Abstract
Key message
We identified 11 SAD genes, and mined their natural variations associated with the conservation of stearic to oleic acid, especially ZmSAD1 supported by both the QTL and an expression QTL.
Abstract
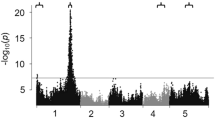
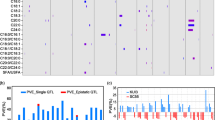
Maize oil is generally regarded as a healthy vegetable oil owing to its low abundance of saturated fatty acids. Stearoyl-ACP desaturase (SAD) is a key rate-limiting enzyme for the conservation of stearic (C18:0) to oleic (C18:1) acid. Here, 11 maize SAD genes were identified to have more divergent functions than Arabidopsis SAD genes. The genomic regional associations in a maize panel including 508 inbred lines identified 6 SAD genes significantly associated (P < 0.01) with the C18:0/C18:1 ratio or the level of C18:0 or C18:1, one gene of which co-localized with a quantitative trait locus (QTL) and 5 of which co-localized with an expression QTL. ZmSAD1, supported by both the QTL and an expression QTL, had the largest effect on C18:0/C18:1. One nonsynonymous single-nucleotide polymorphism in exon 3 and one 5-bp insertion/deletion in the 3′ untranslated region were further shown to contribute to the natural variation in C18:0/C18:1 according to ZmSAD1-based association mapping. Finally, selection tests of ZmSAD1 in teosinte, regular maize, and high-oil maize indicated that ZmSAD1 was not a selection target during the process of maize domestication and high-oil maize development. These results will guide the manipulation of the ratio between saturated and unsaturated fatty acids in maize.





Similar content being viewed by others
References
Alrefai R, Berke TG, Rocheford TR (1995) Quantitative trait locus analysis of fatty acid concentrations in maize. Genome 38:894–901
Beló A, Zheng P, Luck S, Shen B, Meyer DJ, Li B, Tingey S, Rafalski A (2008) Whole genome scan detects an allelic variant of fad2 associated with increased oleic acid levels in maize. Mol Genet Genomics 279:1–10
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Chai YC, Hao XM, Yang XH, Allen WB, Li JM, Yan JB, Shen B, Li JS (2012) Validation of DGAT1-2 polymorphisms associated with oil content and development of functional markers for molecular breeding of high-oil maize. Mol Breed 29:939–949
Chen J, Zeng B, Zhang M, Xie SJ, Wang GK, Hauck A, Lai JS (2014) Dynamic transcriptome landscape of maize embryo and endosperm development. Plant Physiol 166:252–264
Doebley J, Stec A (1991) Genetic analysis of the morphological differences between maize and teosinte. Genetics 129:285–295
Doebley J, Stec A, Gustus C (1995) Teosinte branched1 and the origin of maize: evidence for epistasis and the evolution of dominance. Genetics 141:333–346
Du HW, Yang XH, Yan JB, Li JS (2011) Fatty acid elongase 1 (FAE1) promoter as a candidate for genetic engineering of fatty acids to improve seed oil composition. Afr J Biotechnol 10:19615–19622
Edgar RC (2004a) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Edgar RC (2004b) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC bioinform 5:113
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6:e19379
Fu JJ, Cheng YB, Linghu JJ, Yang XH, Kang L, Zhang ZX, Zhang J, He C, Du XM, Peng ZY, Wang B, Zhai LH, Dai CM, Xu JB, Wang WD, Li XR, Zheng J, Chen L, Luo LH, Liu JJ, Qian XJ, Yan JB, Wang J, Wang GY (2013) RNA sequencing reveals the complex regulatory network in the maize kernel. Nat Commun 4:2832
Ganal MW, Durstewitz G, Polley A, Bérard A, Buckler ES, Charcosset A, Clarke JD, Graner EM, Hansen M, Joets J, Le Paslier MC, McMullen MD, Montalent P, Rose M, Schon CC, Sun Q, Walter H, Martin OC, Falque M (2011) A large maize (Zea mays L) SNP genotyping array: development and germplasm genotyping, and genetic mapping to compare with the B73 reference genome. PLoS One 6:e28334
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hudson RR, Kreitman M, Aguadé M (1987) A test of neutral molecular evolution based on nucleotide data. Genetics 116:153–159
Hufford MB, Xu X, van Heerwaarden J, Pyhäjärvi T, Chia JM, Cartwright RA, Elshire RJ, Glaubitz JC, Guill KE, Kaeppler SM, Lai JS, Morrell PL, Shannon LM, Song C, Springer NM, Swanson-Wagner RA, Tiffin P, Wang J, Zhang GY, Doebley J, McMullen MD, Ware D, Buckler ES, Yang S, Ross-Ibarra J (2012) Comparative population genomics of maize domestication and improvement. Nat Genet 44:808–811
Kachroo A, Shanklin J, Whittle E, Lapchyk L, Hildebrand D, Kachroo P (2007) The Arabidopsis stearoyl-acyl carrier protein-desaturase family and the contribution of leaf isoforms to oleic acid synthesis. Plant Mol Biol 63:257–271
Knutzon DS, Scherer DE, Schreckengost WE (1991) Nucleotide sequence of a complementary DNA clone encoding stearoyl-acyl carrier protein desaturase from castor bean, Ricinus communis. Plant Physiol 96:344–345
Lambert RJ (2001) High-oil corn hybrids. In: Hallau AR (ed) Special corn. CRC Press Inc, Boca Raton, pp 131–153
Li L, Li H, Li Q, Yang XH, Zheng DB, Warburton M, Chai YC, Zhang P, Guo YQ, Yan JB, Li JS (2011) An 11-bp insertion in Zea mays fatb reduces the palmitic acid content of fatty acids in maize grain. PLoS One 6:e24699
Li Q, Yang XH, Xu ST, Cai Y, Zhang DL, Han YJ, Li L, Zhang ZX, Gao SB, Li JS, Yan JB (2012) Genome-wide association studies identified three independent polymorphisms associated with α-tocopherol content in maize kernels. PLoS One 7:e36807
Li H, Peng ZY, Yang XH, Wang WD, Fu JJ, Wang JH, Han YJ, Chai YC, Guo TT, Yang N, Liu J, Warburton ML, Cheng YB, Hao XM, Zhang P, Zhao JY, Liu YJ, Wang GY, Li JS, Yan JB (2013) Genome-wide association study dissects the genetic architecture of oil biosynthesis in maize kernels. Nat Genet 45:43–50
Li F, Bian CS, Xu JF, Pang WF, Liu J, Duan SG, Lei ZG, Jiwan P, Jin LP (2015) Cloning and functional characterization of SAD genes in potato. PLoS One 10:e0122036
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Liu ZJ, Yang XH, Fu Y (2009a) SAD, a stearoyl-acyl carrier protein desaturase highly expressed in high-oil maize inbred lines. Russ J Plant Physiol 56:709–715
Liu ZJ, Yang XH, Fu Y, Zhang YR, Yan JB, Song TM, Rocheford T, Li JS (2009b) Proteomic analysis of early germs with high-oil and normal inbred lines in maize. Mol Biol Rep 36:813–821
Liu HJ, Luo X, Niu LY, Xiao YJ, Chen L, Liu J, Wang XQ, Jin ML, Li WQ, Zhang QH, Yan JB (2016) Distant eQTLs and non-coding sequences play critical roles in regulating gene expression and quantitative trait variation in maize. Mol Plant. doi:10.1016/j.molp.2016.06.016
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
López Alonso D, García-Maroto F, Rodríguez-Ruiz J, Garrido JA, Vilches MA (2003) Evolution of the membrane-bound fatty acid desaturases. Biochem Syst Ecol 31:1111–1124
Matsuoka Y, Vigouroux Y, Goodman MM, Sanchez GJ, Buckler E, Doebley J (2002) A single domestication for maize shown by multilocus microsatellite genotyping. Proc Natl Acad Sci 99:6080–6084
Merlo AO, Cowen N, Delate T, Edington B, Folkerts O, Hopkins N, Lemeiux C, Skokut T, Smith K, Woosley A, Yang YJ, Young S, Zwick M (1998) Ribozymes targeted to stearoyl-ACP Δ9 desaturase mRNA produce heritable increases of stearic acid in transgenic maize leaves. Plant Cell 10:1603–1621
Ohlrogge J, Browse J (1995) Lipid biosynthesis. Plant Cell 7:957–970
Ohlrogge JB, Jaworski JG (1997) Regulation of fatty acid synthesis. Ann Rev Plant Physiol Plant Mol Biol 48:109–136
Piperno DR, Ranere AJ, Holst I, Iriarte J, Dickau R (2009) Starch grain and phytolith evidence for early ninth millennium BP maize from the Central Balsas River Valley, Mexico. Proc Natl Acad Sci 106:5019–5024
Ramsden CE, Zamora D, Leelarthaepin B, Majchrzak-Hong SF, Faurot KR, Suchindran CM, Ringel A, Davis JM, Hibbeln JR (2013) Use of dietary linoleic acid for secondary prevention of coronary heart disease and death: evaluation of recovered data from the Sydney Diet Heart Study and updated meta-analysis. BMJ 346:e8707
Ruddle P II, Whetten R, Cardinal A, Upchurch RG, Miranda L (2014) Effect of Δ9-stearoyl -ACP-desaturase-C mutants in a high oleic background on soybean seed oil composition. Theor Appl Genet 127:349–358
Shanklin J, Mullins C, Somerville C (1991) Sequence of a complementary DNA from Cucumis sativus L encoding the stearoyl-acyl-carrier protein desaturase. Plant Physiol 97:467–468
Shilman F, Brand Y, Brand A, Hedvat I, Hovav R (2011) Identification and molecular characterization of homeologous Δ9-stearoyl acyl carrier protein desaturase 3 genes from the allotetraploid peanut (Arachis hypogaea). Plant Mol Biol Rep 29:232–241
Somerville C, Browse J, Jaworski JG, Ohlrogge JB (2000) Lipids. In: Buchanan BB, Gruissem W, Jones RL (eds) Biochemistry and molecular biology of plants. American Society of Plant Physiologist, Rockville, pp 456–526
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 60. Mol Biol Evol 30:2725–2729
Tenaillon MI, U’Ren J, Tenaillon O, Gaut BS (2004) Selection versus demography: a multilocus investigation of the domestication process in maize. Mol Biol Evol 21:1214–1225
Thambugala D, Cloutier S (2014) Fatty acid composition and desaturase gene expression in flax (Linum usitatissimum L). J Appl Genet 55:423–432
Thompson GA, Scherer DE, Foxall-Van Aken S, Kenny JW, Young HL, Shintani DK, Kridl JC, Knauf VC (1991) Primary structures of the precursor and mature forms of stearoyl-acyl carrier protein desaturase from safflower embryos and requirement of ferredoxin for enzyme activity. Proc Natl Acad Sci 88:2578–2582
van Heerwaarden J, Doebley J, Briggs WH, Glaubitz JC, Goodman MM, Gonzalez S, de Jesus J, Ross-Ibarra J (2011) Genetic signals of origin, spread, and introgression in a large sample of maize landraces. Proc Natl Acad Sci 108:1088–1092
Wang H, Nussbaum-Wagler T, Li BL, Zhao Q, Vigouroux Y, Faller M, Bomblies K, Lukens L, Doebley JF (2005) The origin of the naked grains of maize. Nature 436:714–719
Wassom JJ, Mikkelineni V, Bohn MO, Rocheford TR (2008) QTL for fatty acid composition of maize kernel oil in Illinois high oil × B73 backcross-derived lines. Crop Sci 48:69–78
Weber EJ (1987) Lipids of the kernel. In: Watson SA, Ramstad PE (eds) Corn: chemistry and technology. American Association of Ceral Chemists St Paul, Minn, pp 311–349
Yang XH, Guo YQ, Yan JB, Zhang J, Song TM, Rocheford T, Li JS (2010a) Major and minor QTL and epistasis contribute to fatty acid compositions and oil concentration in high-oil maize. Theor Appl Genet 120:665–678
Yang XH, Yan JB, Shah T, Warburton ML, Li Q, Li L, Gao YF, Chai YC, Fu ZY, Zhou Y, Xu ST, Bai GH, Meng YJ, Zheng YP, Li JS (2010b) Genetic analysis and characterization of a new maize association mapping panel for quantitative trait loci dissection. Theor Appl Genet 121:417–431
Yang XH, Gao SB, Xu ST, Zhang ZX, Prasanna BM, Li L, Li JS, Yan JB (2011) Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize. Mol Breed 28:511–526
Yang N, Lu YL, Yang XH, Huang J, Zhou Y, Ali F, Wen WW, Liu J, Li JS, Yan JB (2014) Genome wide association studies using a new nonparametric model reveal the genetic architecture of 17 agronomic traits in an enlarged maize association panel. PLoS Genet 10:e1004573
Yu JM, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhang YF, Maximova SN, Guiltinan MJ (2015) Characterization of a stearoyl-acyl carrier protein desaturase gene family from chocolate tree, Theobroma cacao L. Front Plant Sci 6:239
Zheng PZ, Allen WB, Roesler K, Williams ME, Zhang SR, Li JM, Glassman K, Ranch J, Nubel D, Solawetz W, Bhattramakki D, Llaca V, Deschamps S, Zhong GY, Tarczynski MC, Shen B (2008) A phenylalanine in DGAT is a key determinant of oil content and composition in maize. Nat Genet 40:367–372
Acknowledgments
We greatly appreciate Dr. Jianbing Yan and Mr. Haijun Liu for providing the genotyping-by-sequencing data for the maize association panel. We also appreciate the helpful comments on the manuscript from three anonymous reviewers. This research was supported by the National Natural Science Foundation of China (Grant Number 31361140362) and the Program for New Century Excellent Talents in University of the Ministry of Education of China (Grant Number 2014FG049).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by A. A. Franco Garcia.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Han, Y., Xu, G., Du, H. et al. Natural variations in stearoyl-acp desaturase genes affect the conversion of stearic to oleic acid in maize kernel. Theor Appl Genet 130, 151–161 (2017). https://doi.org/10.1007/s00122-016-2800-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-016-2800-5