduce mPai-1 gene expression in diabetic mice. Thus, scheduled restricted food intake may be a useful form of treatment for diabetes mellitus patients.
Abbreviations
- PB:
-
phosphate buffer
- RF:
-
restricted feeding
- SCN:
-
suprachiasmatic nucleus
- STZ:
-
streptozotocin
- ZT:
-
Zeitgeber time
References
Takahashi JS, Turek FW, Moore RY (2001) Human circadian rhythms. In: Takahashi JS (ed) Handbook of behavioural neurobiology, vol 12. Kluwer Academic/Plenum Publishers, pp 531–744
Reppert SM, Weaver DR (2002) Coordination of circadian timing in mammals. Nature 418:935–941
Damiola F, Le Minh N, Preitner N, Kornamann B, Fleury-Olela F, Schbler U (2000) Restricted feeding uncouples circadian oscillators in peripheral tissues from the central pacemaker in the suprachiasmatic nucleus. Genes Dev 14:2950–2961
Young ME, Razeghi P, Taegtmeyer H (2001) Clock genes in the heart: characterization and attenuation with hypertrophy. Circ Res 88:1142–1150
Hua X, Miller ZA, Wu G, Shi Y, Lodish HF (1999) Specificity in transforming growth factor β-induced transcription of the plasminogen activator inhibitor-1 gene. Proc Natl Acad Sci USA 96:13130–13135
Maemura K, de la Monte SM, Chin MT et al. (2000) CLIF, a novel cycle-like factor, regulates the circadian oscillation of plasminogen activator inhibitor-1 gene expression. J Biol Chem 275:36847–36851
Pandolf A, Iacoviello L, Capani F, Vitacolonna E, Donati MB, Consoli A (1996) Glucose and insulin independently reduce the fibrinolytic potential of human vascular smooth cells in culture. Diabetologia 39:1425–1431
Maiello M, Boeri D, Podesta F et al. (1992) Increased expression of tissue plasminogen activator and its inhibitor and reduced fibrinolytic potential of human endothelial cells cultured in elevated glucose. Diabetes 41:1009–1015
Chen YQ, Su M, Walia RR, Hao Q, Covington JW, Vaughan DE (1998) Sp1 sites mediate activation of the plasminogen activator inhibitor-1 promoter by glucose in vascular smooth muscle cells. J Biol Chem 273:8225–8231
Moller DE (2001) New drug targets for type 2 diabetes and the metabolic syndrome. Nature 414:821–827
Chen H, Charlat O, Tartaglia LA et al. (1996) Evidence that the diabetes gene encodes the leptin receptor: identification of a mutation in the leptin receptor gene in db/db mice. Cell 84:491–495
Chua SC Jr, Chung WK, Wu-Peng XS et al. (1996) Phenotypes of mouse diabetes and rat fatty due to mutations in the OB (leptin) receptor. Science 271:994–996
Hummel KP, Dickie MM, Coleman DL (1966) Diabetes, a new mutation in the mouse. Science 153:1127–1128
Herberg L, Coleman DL (1977) Laboratory animals exhibiting obesity and diabetes syndromes. Metabolism 26:59–99
Hirota T, Okano T, Kokame K, Shirotani-Ikejima H, Miyata T, Fukada Y (2002) Glucose down regulates Per1 and Per2 mRNA levels and induces circadian gene expression in cultured Rat-1 fibroblasts. J Biol Chem 277:44244–44251
Balsalobre A, Marcacci L, Schibler U (2000) Multiple signaling pathways elicit circadian expression in cultured Rat-1 fibroblasts. Curr Biol 10:1291–1294
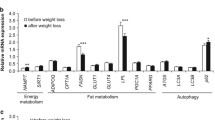
Folsom AR, Qamhieh HT, Wing RR et al. (1993) Impact of weight loss on plasminogen activator inhibitor (PAI-1), factor VII, and other haemostatic factors in moderately overweight adults. Arterioscler Thromb 13:162–169
Gris JC, Schved JF, Feugeas O et al. (1990) Impact of smoking, physical training and weight reduction on FVII, PAI-1 and haemostatic markers in sedentary men. Thromb Haemost 64:516–520
Fonseca VA, Reynolds T, Hemphill D et al. (1998) Effect of troglitazone on fibrinolysis and activated coagulation in patients with non-insulin-dependent diabetes mellitus. J Diabetes Complications 12:181–186
Minami Y, Horikawa K, Akiyama M, Shibata S (2002) Restricted feeding induces daily expression of clock genes and Pai-1 mRNA in the heart of Clock mutant mice. FEBS Lett 526:115–118
Sakamoto K, Ishida N (2000) Light-induced phase-shifts in the circadian expression rhythm of mammalian period genes in the mouse heart. Eur J Neurosci 12:4003–4006
Minami Y, Furuno K, Akiyama M, Moriya T, Shibata S (2002) Pituitary adenylate cyclase-activating polypeptide produces a phase shift associated with induction of mPer expression in the mouse suprachiasmatic nucleus. Neuroscience 113:37–45
Wakamatsu H, Yoshinobu Y, Aida R, Moriya T, Akiyama M, Shibata S (2001) Restricted feeding-induced anticipatory activity rhythm is associated with a phase-shift of the expression of mPer1 and mPer2 mRNA in the cerebral cortex and hippocampus but not in the suprachiasmatic nucleus of mice. Eur J Neurosci 13:1190–1196
Sokolove PG, Bushell WN (1978) The chi square periodogram: its utility for analysis of circadian rhythms. J Theor Biol 72:131–160
Refinetti R (1996) Comparison of the body temperature rhythms of diurnal and nocturnal rodents. J Exp Zool 275:67–70
Young ME, Wilson CR, Razeghi P, Guthrie PH, Taegtmeyer H (2002) Alterations of the circadian clock in the heart by streptozotocin-induced diabetes. J Mol Cell Cardiol 34:223–231
Ishida H, Takizawa M, Ozawa S et al. (2004) Pioglitazone improves secretary and prevents the loss of β-cell mass in obese diabetic db/db mice: possible protection of β cells from oxidative stress. Metabolism 53:488–494
Szkudelski T (2001) The mechanism of alloxan and streptozotocin action in B cells of the rat pancreas. Physiol Res 50:537–546
Lyon CJ, Hsueh WA (2003) Effect of plasminogen activator inhibitor-1 in diabetes mellitus and cardiovascular disease. Am J Med 115:S62–S68
Schneider DJ, Sobel BE (1996) Synergistic augmentation of expression of plasminogen activator inhibitor type-1 induced by insulin, very low density lipoprotein, and fatty acids. Coron Artery Dis 7:813–817
Asai M, Yoshinobu Y, Kaneko S et al. (2001) Circadian profile of Per gene mRNA expression in the suprachiasmatic nucleus, paraventricular nucleus, and pineal body of aged rats. J Neurosci Res 66:1133–1139
Murakami DM, Horwitz BA, Fuller CA (1995) Circadian rhythms of temperature and activity in obese and lean Zucker rats. Am J Physiol 269:R1038–R1043
Clark JB, Palmer CJ, Shaw WN (1983) The diabetic Zucker fatty rat. Proc Soc Exp Biol Med 173:68–75
Phillips MS, Liu Q, Hammond HA et al. (1996) Leptin receptor missense mutation in the fatty Zucker rat. Nat Genet 13:18–19
Zucker TF, Zucker LM (1996) Fat accretion and growth in the rat. Obes Res 4:102–108
Ionescu E, Sauter JF, Jeanrenaud B (1985) Abnormal oral glucose tolerance in genetically obese (fa/fa) rats. Am J Physiol 248:E500–E506
Stern J, Johnson PR, Greenwood MR, Zucker LM, Hirsch J (1972) Insulin resistance and pancreatic insulin release in the genetically obese Zucker rat. Proc Soc Exp Biol Med 139:66–69
Oishi K, Kasamatsu M, Ishida N (2004) Gene- and tissue-specific alterations of circadian clock gene expression in streptozotocin-induced diabetic mice under restricted feeding. BBRC 317:330–334
Suzuki Y, Urano T, Ihara H et al. (2001) Bezafibrate attenuates the overexpression of plasminogen activator inhibitor-1 messenger RNA by a combination of mono-unsaturated fatty acid and insulin in HepG2 cells. Life Sciences 68:1827–1837
Ihara H, Urano T, Takada A, Loskutoff DJ (2001) Induction of plasminogen activator inhibitor 1 gene expression in adipocytes by thiazolidinediones. FASEB J 15:1233–1235
Acknowledgements
This work was partially supported by grants awarded to S. Shibata from the Japanese Ministry of Education, Sports, and Culture (15390074, 14657621), the Special Coordination Funds of the Japanese Science and Technology Agency, and Waseda University. We thank Ms Hayasaka for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kudo, T., Akiyama, M., Kuriyama, K. et al. Night-time restricted feeding normalises clock genes and Pai-1 gene expression in the db/db mouse liver. Diabetologia 47, 1425–1436 (2004). https://doi.org/10.1007/s00125-004-1461-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-004-1461-0