Abstract
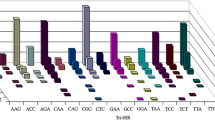
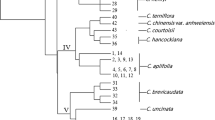
We have generated 66 sequence-tagged-site (STS) markers from cDNA clones of Cryptomeria japonica, and 60% of them have already been mapped into C. japonica linkage groups. All of the STS markers showed a single fragment following polymerase chain reaction (PCR) amplification. We investigated by polymorphism of these STS markers in a mapped F2 population and 15 plus trees by means of a restriction endonuclease analysis. Polymorphism levels were 10.6% and 22.7% in the F2 population and the 15 plus trees, respectively. PCR amplification levels of the 66 STS markers in 14 conifer species varied depending on their genetic relationship with C. japonica. Taxodium, which is closely related to C. japonica, had the most amplifications (31.82%), followed by Sequoiadendron giganteum, which is of the same family. The average proportion of PCR amplifications in each family gradually declined in the following order: from Taxodiaceae to Cuppresaceae, Sciadopityaceae, Pinaceae, and Taxaceae. These results are in general agreement with a molecular phylogenetic relationship based on chloroplast DNA. The 66 STS markers will be useful as on anchor point for genome mapping and population genetics, and some of them will also be useful when studying other conifers.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 8 September 1996 / Accepted: 8 November 1996
Rights and permissions
About this article
Cite this article
Tsumura, Y., Suyama, Y., Yoshimura, K. et al. Sequence-tagged-sites (STSs) of cDNA clones in Cryptomeria japonica and their evaluation as molecular markers in conifers. Theor Appl Genet 94, 764–772 (1997). https://doi.org/10.1007/s001220050476
Issue Date:
DOI: https://doi.org/10.1007/s001220050476