Abstract
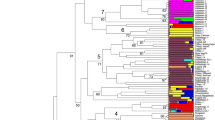
Twenty-four simple sequence repeat (SSR) markers were used to detect molecular polymorphisms among 370 mostly sexually derived Citrus accessions from the collection of citrus germplasm maintained at the University of California, Riverside. A total of 275 alleles were detected with an average of 11.5 alleles per locus and an average polymorphism information content of 0.625. Genetic diversity statistics were calculated for each individual SSR marker, the entire population, and for specified Citrus groups. Phylogenetic relationships among all citrus accessions and putative non-hybrid Citrus accessions were determined by constructing neighbor-joining trees. There was strong support for monophyly at the species level when hybrid taxa were removed from the data set. Both of these trees indicate that Fortunella clusters within the genus Citrus but Poncirus is a sister genus to Citrus. Additionally, Citrus accessions were probabilistically assigned to populations or multiple populations if their genotype indicated an admixture by a model-based clustering approach. This approach identified five populations in this data set. These separate analyses (distance and model based) both support the hypothesis that there are only a few naturally occurring species of Citrus and most other types of Citrus arose through various hybridization events between these naturally occurring forms.


Similar content being viewed by others
References
Ahmad R, Struss D, Southwick SM (2003) Development and characterization of microsatellite markers in Citrus. J Amer Soc Hort Sci 128:584–590
Barrett HC, Rhodes AM (1976) A numerical taxonomic study of affinity relationships in cultivated Citrus and its close relatives. Syst Bot 1:105–136
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Bowcock AM, Ruiz-Linares A, Tomfohrde J, Minch E, Kidd JR, Cavalli-Sforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Nature 368:455–457
Bretó MP, Ruiz C, Pina JA, Asins MJ (2001) The diversification of Citrus clementina Hort. Ex Tan., a vegetatively propagated crop species. Mol Phyl and Evol 21:285–293
Brown SM, Hopkins MS, Mitchell SE, Senior ML, Wang TY, Duncan RR, Gonzalez-Candelas F, Kresovich S (1996) Multiple methods for the identification of polymorphic simple sequence repeats (SSRs) in sorghum. Theor Appl Genet 93:190–198
Dangl GS, Mendum ML, Prins BH, Walker MA, Meredith CP, Simon CJ (2001) Simple sequence repeat analysis of a clonally propagated species: a tool for managing a grape germplasm collection. Genome 44:432–438
Fang DQ, Roose ML (1997) Identification of closely related Citrus cultivars with inter-simple sequence repeat markers. Theor Appl Genet 95:408–417
Fang DQ, Roose ML, Krueger RR, Federici CT (1997) Fingerprinting trifoliate orange germplasm accessions with isozymes, RFLPs, and inter-simple sequence repeat markers. Theor Appl Genet 95:211–219
Federici CT, Fang DQ, Scora RW, Roose ML (1998) Phylogenetic relationships within the genus Citrus (Rutaceae) and related genera as revealed by RFLP and RAPD analysis. Theor Appl Genet 94:812–822
Green RM, Vardi A, Galun E, (1986) The plastome of Citrus: physical map, variation among Citrus cultivars and species and comparison with related genera. Theor Appl Genet 72:170–177
Gulsen O, Roose ML (2001a) Lemons: diversity and relationships with selected Citrus genotypes as measured with nuclear genome markers. J Amer Soc Hort Sci 126:309–327
Gulsen O, Roose ML (2001b) Chloroplast and nuclear genome analysis of the parentage of lemons. J Amer Soc Hort Sci 126:210–215
Herrero R, Asins MJ, Carbonell EA, Navarro L (1996) Genetic diversity in the orange subfamily Aurantioideae. I. Intraspecies and intragenus genetic variability. Theor Appl Genet 92:599–609
Hodgson RW (1967) Horticultural varieties of citrus. In: Reuther W, Webber HJ, Batchelor LD (eds) The Citrus Industry. Vol I. University of California Press, Berkeley, pp 431–591
Hokanson SC, Szewc-McFadden AK, Lamboy WF, McFerson JR (1998) Microsatellite (SSR) markers reveal genetic identities, genetic diversity and relationships in a Malus × domestica borkh. Core subset collection. Theor Appl Genet 97:671–683
Jain MS, Brar DS, Ahloowalia BS (2002) Microsatellites and molecular breeding: exploitation of microsatellite variability for the analysis of a monotonous genome. In: Molecular techniques in crop improvement. Kluwer, London, pp 85–135
Jarrell DC, Roose ML, Traugh SN, Kupper RS (1992) A genetic map of Citrus based on the segregation of isozymes and RFLPs in an intergeneric cross. Theor Appl Genet 84:49–56
Kijas JM, Thomas MR, Fowler JC, Roose ML (1997) Integration of trinucleotide microsatellites into a linkage map of Citrus. Theor Appl Genet 94:701–706
Liu K, Goodman M, Muse S, Smith JS, Buckler E, Doebley J (2003) Genetic structure and diversity among Maize inbred lines as inferred from DNA microsatellites. Genetics 165:2117–2128
Luro F, Laigret F, Bove JM, Ollitrault P (1995) DNA amplified fingerprinting, a useful tool for determination of genetic origin and diversity analysis in Citrus. Hort Sci 30:1063–1067
Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655
Minch E, Ruiz-Linares A, Goldstein D, Feldman M, Kidd JR, Cavalli-Sforza LL (1997) Microsat 1.5: a computer program for calculating various statistics on microsatellite allele data. Available via http://www.hpgl.stanford.edu/projects/microsat/programs/
Nicolosi E, Deng ZN, Gentile A, La Malfa S, Continella G, Tribulato E (2000) Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor Appl Genet 100:1155–1166
Pritchard J, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Roose ML, Soost RK, Cameron JW (1995) Citrus. In: Smart J, Simmonds NW (eds) The evolution of crop plants. 2nd edn, Longman, Essex, pp 443–449
Roose ML, Fang D, Cheng FS, Tayyar RI, Federici CT, Kupper RS (2000) Mapping of the Citrus genome. Acta Hortic 535:25–32
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Scora RW (1975) On the history and origin of Citrus. Bull Torr Bot Club 102:369–375
Scora RW, Kumamoto J, Esen A, Stone BC (1976) A phytochemical investigation of Citrus halimii. Biochem Sys and Ecol 4:255–258
Singh R, Schroeder CA (1962) Taxonomic and physiological relationships of the so-called mandarin-lime group of citrus. Proc Amer Soc Hort Sci 80:291–295
Stam P (1993) Construction of integrated genetic linkage maps by means of a new computer package: JoinMap. Plant J 3:739–744
Swingle WT, Reece PC (1967) The botany of Citrus and its wild relatives. In: Reuther W, Webber HJ, Batchelor LD (eds) The Citrus Industry. Vol 1. University of California Press, Berkeley, pp 190–430
Swofford DL (1998) PAUP*. Phylogenetic analysis using parismony (*and other methods). Version 4. Sinauer Associates, Sunderland
Tanaka T (1977) Fundamental discussion of Citrus classification. Stud Citrologia 14:1–6
Webb DM, Knapp SJ (1990) DNA extraction from a previously recalcitrant plant genus. Plant Mol Bio 8:180–185
Weber JL, Wong C (1993) Mutation of human short tandem repeats. Hum Mol Genet 2:1123–1128
Webber JH (1943) Cultivated varieties of citrus. In: Webber HJ, Batchelor LD (eds) The Citrus Industry. Vol 1. University of California Press, Berkeley. pp 475–668
Acknowledgements
We would like to thank the U.S. Department of Agriculture for the funding to support this project. In addition, we thank the Genetic Graduate Group at the University of California, Riverside for providing partial funding of this project. Also, thanks to Dr. Mark Springer and Dr. Angela Burk-Herrick (University of California, Riverside) for providing assistance on PAUP and phylogenetic interpretation. Thanks to Dr. Tracy Kahn and Ms. Toots Bier for their support in providing historical and morphological information on Citrus varieties from the Citrus Variety Collection (CVC archival data). Lastly, thanks to colleagues at USDA-ARS Plant Genetic Resource Conservation Unit in Griffin, GA, USA for reviewing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. S. Heslop-Harrison
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Barkley, N.A., Roose, M.L., Krueger, R.R. et al. Assessing genetic diversity and population structure in a citrus germplasm collection utilizing simple sequence repeat markers (SSRs). Theor Appl Genet 112, 1519–1531 (2006). https://doi.org/10.1007/s00122-006-0255-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-006-0255-9