Abstract
Key message
Fine mapping of Yr47 and Lr52 in chromosome arm 5BS of wheat identified close linkage of the marker sun180 to both genes and its robustness for marker-assisted selection was demonstrated.
Abstract
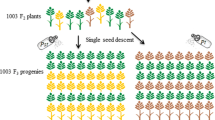
The widely effective and genetically linked rust resistance genes Yr47 and Lr52 have previously been mapped in the short arm of chromosome 5B in two F3 populations (Aus28183/Aus27229 and Aus28187/Aus27229). The Aus28183/Aus27229 F3 population was advanced to generate an F6 recombinant inbred line (RIL) population to identify markers closely linked with Yr47 and Lr52. Diverse genomic resources including flow-sorted chromosome survey sequence contigs representing the orthologous region in Brachypodium distachyon, the physical map of chromosome arm 5BS, expressed sequence tags (ESTs) located in the 5BS6-0.81-1.00 deletion bin and resistance gene analog contigs of chromosome arm 5BS were used to develop markers to saturate the target region. Selective genotyping was also performed using the iSelect 90 K Infinium wheat SNP assay. A set of SSR, STS, gene-based and SNP markers were developed and genotyped on the Aus28183/Aus27229 RIL population. Yr47 and Lr52 are genetically distinct genes that mapped 0.4 cM apart in the RIL population. The SSR marker sun180 co-segregated with Lr52 and mapped 0.4 cM distal to Yr47. In a high resolution mapping population of 600 F2 genotypes Yr47 and Lr52 mapped 0.2 cM apart and marker sun180 was placed 0.4 cM distal to Lr52. The amplification of a different sun180 amplicon (195 bp) than that linked with Yr47 and Lr52 (200 bp) in 204 diverse wheat genotypes demonstrated its robustness for marker-assisted selection of these genes.



Similar content being viewed by others
References
Alfares W, Bouguennec A, Balfourier F, Gay G, Bergès H, Vautrin S, Sourdille P, Bernard M, Feuillet C (2009) Fine mapping and marker development for the crossability gene SKr on chromosome 5BS of hexaploid wheat (Triticum aestivum L.). Genetics 183:469–481
Bansal UK, Forrest K, Hayden M, Miah H, Singh D, Bariana H (2011) Characterisation of a new stripe rust resistance gene Yr47 and its genetic association with the leaf rust resistance gene Lr52. Theor Appl Genet 122:1461–1466
Bansal UK, Kazi AG, Singh B, Hare RA, Bariana HS (2014a) Mapping of durable stripe rust resistance in a durum wheat cultivar Wollaroi. Mol Breed 33:51–59
Bansal U, Bariana H, Wong D, Randhawa M, Wicker T, Hayden M, Keller B (2014b) Molecular mapping of an adult plant stem rust resistance gene Sr56 in winter wheat cultivar Arina. Theor Appl Genet 127:1441–1448
Bansal M, Kaur S, Dhaliwal H, Bariana HS, Bansal UK, Chhuneja P (2015) Introgression of linked rust resistance genes Lr76 and Yr70 from Aegilops umbellulata to wheat chromosome 5DS. Plant Pathol. doi:10.1111/ppa.12549
Bariana HS (2003) Breeding for disease resistance. In: Thomas B, Murphy DJ, Murray BG (eds) Encyclopedia of applied plant sciences. Harcourt, Academic Press, UK, pp 244–253
Bariana H, McIntosh R (1995) Genetics of adult plant stripe rust resistance in four Australian wheats and the French cultivar ‘Hybride-de-Bersée’. Plant Breed 114:485–491
Bariana HS, Brown GN, Bansal UK, Miah H, Standen GE, Lu M (2007) Breeding for triple rust resistance wheat cultivars for Australia using conventional and marker assisted selection technologies. Aust J Agric Res 58:576–587
Bariana HS, Forrest K, Qureshi N, Miah H, Hayden M, Bansak UK (2016) Adult plant stripe rust resistance Yr71 maps close Lr24 in chromosome 3D of common wheat. Mol Breed. doi:10.1007/s11032-016-0528-1
Choudhary K, Choudhary O, Shekhawat N (2008) Marker assisted selection: a novel approach for crop improvement. Am-Eurasian J Agron 1:26–30
Dawson AM, Ferguson JN, Gardiner M, Green P, Hubbard A, Moscou MJ (2016) Isolation and fine mapping of Rps6: an intermediate host resistance gene in barley to wheat stripe rust. Theor Appl Genet 129:1–13
Gustafson P, Raskina O, Ma X, Nevo E (2009) Wheat evolution, domestication, and improvement. Sci Trade, Wheat, pp 5–30
Hiebert C, Thomas J, McCallum B (2005) Locating the broadspectrum wheat leaf rust resistance gene Lr52 (LrW) to chromosome 5B by a new cytogenetic method. Theor Appl Genet 110:1453–1457
International Wheat Genome Sequencing Consortium (2014) A chromosome-based draft sequence of the hexaploid bread wheat genome. Science 345:1251788
Jin Y, Szabo L, Pretorius Z, Singh R, Ward R, Fetch T Jr (2008) Detection of virulence to resistance gene Sr24 within race TTKS of Puccinia graminis f. sp. tritici. Plant Dis 92:923–926
Kolmer J (2013) Leaf rust of wheat: pathogen biology, variation and host resistance. Forests 4:70–84
Kosambi DD (1943) The estimation of map distances from recombination values. Annu Eugen 12:172–175
Kosina P, Reynolds M, Dixon J, Joshi A (2007) Stakeholder perception of wheat production constraints, capacity building needs, and research partnerships in developing countries. Euphytica 157:475–483
Kuraparthy V, Chhuneja P, Dhaliwal HS, Kaur S, Bowden RL, Gill BS (2007) Characterization and mapping of cryptic alien introgression from Aegilops geniculata with new leaf rust and stripe rust resistance genes Lr57 and Yr40 in wheat. Theor Appl Genet 114:1379–1389
Lorieux M (2012) MapDisto: fast and efficient computation of genetic linkage maps. Mol Breed 30:1231–1235
Lu HJ, Fellers JP, Friesen TL, Meinhardt SW, Faris JD (2006) Genomic analysis and marker development for the Tsn1 locus in wheat using bin-mapped ESTs and flanking BAC contigs. Theor Appl Genet 112:1132–1142
Marais G, Pretorius Z, Wellings C, McCallum B, Marais A (2005) Leaf rust and stripe rust resistance genes transferred to common wheat from Triticum dicoccoides. Euphytica 143:115–123
Marais F, Marais A, McCallum B, Pretorius Z (2009) Transfer of leaf rust and stripe rust resistance genes and from req. ex Bertol. to common wheat. Crop Sci 49:871–879
McIntosh RA, Welling CR, Park RF (1995) Wheat rusts: an atlas of resistance genes. CSIRO, Melbourne, p 200
Murray GM, Brennan JP (2009) Estimating disease losses to the Australian wheat industry. Aust Plant Pathol 38:558–570
Nesterov MA, Afonnikov DA, Sergeeva EM, Miroshnichenko LA, Bragina MK, Bragin AO, Vasiliev GV, Salina EA (2015) Identification of microsatellite loci according to BAC sequencing data and their physical mapping to the bread wheat 5B chromosome. Vavilov J Genet Breed 19:707–714
Pretorius Z, Singh R, Wagoire W, Payne T (2000) Detection of virulence to wheat stem rust resistance gene Sr31 in Puccinia graminis f. sp. tritici in Uganda. Plant Dis 84:203
Randhawa M, Bariana H, Mago R, Bansal U (2015) Mapping of a new stripe rust resistance locus Yr57 on chromosome 3BS of wheat. Mol Breed 35:1–8
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Rosegrant MW, Msangi S, Ringler C, Sulser TB, Zhu T, Cline SA (2008) International model for policy analysis of agricultural commodities and trade (IMPACT): model description. International Food Policy Research Institute, Washington DC
Sharp PJ, Johnston S, Brown G, McIntosh RA, Palotta M, Carter M, Bariana HS, Khatkar S, Lagudah ES, Singh RP, Khairallah M, Potter R, Jones MGK (2001) Validation of molecular markers for wheat breeding. Aust J Agric Res 52:1357–1366
Shi G, Zhang Z, Friesen TL, Bansal U, Cloutier S, Wicker T, Rasmussen JB, Faris JD (2016) Marker development, saturation mapping, and high-resolution mapping of the Septoria nodorum blotch susceptibility gene Snn3-B1 in wheat. Mol Genet Genomics 291:107–119
Singh RP, Huerta-Espino J, Rajaram S (2000) Achieving near immunity to leaf and stripe rusts in wheat by combining slow rusting resistance genes. Acta Phytopathol Entomol Hung 35:133–139
Singh RP, Hodson DP, Jin Y, Huerta-Espino J, Kinyua MG, Wanyera R, Njau P, Ward RW (2006) Current status, likely migration and strategies to mitigate the threat to wheat production from race Ug99 (TTKS) of stem rust pathogen. CAB reviews: perspectives in agriculture, veterinary science, nutrition and natural resources 1:1–13
Voorrips R (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, International Wheat Genome Sequencing C, Lillemo M, Mather D, Appels R, Dolferus R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo MC, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E (2014) Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotech J 12:787–796
Wellings C (2007) Puccinia striiformis in Australia: a review of the incursion, evolution, and adaptation of stripe rust in the period 1979–2006. Aust J Agric Res 58:567–575
Xue S, Zhang Z, Lin F, Kong Z, Cao Y, Li C, Yi H, Mei M, Zhu H, Wu J, Xu H, Zhao D, Tian D, Zhang C, Ma Z (2008) A high-density intervarietal map of the wheat genome enriched with markers derived from expressed sequence tags. Theor Appl Genet 117:181–189
Yang H, Li C, Lam HM, Clements J, Yan G, Zhao S (2015) Sequencing consolidates molecular markers with plant breeding practice. Theor Appl Genet 128:779–795
Acknowledgements
Naeela Qureshi acknowledges the University of Sydney for the USYDIS award. We thank the Grains Research and Development Corporation (GRDC) Australia for financial support through the Australian Cereal Rust Control Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors have read the manuscript and declare that they have no conflict of interest.
Additional information
Communicated by A Zhang.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Qureshi, N., Bariana, H., Forrest, K. et al. Fine mapping of the chromosome 5B region carrying closely linked rust resistance genes Yr47 and Lr52 in wheat. Theor Appl Genet 130, 495–504 (2017). https://doi.org/10.1007/s00122-016-2829-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-016-2829-5