Abstract
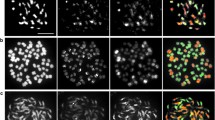
When a crop hybridizes with a wild relative, the potential for stable transmission to the wild of any crop gene is directly related to the frequency of crop–wild homoeologous pairing for the chromosomal region where it is located within the crop genome. Pairing pattern at metaphase I (MI) has been examined in durum wheat × Aegilops geniculata interspecific hybrids (2n=4x=ABUgMg) by means of a genomic in-situ hybridization procedure that resulted in simultaneous discrimination of A, B and wild genomes. The level of MI pairing in the hybrids varied greatly depending on the crop genotype. However, their pattern of homoeologous association was very similar, with a frequency of wheat–wild association close to 60% in all genotype combinations. A–wild represented 80–85% of wheat–wild associations which supports that, on average, A genome sequences are much more likely to be transferred to this wild relative following interspecific hybridization and backcrossing. Combination of genomic DNA probes and the ribosomal pTa71 probe has allowed to determine the MI pairing behaviour of the major NOR-bearing chromosomes in these hybrids (1B, 6B, 1Ug and 5Ug), in addition to wheat chromosome 4A which could be identified with the sole use of genomic probes. The MI pairing pattern of the wild chromosome arms individually examined has confirmed a higher chance of gene escape from the wheat A genome. However, a wide variation regarding the amount of wheat–wild MI pairing among the specific wheat chromosome regions under analysis suggests that the study should be extended to other homoeologous groups.

Similar content being viewed by others
References
Badaeva ED, Amosova AV, Samatadze TE, Zoshchuk SA, Shostak NG, Chikida NN, Zelenin AV, Raupp WJ, Friebe B, Gill BS (2004) Genome differentiation in Aegilops. 4. Evolution of the U-genome cluster. Plant Syst Evol 246:45–76
Benavente E, Orellana J, Fernandez-Calvin B (1998) Comparative analysis of the meiotic effects of wheat ph1b and ph2b mutations in wheat × rye hybrids. Theor Appl Genet 96:1200–1204
Benavente E, Alix K, Dusautoir JC, Orellana J, David JL (2001) Early evolution of the chromosomal structure of Triticum turgidum–Aegilops ovata amphiploids carrying and lacking the Ph1 gene. Theor Appl Genet 103:1123–1128
Cai X, Jones S (1997) Direct evidence for high level of autosyndetic pairing in hybrids of Thinopyrum intermedium and Th. ponticum with Triticum aestivum. Theor Appl Genet 95:568–572
Chen Q, Conner RL, Laroche A, Ahmad F (2001) Molecular cytogenetic evidence for a high level of chromosome pairing among different genomes in Triticum aestivum–Thinopyrum intermedium hybrids. Theor Appl Genet 102:847–852
Cuadrado A, Vitellozzi F, Jouve N, Ceoloni C (1997) Fluorescence in situ hybridization with multiple repeated DNA probes applied to the analysis of wheat–rye chromosome pairing. Theor Appl Genet 94:347–355
Dale PJ, Clarke B, Fontes EM (2002) Potential for the environmental impact of transgenic crops. Nat Biotechnol 20:567–574
David JL, Benavente E, Bres-Patry C, Dusautoir JC, Echaide M (2004) Are neopolyploids a likely route for a transgene walk to the wild? The Aegilops ovata × Triticum turgidum durum case. Biol J Linn Soc 82:503–510
Ellstrand NC, Prentice HC, Hancock JF (1999) Gene flow and introgression from domesticated plants into their wild relatives. Annu Rev Ecol Syst 30:539–563
ESF (2004) Programme ‘assessment of the impacts of genetically modified plants (AIGM)’ of European Science Foundation (1999–2004). http://www.esf.org/esf_article.php?language=0&activity=1&domain=3&article=49&page=97
FAO (2002) Gene flow from GM to non-GM populations in the crop, forestry, animal and fishery sectors. Electronic forum on biotechnology in food and agriculture, 7th conference (May 31–July 6, 2002). Summary in http://www.fao.org/biotech/logs/C7/summary.htm
Fernandez-Calvin B, Orellana J (1992) Relationship between pairing frequencies and genome affinity estimations in Aegilops ovata × Triticum aestivum hybrid plants. Heredity 68:165–172
Gerlach WL, Bedbrook JR (1979) Cloning and characterization of ribosomal-RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1885
Guadagnuolo R, SavovaBianchi D, Felber F (2001) Gene flow from wheat (Triticum aestivum L.) to jointed goatgrass (Aegilops cylindrica Host.), as revealed by RAPD and microsatellite markers. Theor Appl Genet 103:1–8
Jauhar PP, Riera-Lizarazu O, Dewey WG, Gill BS, Crane CF, Bennett JH (1991) Chromosome pairing relationships among the A, B, and D genomes of bread wheat. Theor Appl Genet 82:441–449
Jauhar PP, Almouslem AB, Peterson TS, Joppa LR (1999) Inter- and intragenomic chromosome pairing in haploids of durum wheat. J Hered 90:437–445
Jauhar PP, Dogramaci M, Peterson TS (2004) Synthesis and cytological characterization of trigeneric hybrids of durum wheat with and without Ph1. Genome 47:1173–1181
Jones HD (2005) Wheat transformation: current technology and applications to grain development and composition. J Cereal Sci 41:137–147
Lucas H, Jahier J (1988) Phylogenetic relationships in some diploid species of Triticineae: cytogenetic analysis of interspecific hybrids. Theor Appl Genet 75:498–502
Lukaszewski AJ, Lapinski B, Rybka K (2005) Limitations of in situ hybridization with total genomic DNA in routine screening for alien introgressions in wheat. Cytogenet Genome Res 109:373–377
Maestra B, Naranjo T (2000) Genome evolution in Triticeae. In: Olmo E, Redi CA (eds) Chromosomes today, vol 13. Birkhäuser Verlag, Switzerland, pp 155–167
Martinez M, Cuadrado C, Laurie DA, Romero C (2005) Synaptic behaviour of hexaploid wheat haploids with different effectiveness of the diploidizing mechanism. Cytogenet Genome Res 109:210–214
Mukai Y, Endo TR, Gill BS (1991) Physical mapping of the 18S.26S rRNA multigene family in common wheat: identification of a new locus. Chromosoma 100:71–78
Naranjo T (1990) Chromosome structure of durum wheat. Theor Appl Genet 79:397–400
Naranjo T, Fernandez-Rueda P (1996) Pairing and recombination between individual chromosomes of wheat and rye in hybrids carrying the ph1b mutation. Theor Appl Genet 93:242–248
Naranjo T, Roca A, Goicoechea PG, Giraldez R (1987) Arm homoeology of wheat and rye chromosomes. Genome 29:873–882
Ozkan H, Feldman M (2001) Genotypic variation in tetraploid wheat affecting homoeologous pairing in hybrids with Aegilops peregrina. Genome 44:1000–1006
Sahrawat AK, Becker D, Lutticke S, Lorz H (2003) Genetic improvement of wheat via alien gene transfer, an assessment. Plant Sci 165:1147–1168
Sanchez-Moran E, Benavente E, Orellana J (1999) Simultaneous identification of A, B, D and R genomes by genomic in situ hybridization in wheat–rye derivatives. Heredity 83:249–252
Sanchez-Moran E, Benavente E, Orellana J (2001) Analysis of karyotypic stability of homoeologous-pairing (ph) mutants in allopolyploid wheats. Chromosoma 110:371–377
van Slageren MW (1994) Wild wheats: a monograph of Aegilops L. and Amblyopyrum (Jaub and Spach) Eig (Poaceae). Agricultural University of Wageningen-ICARDA
Tomiuk J, Hauser TP, BaggerJorgensen R (2000) A- or C-chromosomes, does it matter for the transfer of transgenes from Brassica napus. Theor Appl Genet 100:750–754
Wang ZN, Hang A, Hansen J, Burton C, Mallory-Smith CA, Zemetra RS (2000) Visualization of A- and B-genome chromosomes in wheat (Triticum aestivum L.) × jointed goatgrass (Aegilops cylindrica Host) backcross progenies. Genome 43:1038–1044
Zhang H, Jia J, Gale MD, Devos KM (1998) Relationships between the chromosomes of Aegilops umbellulata and wheat. Theor Appl Genet 96:69–75
Acknowledgements
The authors are indebted to Dr. Jacques David for having provided access to the wild hybrid meiosis. We thank also Jean-Claude Dusautoir for his invaluable help in the production and growing of some of the interspecific hybrids. This study has been funded by grant AGL2002-1593 from the Dirección General de Investigación of Ministerio de Educación y Ciencia (formerly Ministerio de Ciencia y Tecnología) of Spain.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Friebe
Rights and permissions
About this article
Cite this article
Cifuentes, M., Blein, M. & Benavente, E. A cytomolecular approach to assess the potential of gene transfer from a crop (Triticum turgidum L.) to a wild relative (Aegilops geniculata Roth.). Theor Appl Genet 112, 657–664 (2006). https://doi.org/10.1007/s00122-005-0168-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-005-0168-z