Abstract
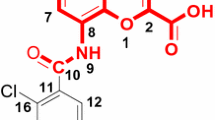
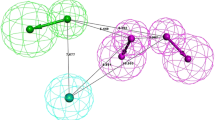
Pharmacophore modeling, comparative molecular field analysis (CoMFA), and comparative molecular similarity indices analysis (CoMSIA) studies have been carried out on 5-(4-piperidyl)-3-isoxazolol (4-PIOL) analogs as GABAA receptor antagonists in this study. The best pharmacophore hypothesis generated by PHASE was ADHPR.6, which comprised a hydrogen bond acceptor (A), a hydrogen bond donor (D), a hydrophobic group (H), a positively charged group (P), and an aromatic ring (R). The pharmacophore model provided a good alignment for the further 3D-QSAR analyses, which presented a good R 2 value of 0.943, 0.930, and 0.916 for atom-based QSAR model, CoMFA model, and CoMSIA model, respectively. All QSAR models presented good statistical significance and predictivity, the corresponding Q 2 values for each 3D-QSAR model are 0.794, 0.569, and 0.637, respectively. Both pharmacophore and CoMSIA results showed that the hydrophobic sites are the key structural feature for GABAA receptor antagonists with high activities.




Similar content being viewed by others
References
Ai Y, Wang ST, Tang C, Sun PH, Song FJ (2011) 3D-QSAR and docking studies on pyridopyrazinones as BRAF inhibitors. Med Chem Res 20(8):1298–1317
Almerico AM, Tutone M, Lauria A (2010) 3D-QSAR pharmacophore modeling and in silico screening of new Bcl-xl inhibitors. Eur J Med Chem 45(11):4774–4782. doi:10.1016/j.ejmech.2010.07.042
Chebib M, Johnston GAR (1999) The ‘ABC’ of GABA receptors: a brief review. Clin Exp Pharmacol Physiol 26(11):937–940. doi:10.1046/j.1440-1681.1999.03151.x
Cramer RD (2010) Rethinking 3D-QSAR. J Comput Aided Mol Des 25(3):197–201. doi:10.1007/s10822-010-9403-z
D’Hulst C, Atack JR, Kooy RF (2009) The complexity of the GABAA receptor shapes unique pharmacological profiles. Drug Discov Today 14(17–18):866–875. doi:10.1016/j.drudis.2009.06.009
Dixon SL, Smondyrev AM, Knoll EH, Rao SN, Shaw DE, Friesner RA (2006) PHASE: a new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening: 1. Methodology and preliminary results. J Comput Aided Mol Des 20(10–11):647–671. doi:10.1007/s10822-006-9087-6
Frolund B, Kristiansen U, Brehm L, Hansen AB, Krogsgaardlarsen P, Falch E (1995) PARTIAL GABA(A) RECEPTOR AGONISTS - SYNTHESIS AND IN-VITRO PHARMACOLOGY OF A SERIES OF NONANNULATED ANALOGS OF 4,5,6,7-TETRAHYDROISOXAZOLO 5,4-C PYRIDIN-3-OL. J Med Chem 38(17):3287–3296. doi:10.1021/jm00017a014
Frolund B, Tagmose L, Liljefors T, Stensbol TB, Engblom C, Kristiansen U, Krogsgaard-Larsen P (2000) A novel class of potent 3-isoxazolol GABA(A) antagonists: design, synthesis, and pharmacology. J Med Chem 43(26):4930–4933. doi:10.1021/jm000371q
Frolund B, Jorgensen AT, Tagmose L, Stensbol TB, Vestergaard HT, Engblom C, Kristiansen U, Sanchez C, Krogsgaard-Larsen P, Liljefors T (2002) Novel class of potent 4-arylalkyl substituted 3-isoxazolol GABA(A) antagonists: synthesis, pharmacology, and molecular modeling. J Med Chem 45(12):2454–2468. doi:10.1021/jm020027o
Frolund B, Tagmose L, Jorgensen AT, Kristiansen U, Stensbol TB, Lijefors T, Krogsgaard-Larsen P (2003) Design and synthesis of a new series of 4-alkylated 3-isoxazolol GABA(A) antagonists. Eur J Med Chem 38(4):447–449. doi:10.1016/s0223-5234(03)00056-4
Frolund B, Jensen LS, Guandalini L, Canillo C, Vestergaard HT, Kristiansen U, Nielsen B, Stensbol TB, Madsen C, Krogsgaard-Larszen P, Liljefors T (2005) Potent 4-aryl- or 4-arylalkyl-substituted 3-isoxazolol GABA(A) antagonists: synthesis, pharmacology, and molecular modeling. J Med Chem 48(2):427–439. doi:10.1021/jm049256w
Frolund B, Jensen LS, Storustovu SI, Stensbol TB, Ebert B, Kehler J, Krogsgaard-Larsen P, Liljefors T (2007) 4-aryl-5-(4-piperidyl)-3-isoxazolol GABA(A) antagonists: synthesis, pharmacology, and structure-activity relationships. J Med Chem 50(8):1988–1992. doi:10.1021/jm070038n
Ghoshal N, Vijayan RSK (2010) Pharmacophore models for GABAA modulators: implications in CNS drug discovery. Expert Opin Drug Discov 5(5):441–460
Golbraikh A, Tropsha A (2002) Beware of q2! J Mol Graph Model 20(4):269–276
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26(5):694–701
Guasch L, Sala E, Valls C, Mulero M, Pujadas G, Garcia-Vallvé S (2012) Development of docking-based 3D-QSAR models for PPARgamma full agonists. J Mol Graph Model 36:1–9
Korpi ER, Sinkkonen ST (2006) GABAA receptor subtypes as targets for neuropsychiatric drug development. Pharmacol Ther 109(1–2):12–32. doi:10.1016/j.pharmthera.2005.05.009
Korpi ER, Grunder G, Luddens H (2002) Drug interactions at GABAA receptors. Prog Neurobiol 67(2):113–159
Krehan D, Storustovu SI, Lijefors T, Ebert B, Nielsen B, Krogsgaard-Larsen P, Frolund B (2006) Potent 4-arylalkyl-substituted 3-isothiazolol GABA(A) competitive/noncompetitive antagonists: synthesis and pharmacology. J Med Chem 49(4):1388–1396. doi:10.1021/jm050987l
Lan P, Chen WN, Huang ZJ, Sun PH, Chen WM (2011) Understanding the structure-activity relationship of betulinic acid derivatives as anti-HIV-1 agents by using 3D-QSAR and docking. J Mol Model 17(7):1643–1659
Li YS, Zhou L, Ma X (2012) Molecular docking and 3D QSAR studies of substituted 4-amino-1H-pyrazolo 3,4-d pyrimidines as insulin-like growth factor-1 receptor (IGF1R) inhibitors. Med Chem Res 21(10):3301–3311. doi:10.1007/s00044-011-9877-9
Lu P, Wei X, Zhang R (2010) CoMFA and CoMSIA 3D-QSAR studies on quionolone caroxylic acid derivatives inhibitors of HIV-1 integrase. Eur J Med Chem 45(8):3413–3419
Mehta N, Chand S, Bahia MS, Silakari O (2012) Elaboration of new anti-inflammatory agents using pharmacophore based 3D QSAR of 4, 5-diaryl imidazoline as P2X(7) receptor antagonists. Lett Drug Des Discov 9(2):185–198
Møller HA, Sander T, Kristensen JL, Nielsen B, Krall J, Bergmann ML, Christiansen B, Balle T, Jensen AA, Frølund B (2010) Novel 4-(Piperidin-4-yl)-1-hydroxypyrazoles as γ-aminobutyric AcidAReceptor ligands: synthesis, pharmacology, and structure–activity relationships. J Med Chem 53(8):3417–3421. doi:10.1021/jm100106r
Nagamani S, Muthusamy K, Kirubakaran P, Singh KD, Krishnasamy G (2012) Theoretical studies on benzimidazole derivatives as E. coli biotin carboxylase inhibitors. Med Chem Res 21(9):2169–2180. doi:10.1007/s00044-011-9738-6
Nair SB, Teli MK, Pradeep H, Rajanikant G (2012) Computational identification of novel histone deacetylase inhibitors by docking based QSAR. Comput Biol Med 42(6):697–705
Neaz M, Pasha F, Muddassar M, Lee SH, Sim T, Hah JM, Cho SJ (2009) Pharmacophore based 3D-QSAR study of VEGFR-2 inhibitors. Med Chem Res 18(2):127–142
Olsen RW, Sieghart W (2009) GABAA receptors: subtypes provide diversity of function and pharmacology. Neuropharmacology 56(1):141–148. doi:10.1016/j.neuropharm.2008.07.045
PHASE 3.1 (2009) Usermanual. Schrödinger. LLC, New York
Reddy KK, Singh SK, Dessalew N, Tripathi SK, Selvaraj C (2012) Pharmacophore modelling and atom-based 3D-QSAR studies on N-methyl pyrimidones as HIV-1 integrase inhibitors. J Enzyme Inhib Med Chem 27(3):339–347. doi:10.3109/14756366.2011.590803
Sander T, Frolund B, Bruun AT, Ivanov I, McCammon JA, Balle T (2011) New insights into the GABA(A) receptor structure and orthosteric ligand binding: receptor modeling guided by experimental data. Proteins-Struct Funct Bioinf 79(5):1458–1477. doi:10.1002/prot.22975
Schwab CH (2010) Conformations and 3D pharmacophore searching. Drug Discov Today Technol 7(4):e245–e253. doi:10.1016/j.ddtec.2010.10.003
Sun X, Li Y, Li W, Xu Z, Tang Y (2011) Computational investigation of interactions between human H2 receptor and its agonists. J Mol Graph Model 29(5):693–701. doi:10.1016/j.jmgm.2010.12.001
Telvekar VN, Chaudhari HK (2012) 3D-QSAR and docking-based combined in silico study on C-5 methyl substituted 4-arylthio and 4-aryloxy-3-iodopyridin-2-(1H)-one as HIV-1 RT inhibitors. Med Chem Res 21(8):2032–2043. doi:10.1007/s00044-011-9720-3
Vestergaard HT, Cannillo C, Frølund B, Kristiansen U (2007) Differences in kinetics of structurally related competitive GABAA receptor antagonists. Neuropharmacology 52(3):873–882. doi:10.1016/j.neuropharm.2006.10.004
Weber KC, Salum LB, Honório KM, Andricopulo AD, da Silva ABF (2010) Pharmacophore-based 3D QSAR studies on a series of high affinity 5-HT1A receptor ligands. Eur J Med Chem 45(4):1508–1514. doi:10.1016/j.ejmech.2009.12.059
Zhu H, Fang H, Cheng X, Wang Q, Zhang L, Feng J, Xu W (2009) 3D-QSAR study of pyrrolidine derivatives as matrix metalloproteinase-2 inhibitors. Med Chem Res 18(8):683–701
Acknowledgments
We thank for the financial supports from National Natural Science Foundation of China (21172070), National Key Technology R&D Program of China (2011BAE06B05), National High Technology Research Development Program of China (2011AA10A207), National Basic Research Program of China (2010CB126100), and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhang, W., Xia, S., Ye, J. et al. Structural features of GABAA receptor antagonists: pharmacophore modeling and 3D-QSAR studies. Med Chem Res 22, 5961–5972 (2013). https://doi.org/10.1007/s00044-013-0583-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-013-0583-7