Abstract
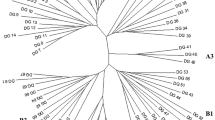
Amplified fragment length polymorphism (AFLP) analysis is a rapid and efficient method for producing DNA fingerprints. The AFLP diversity of sunflower has not been described, and much of the public germ plasm of sunflower has not yet been fingerprinted. Our objectives were to: (1) estimate genetic similarities, polymorphism rates, and polymorphic information contents (PICs) for AFLP markers among elite public oilseed inbred lines, and (2) assess the genetic diversity of inbred lines using genetic similarities estimated from AFLP fingerprints. We produced fingerprints for 24 public inbred lines of sunflower (Helianthus annuus L.) using six AFLP primer combinations. These primers produced a total of 359 AFLP markers or about 60 markers per primer combination. Genetic similarities ranged from 0.70 to 0.91, polymorphism rates ranged from 7 to 24%, and PICs ranged from 0.0 to 0.5. Genetic similarities were lower overall for maintainer (B)×restorer (R) crosses than for B×B or R×R crosses. Principal-coordinate and cluster analyses separated lines into two groups, one for B-lines and another for R-lines. These groupings illustrate the breeding history and basic heterotic pattern (B×R) of sunflower and the widespread practice of using B×B and R×R crosses to develop new lines. There were, nevertheless, distinct subgroups within these groups. These subgroups may represent unique heterotic groups and create a basis for formally describing heterotic patterns in sunflower.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 10 June 1996 / Accepted: 4 April 1997
Rights and permissions
About this article
Cite this article
Hongtrakul, V., Huestis, G. & Knapp, S. Amplified fragment length polymorphisms as a tool for DNA fingerprinting sunflower germplasm: genetic diversity among oilseed inbred lines. Theor Appl Genet 95, 400–407 (1997). https://doi.org/10.1007/s001220050576
Issue Date:
DOI: https://doi.org/10.1007/s001220050576