Abstract
Antiviral responses of interferons (IFNs) are crucial in the host immune response, playing a relevant role in controlling viralw infections. Three types of IFNs, type I (IFN-α, IFN-β), II (IFN-γ) and III (IFN-λ), are classified according to their receptor usage, mode of induction, biological activity and amino acid sequence. Here, we provide a comprehensive review of type I IFN responses and different mechanisms that viruses employ to circumvent this response. In the first part, we will give an overview of the different induction and signaling cascades induced in the cell by IFN-I after virus encounter. Next, highlights of some of the mechanisms used by viruses to counteract the IFN induction will be described. And finally, we will address different mechanism used by viruses to interference with the IFN signaling cascade and the blockade of IFN induced antiviral activities.
Similar content being viewed by others
Introduction
Innate immune responses are the first host defense against viral infections. Conserved pathogen structures are recognized by pattern recognition receptors (PRRs) on the host cells [1], that recruit a variety of adaptor proteins to signal downstream and activate the IFN response. The IFN system is present in all vertebrates and is central to antiviral responses [2]. IFNs are classified into three different families according to their receptor usage, mode of induction, biological activity and amino acid sequence [3]: type I, type II and type III IFNs. Type I IFNs, originally identified by their antiviral activity [4], include multiple IFN-α subtypes (13 in humans and 14 in mice), and a single IFN-β, and IFN-ε, IFN-κ, IFN-ω (humans) and IFN-ξ (mice) subtypes [5]. In mammals, type I IFN (IFN-I) response is essential for innate antiviral responses. They all bind to the same ubiquitously expressed receptor, IFNAR receptor, but they differ in their biological functions, due partially to the different binding affinities to the IFNAR receptor [6]. This differences in affinity results in different downstream signaling cascades [7]. For IFN-α subtypes, the quantity of the receptor on the surface of a target cell correlates also with their biological activities suggesting that the amount of IFNAR expression might compensate the weak affinity of some IFN-α subtypes [6, 8].
Type II IFNs include only one member, IFN-γ, secreted by activated T cells, natural killer (NK), NKT cells and dendritic cells with pro-inflammatory and immunomodulatory functions [9]. In general, type II IFN acts as a link between the innate immune response and the activation of the adaptive immune response [10]. Type III IFNs include IFN-λ1, IFN-λ2 and IFN-λ3, and, although genetically different to type I IFNs and signaling through different receptors, they are induced by PPRs and activates antiviral pathways similar to type I IFNs [11, 12].
Viruses use multiple mechanisms to by-pass the host IFN responses so that they can replicate and continue their infectious cycle. The present review will focus on how viruses interfere with IFN-I responses. Viruses can act at different levels of the signaling cascades involved in IFN-I responses. They can inhibit the induction of the IFN response, block the IFN signaling, and/or interfere with the antiviral activities induced by IFN. We will review some of the emerging themes and new insights from the past years of the IFN evasion mechanism employed by viruses in these contexts.
Type I IFN induction
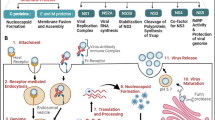
The antiviral state of an infected cells is attained by the initial induction of type I IFN expression, followed by IFN signaling transduction, which finally leads to the expression of multiple genes (Fig. 1). IFNs are the main group of cytokines secreted by host cells in response to the presence of “aberrant” nucleic acids such as double-stranded RNA (dsRNA) molecules generated as viral intermediates during viral transcription in infected cells, to CpG DNA, or uncapped ssRNA with 5′ triphosphate present in some viruses. These elements are known as pathogen-associated molecular patterns (PAMP) [13, 14] that can be recognized by PRRs. Four main types of PRRs have been described to detect virus-derived genetic materials: Toll-like receptors (TLRs) 3/7/8/9 [15]; retinoic acid-inducible gene-I (RIG-I) like receptors (RLRs) which include the cytosolic sensor RIG-I, the melanoma differentiation-associated factor 5 (MDA-5) and laboratory of genetics and physiology (LGP2) [16]; and nucleotide oligomerization domain-like receptors (NLRs) [17] and the cytosolic DNA sensors [18]. For a host to establish an antiviral state it first requires the production of type I (α/β) IFNs in direct response to virus infection and recognition of virus-derived genetic material. Hence, TLR-7 and 8 become active with the detection of ssRNA in the acidic compartment of endosomes [19] while TLR3 and 9 are receptors for dsRNA and unmethylated DNA, respectively [20].
IFN-I induction and viral counteracting actions. Viral motifs (e.g. dsRNA), are recognized by PPRs. This leads to the activation of adaptor proteins such as NFκB, TBK-1, IKKε and AP-1. TBK-1 and IKKε phosphorylate IRF3, which translocate to the nucleus and induce IFN-I expression. In a second IFN signaling wave, IRF7 is phosphorylated and translocated to the nucleus, creating an amplification loop of IFN-I induction. Several viruses have developed mechanisms to block several steps of this IFN induction cascade, indicated in red blades in the figure
Both RIG-I and MDA-5 sense cytosolic dsRNA. RIG-I also specifically senses 5′ triphosphate RNA generated during infection, while MDA-5 detects longer dsRNA sequence generated during virus replication. RIG-I and MDA5 contain two caspase activation and recruitment domains (CARDs) at the N-terminal. Activation of RIG-I and MDA5 liberates these CARD domains and drives the interaction of these tandem CARDs with the CARD of the mitochondrial activation signaling (MAVS) protein [21]. MAVS aggregates in filaments that provide a platform for the recruitment of the elements involved in the subsequent signaling cascade such as the tumor necrosis factor receptor-associated factor (TRAF) 3 and TRAF6, the TANK binding kinase 1 (TBK1) and IKKε ultimately drives the activation of transcription factors as IRF3/7, NFκB and AP-1, leading the production of type I IFN and pro-inflammatory cytokines [22]. IRF3 is constitutively expressed in many cell types, and after phosphorylation, IRF-3 forms a homodimer that translocates into the nucleus, activating the transcription of early transcribed type I and III IFN genes, IFN-β, IFN-α4 and IFN-α1 [23, 24].
The activation of IRF-7, expressed only in B cells, monocytes and plasmacytoid dendritic cells (pDCs) [25], with high levels of expression on pDCs [26] requires a second IFN signaling wave to be activated by phosphorylation. IRF7 induces the transcription of a set of delayed IFNα genes [24]. This creates an amplification loop in which type I IFN induces more IRF7, leading to the production of more IFN, with an important role in the generation of a potent response to viral infections [26].
The aberrant presence of DNA can also be sensed by PRRs during viral infection. Two main sensors for DNA during viral infection have been described so far: the IFN-γ inducible protein 16 (IFI16) and the cGAS cyclic-GMP-AMP synthetase (cGAS) (reviewed in [18]). IFI16 appears to have a preferential affinity for quadruplex DNA structures [27] that can be found in some viral genomes [28]. cGAS recognizes the presence of DNA in the cytosol. This includes viral, bacterial, or leaked cellular DNA [29, 30]. Upon the recognition cGAS catalyze the production of cGAMP that interacts with the stimulator of IFN gene (STING). STING possesses a pocket where it binds cyclic dinucleotides such as cGAMP resulting in conformational changes that lead to activation [31]. Once activated, STING acts as an adaptor protein located in the ER that traffics through the Golgi to perinuclear regions. During trafficking, STING recruits and activates TBK1, which leads to IRF3 activation and type I IFN induction [32]. While cGAS activation of STING involves the second messenger cGAMP, IFI16 appears to interact with STING to induce TBK1-dependent IFN induction [33].
IFN signaling in antiviral defense
All type I IFNs signal through the same heterodimeric transmembrane receptor termed the IFNα receptor (IFNAR), containing the subunit 1 and 2 (IFNAR1 and IFNAR2). In a first step, IFN-I binds with high affinity to IFNAR1, and then recruit IFNAR2 [34]. IFNAR engagement with IFN-I promotes the induction of an antiviral state in cells. This involves the upregulation of products from a large subset of genes named IFN-stimulated genes (ISG) that protect the cell from viral replication. Broadly speaking, ISG products modulate and mediate IFN activity in the cells. This includes for instance cooperating in PRR recognition of viral PAMPs, stabilizing signaling complexes to improve their resistance to degradation, stopping virus entry, blocking viral capsid formation, impairing trafficking and budding of virions from the infected cells, but also modulating the IFN response to avoid the toxicity of these potent immune mediators. An important feature of the IFN signaling is the rapid speed at which the response happens, which is possible because protein synthesis is not required in an initial stage.
The interaction of type I IFNs with their universally expressed receptor (IFNAR) elicits an intracellular signaling cascade through the Janus protein kinase (JAK) family members, JAK and Tyk2, that successively phosphorylate signal transducer and transcription activator (STAT) family proteins [35]. The phosphorylated STAT1/STAT2 heterodimer associates with interferon regulatory factor 9 (IRF9) to form the transcriptional factor complex ISGF3, which translocate to the nucleus and binds the IFN-response elements (ISRE) in ISG promoters leading to the expression of ISG products [36] (Fig. 2). ISGs target different steps of viral replication, amplifying the IFN signaling cascades to strength the antiviral activities [37]. Hence, the ISGs can exert a complex and wide range of functions, providing a significant redundancy in the system that fights against viral infections (reviewed in [38]).
Type I IFN signaling and viral countermeasures. IFN binds to the receptor IFNAR and triggers a signaling cascade that is summarized in this figure. Viruses has developed strategies to counteract different steps on this signaling cascade. It is marked in red blades the main signaling targets of viruses
The so-called classical ISGs pathways belong to one of the following three gene families: Mx proteins, 2′,5′-oligoadenylate synthetase (2-5OAS) or ds RNA-activated protein kinase (PKR) [39, 40]. Evidences of the role of these genes in establishing an antiviral state came from studies of infected knock out mice for PKR, 2-5OAS or Mx, individually and/or together [41]. The Mx family GTPases includes Mx1 and Mx2 proteins that function as an inhibitor of viral entry. Mx1 acts before genome replication at a very early time of the virus life cycle, inhibiting the replication of several viruses belonging to Orthomyxo-, Paramyxo-, Rhabdo- and Bunyaviridae families by interfering with the traffic or synthesis of the viral ribonucleoprotein complexes [42]. Mx2 acts as an antiretroviral protein by inhibiting chromosomal integration, a key event in the retrovirus life cycle [43].
The oligoadenylate synthetase (OAS)-latent RNase (RNase L) pathway is another IFN-inducible pathway that provides the cell with an effector mechanism upon recognition of viral dsRNA (reviewed in [44]). When the OAS senses dsRNA activates the production of 2′,5′-oligoadenylates that act as a second messenger on the inactive monomeric RNaseL [45]. The 2′,5-oligoadenylates binding to RNase L produces a catalytically active dimer that cleaves ssRNA [46]. This leads to the translational arrest and prevent viral replication and spreading [47].
RNA-activated protein kinase is an ISG product that detects cytosolic dsRNA. PKR recognition of its dsRNA substrate leads to dimerization and autophosphorylation which in turn leads to the phosphorylation of the eukaryotic initiation factor 2α (eIF2α) required for translation initiation [48]. eIF2α phosphorylation results in the shutdown of all translation of 5′capped mRNA, thus preventing the synthesis of viral proteins. This also usually results in the formation of stress granules (SG) that consist of the accumulation of RNA and proteins from the stalled translation complexes. SG formation has been linked to antiviral responses, and their formation is often inhibited in viral infections. In general, the antiviral activity of PKR is related to apoptosis induction [49], regulation of IFN-β synthesis and NF-κΒ pathway [50,51,52], serine kinase activity for STAT1 that also regulate the IFN-I signaling pathway [53].
A family of proteins with a wide range of anti-viral functions are the interferon-induced proteins with tetratricopeptide repeats (IFITs) [54]. These genes are expressed at very low basal levels, but their transcription is rapidly increased after activation by IFN signaling. IFIT detect the lack of 2′-O-methylation on RNAs species, a methylation absent in some viral RNA but present in eukaryotic mRNA [55]. IFIT1 has also been shown to bind to the 5′-triP end of some viral RNA [56]. IFIT1 can sequester viral RNA or interact with the eukaryotic translation initiation factor 3 to inhibit the translation initiation of IFIT1-bound RNA species.
In addition to these ISGs, one highly upregulated gene in the initial stage of the antiviral immune response is ISG15 (interferon-stimulated gene 15), which encodes an ubiquitin-like protein involved in a post-translational process termed ISGylation [57]. This process allows ISG15 to bind covalently to a range of target proteins, both viral and cellular [58], by a process that is reversible due to the action of the ubiquitin-specific protease 18 (USP18), an event regulated by type I IFN [59]. ISGylation appears to modulate the activity of multiple elements involved in the IFN response. For instance, ISGylation has been shown to sustain STAT1 or IRF3 activity [60, 61], downregulate the turnover of ubiquitinated proteins [62], but also to negatively regulate RIG-I signaling [63]. ISG15 acts during viral replication by interfering with the endogenous proteins that the virus needs to replicate. Thus, ISG15 conjugates to the eukaryotic factor 4E (eIF4E) homologous protein (4EHP) that binds to the capped mRNA, inhibiting in this way the viral RNA translation of those viruses that contains a capped positive-sense RNA such as flaviviruses [64]. ISG15 also exists as an unconjugated protein that acts as a cytokine, regulating viral replication and host responses [65, 66].
The role of the ISGs members mentioned above clearly illustrates the breadth and diversity in the function of this protein group. A database has been created, called Interferome (https://www.interferome.org/interferome/home.jspx), in which ISGs are catalogued and incorporated into a database, based on the information obtained from all published reports where cells were treated with IFN, Thus, this database will allow to identify ISG signatures from high-throughput data, having implications for determining the role of ISGs. Viruses employ mechanisms that impair ISG activity to enhance their evasion from the IFN system. The mechanisms will be discussed in another section of this review.
Viral evasion strategies: inhibition of interferon induction
As discussed previously, PRR activation leads to the production of IFN-I, -III, and pro-inflammatory cytokines such as IL-1β. The present section will be centered on how viruses affect IFN-I induction. Typically, virus genetic material triggers IFN-I induction when recognized by viral nucleic acid sensors that are membrane bound or present in the cytoplasm/nucleus. This recognition leads to activation of signaling cascades that converge towards the induction of IFN-I production in infected cells. Viruses are known to interfere at every point of this process. Viruses can interfere with the sensing of their genetic material, impair the signaling cascade that leads to IFN-I induction, and/or antagonize the activity of the transcription factors involved in IFN-I gene expression. The present section aims at providing a non-exhaustive overview of some of the most commonly used viral mechanisms to counter IFN-I induction in the host with a focus on recent findings in the field.
Virus interference with viral sensors of genetic material
Viruses use different mechanisms to counteract the recognition of their genetic material by the host cell so that IFNs are not induced (Table 1). Viruses can sequester, modify or even degrade their nucleic acids to avoid detection by PRRs. For instance, during replication most flaviviruses create vesicular structures in the ER membrane which physically shield the viral genetic material from cytosolic RLRs [67,68,69]. Influenza A virus (IAV) uses the nucleus for replication, atypically for an RNA virus, so that its genetic material remains hidden from cytosolic RLRs [70]. Vaccinia virus (VV), a large double-stranded DNA virus has the peculiarity of replicating in the cytoplasm, where DNA sensors like cGAS are present, which potentially renders the viral genetic material susceptible to recognition by PRR. VV replication, however, occurs in organelles similar to micronuclei [71] that probably render viral DNA inaccessible to recognition by cytoplasmic DNA sensors. Rotaviruses (RV) concentrate their replication in a cytoplasmic structure called viroplasms where dsRNA genome is generated for packaging so that it is not exposed to the cytoplasmic PRR [72].
Many viruses also encode proteins that help conceal their genetic material from PRR. Some viruses possess dsRNA binding proteins that could potentially sequester these PAMPs from PPR recognition, such as VP35 from Ebola virus or σ3 from reovirus [73, 74]. The encapsidation of the viral RNA can also impair RLR recognition. For instance, IAV nucleoprotein and polymerase prevents RIG-I binding to viral RNA during transit through the cytoplasm [75]. IAV NS1 protein also possesses a dsRNA binding site that prevents recognition by RIG-I [76]. Calicivirus and picornavirus ssRNA ( +) is covalently linked to a capping protein that could prevent recognition of the 5′ viral RNA extremity by RIG-I [77]. Lassa virus (LASV) nucleoprotein (NP) can act as a capping enzyme with exonuclease activity specific for dsRNA, which has been shown to antagonize IFN induction [78, 79]. Other viruses can modify their 5′tri-P motifs recognized by RIG-I to evade this cytoplasmic RNA sensor. Hantaan virus, Crimean-Congo hemorrhagic fever virus and Borna disease virus can process their 5′ genome extremity to form 5′mono-P forms, evading RIG-I recognition [80]. Poxvirus decapping enzymes D9 and D10 can prevent the accumulation of dsRNA, an intermediate necessary in viral replication, and thereby evade RLR recognition [81]. Measles virus (MeV) encodes for the non-structural C protein that can impair IFN response by modulating viral RNA replication [82] and improving the polymerase processivity [83], thus probably limiting the amount of viral material recognizable by cytosolic PRR.
Another mechanism employed by viruses to limit detection by PRR is to interact with these sensors to impair their activation. The Kaposi’s sarcoma-associated virus (KSHV) uses the tegument protein ORF52 to bind to cGAS and inhibit cGAMP production, the second messenger used for STING (stimulator of interferon response cGAMP interactor 1) activation [84]. Homologues of ORF52 in other gammaherpesviruses have also been described to act similarly, indicating that inhibition of this PRR pathway is probably shared by gammaherpesvirus. The Herpes simplex virus 1 (HSV-1) tegument protein VP22 has also been shown to inhibit cGAS enzymatic activity, indicating that other Herpesviridae can directly target cGAS [85]. Other viruses can sequester PRR so that they are unable to relocate to their activity site. For instance, the protein Z of new world arenaviruses binds to RIG-I and prevents its association with the signaling platform MAVS protein [86]. Severe acute respiratory syndrome coronavirus (SARS-CoV) M protein has been shown to associate with RIG-I and can potentially sequester this PRR [87].
Viruses can also promote PRR degradation, thus reducing the number of cellular sensors capable of detecting viral infection (Fig. 3). This can be done directly by proteases encoded by the viral genome. Foot and mouth disease virus (FMDV) Lpro and 3Cpro protease can reduce RIG-I intracellular protein levels [88]. The 3Cpro protease of other picornaviruses has also been shown to degrade RIG-I [89], indicating that this is a shared mechanism of RLR evasion by this viral family. RIG-I is not the sole PRR that picornavirus proteases can target; the poliovirus and enterovirus 71 (EV71) 2Apro protease can also degrade MDA5 [90]. Viruses also encode for proteins that indirectly promote PRR degradation. The nuclear sensor of DNA IFI16 is degraded during HSV-1 infection through a mechanism dependent on the viral ICP0 protein that is not fully understood but probably involves targeting the DNA sensor for proteasomal degradation [91]. A E3 ligase activity on the NSs protein of the phlebovirus Toscana virus was recently identified that allowed the ubiquitination of RIG-I CARD domains and the subsequent proteasomal degradation of the PRR [92]. The NS2B protein of the flavivirus Dengue virus (DENV) can target the cytosolic DNA sensor cGAS for lysosomal degradation. Although this mechanism could at first glance seem counterproductive for an RNA virus, it actually allows DenV to evade the recognition of the mitochondrial DNA that becomes exposed during infection [93].
Viral interference with accessory cellular components involved in PRR activation. MeV can interfere with RLR activation by targeting the phosphatase PP1 using 2 distinct mechanisms. MeV V protein can interact with PP1 to prevent the dephosphorylation of MDA-5 required for activation. MeV can interact on the cell surface with the C-lectin receptor DC-SIGN which results in the association of PP1-inhibitor 1 with PP1 thus preventing RLR dephosphorylation. Ebola virus VP35 protein, MERS-CoV 4a protein and arenavirus NP can interfere with PACT binding to dsRNA, a mechanism that potentiates RLR activation. RLR ubiquitination is also essential for adequate activation and transport to MAVS for subsequent IFN signaling events to take place. Riplet and TRIM25 are critical to RIG-I ubiquitination. IAV-NS1 and Denv sfRNA can interfere with TRIM25 activity, whereas Hepatitis C NS3-4A protease can cleave Riplet to impair RIG-I ubiquitination. The mitochondrial-targeting chaperone 14-3-3ε is responsible for RIG-I translocation to the mitochondrial membrane. DenV NS3 protein targets 14-3-3ε using a phosphomimetic domain that displaces activated RIG-I from this chaperone. WNV NS3 possesses a similar domain
Other viruses alter the post-translational modifications essential to PRR signaling so that the PPR remains inactive. Some viral-encoded proteins achieve this directly, as in the case of the deubiquitinase activity of coronaviruses papain-like protease (PLP) that removes the K63 ubiquitin tail from RIG-I that is essential for RIG-I translocation to the MAVS protein platform [94]. This deubiquitination activity has been characterized in several coronaviruses thus suggesting that this mechanism is central to coronavirus evasion from the IFN system [95,96,97]. Paramyxovirus V protein binds to MDA5 and impairs its dephosphorylation by blocking the ATP hydrolysis necessary for MDA5 folding to its active state, thus impairing the adequate activation of this PRR [98].
Viruses can also impair PRR activity by interfering with the functionality of accessory cellular components required for PRR activation. MeV has been shown to act on the phosphatase PP1 required for RLR activation using two distinct mechanisms. The MeV V protein can bind PP1 which prevents MDA5 dephosphorylation [99]. MeV infection in dendritic cells also produces recognition of the viral particle through the C-lectin receptor DC-SIGN [100]. This triggers a signaling cascade that results in Raf-1 kinase activation and the association of the PP1 inhibitor I-1 with PP1 that prevented RLR dephosphorylation thus impairing IFN induction. Several viruses have also been shown to interact with the dsRNA binding protein PACT that potentiates RLR activation [101, 102]. For instance, Ebola virus VP35 protein, Middle East respiratory syndrome coronavirus (MERS-CoV) 4a protein and arenavirus nucleoproteins have been shown to interfere with PACT binding to RLR [103,104,105]. As previously stated, RIG-I activation is associated to its ubiquitination with K63 Ub chains that liberates its autorepressed N-terminal CARD domains, a mechanism dependent on the activity of the cellular proteins TRIM25 and Riplet [106, 107]. This mechanism can be targeted by viral products such as the IAV NS1 protein that impairs TRIM25-mediated RIG-I ubiquitination [108, 109], or the hepatitis C virus NS3-4A protease that cleaves Riplet [106]. Intriguingly, not only protein products appear to interfere with the RIG-I activation complex. The subgenomic flavivirus RNA (sfRNA) generated during DenV infection can bind to TRIM25 and prevent the deubiquitination step required for TRIM25 activity to take place [110]. DenV uses yet another mechanism to avoid PRR detection. DenV NS3 protein possesses a phosphomimetic domain that binds the mitochondrial-targeting chaperone protein 14-3-3ε [111]. 14-3-3ε is responsible for RIG-I translocation to the mitochondrial membrane where the subsequent steps of the IFN induction signaling cascade take place. By binding 14-3-3ε, DenV NS3 displaces the activated RIG-I complex and prevents IFN induction. Interestingly, a similar phosphomimetic domain is also present in West Nile Virus (WNV) NS3 protein [111].
Virus interference with the IFN-I induction signaling cascade
Viruses not only interfere with the PRR capable of detecting their presence during infection, they also commonly affect the activity of the signaling complexes in charge of signal transduction. Indeed, this is a strategy employed by most viruses to limit IFN responses. Viruses can antagonize these signaling cascades at multiple levels and through varied mechanisms (Table 2). This can be achieved by impairing the post-translational modifications required for signaling, i.e. by altering the phosphorylation or ubiquitinylation status of signaling intermediates. For instance, SARS-CoV and human coronavirus NL63 (HCoV-NL63) PLPs have been shown to prevent STING dimerization and thus subsequent activation of the TBK1 pathway possibly through the PLP deubiquitinase activity [112, 113], as STING dimerization is dependent on the attachment of K63Ub chains [114]. Similarly, SARS-CoV PLP can inhibit TLR7 signaling by removing K63-Ub chains from TRAF3 and TRAF6 and thus blocking TBK1 activation [115]. Another strategy to prevent post-translational modification of IFN-I pathway signaling components consists of sequestering these components so that they do not reach the adequate cellular location for activation. The NS3 protein from the economically important orbivirus Bluetongue Virus (BTV) binds to the ubiquitin-binding protein optineurin in the Golgi apparatus [116]. This prevents optineurin from recruiting ubiquitinated TBK1 at the Golgi apparatus, a necessary step for subsequent TBK1 phosphorylation to occur [116]. Recently, it has been described that cGAMP, the second messenger generated by cGAS and that activates STING, can be cleaved by poxvirus-encoded nucleases (named poxins) [117]. This allows poxvirus blockade of the cGAS-STING signaling axis.
Viral proteins can also prevent the adequate formation of signaling complexes by steric hindrance. For instance, human adenovirus type 5 E1A protein has been shown to bind to STING and thus antagonize IFN signaling [118]. MERS-CoV and SARS-CoV M proteins can interact with TRAF3 and thus disrupt the TRAF3-TBK1 association [87, 119, 120]. MERS-CoV ORF4b protein can associate with the TBK1/IKKε complex to impair signaling [121], indicating that coronaviruses encode for multiple proteins that affect IFN induction at different stages of the signaling cascade. KSHV encodes for viral interferon regulatory factor 1 (vIRF1), a protein that binds to STING and block the association of TBK1 with STING, thus hindering the consequent IRF3 phosphorylation [122]. HSV-1 encodes for the ICP27 protein that can interact with the active STING-TBK1 complex and inhibit the subsequent TBK1-mediated phosphorylation of IRF3 [123]. The NSs protein from some phleboviruses has also been described to antagonize IFN induction by targeting TBK1 [124]. TBK1 activity is thus commonly targeted by both RNA and DNA viruses to antagonize IFN induction.
Viruses can also promote the degradation of signaling proteins involved in IFN induction. Virally-encoded proteases can directly cleave some of the components of these pathways. STING can be directly cleaved by the NS2B3 protease from several flaviviruses such as DenV, Zika virus, WNV, or Japanese encephalitis virus (JEV), but not others like yellow fever virus [125,126,127]. Indeed, this specific cleavage could partially explain some of the host range and pathogenicity of these flaviviruses in humans, as the identified STING cleavage site for the NS2B3 protease is only partially conserved among species [125]. Similarly, the 3Cpro from several Picornoviridae has been described to cleave MAVS proteins, thus inhibiting signal transduction [128, 129]. The 3C-like protease from the arterivirus porcine reproductive respiratory syndrome virus (PRRSV) and the NS3/4A protease complex from the flavivirus hepatitis C virus (HCV) have also been described to cleave MAVS protein [130, 131]. Other viruses encode proteins that promote the degradation of signaling complexes involved in the IFN responses. RV NSP1 and VP3 proteins have been shown to target MAVS protein for proteasome-dependent degradation and thus impair IFN induction [132, 133]. SARS-CoV ORF9b protein localizes in the mitochondrial membrane where it interacts with MAVS protein and promote its degradation [134]. This mechanism probably involves the recruitment of the MAVS protein cellular regulator PCBP2 [135] to the mitochondrial membrane by SARS-CoV ORF9b protein, which favors MAVS protein ubiquitination through K48-Ub chains and thus subsequent proteasomal degradation.
Since MAVS protein represents an important signaling platform in the IFN induction cascade triggered by RLR activation, some viruses use strategies to alter mitochondria structure to impair MAVS protein assembly. DenV NS2B3 protease can cleave mitofusins, which alters mitochondria dynamics and impair their fusion [136]. In other cases, viruses can also promote mitophagy to promote their replication and potentially impair IFN responses. Mitophagy has been described in MeV, hepatitis B virus (HBV) or Newcastle disease virus infections [137,138,139]. The protein M from the human parainfluenza virus 3 has also been shown to promote mitophagy and thus trigger MAVS protein degradation to antagonize IFN induction [140]. Recently, IAV has also been shown to employ a similar mitophagic strategy to reduce MAVS protein levels through the expression of its PB1-F2 protein [141].
Virus interference with IFN-I transcription factors
Viruses can also act on the transcription factors that bind to the IFN-I promoter and trigger the expression of ISGs. Viruses use multiple strategies to impair the binding of the transcription factors to the IFN-I promoters (Table 3). They can impair the phosphorylation by the TBK1/IKKε complex of IRF3/7 that activates dimerization and transport to the nucleus. For instance, the paramyxovirus C protein inhibits IRF7 phosphorylation and thus its activation [142]. The DenV NS2B3 protease can impair IRF3 phosphorylation by masking the IKKε kinase domain necessary for IRF3 activation [143]. Lymphocytic choriomeningitis virus (LCMV) nucleoprotein was also shown to bind to the IKKε kinase domain, thus preventing IRF3 phosphorylation [144] indicating that arenavirus NP-mediated inhibition of IRF3 phosphorylation [145] uses a similar mechanism.
Viruses not only target IRF3/7 activation to antagonize IFN induction, they can also interfere with the activation of NF-κB, as this transcription factor collaborates with IRF3 in the early activation of the IFN-β promoter [146]. Most arenavirus nucleoproteins not only block IRF3 phosphorylation but they can also impair NF-κB activity [147]. Viruses can also target adaptor molecules that regulate NF-κB activity. The NF-κB essential modulator (NEMO) is part of the kinase complex that controls NF-κB release from its inhibitor IκB. The 3Cpro from Hepatitis A virus or FMDV and the 3C-like proteases from porcine epidemic diarrhea virus or PRRSV cleave NEMO thus preventing IκB release from NF-κB and consequently antagonizing IFN-β production [148,149,150,151].
The degradation of the transcription factors involved in IFN induction is also the target of different viral proteins, as in the case of the NSP1 from RV that promote the degradation of several IRFs including IRF3 and 7 [152]. RV NSP1 appears to target IRFs by interacting with their dimerization domain and thus preventing their association in active form [153]. NSP1 has also been described to impair NF-κB by promoting the degradation of β-transducing repeat-containing protein (β-TrCP) [154], the protein responsible for substrate recognition of the E3 ligase complex that targets for degradation IκB, the associated inhibitor of NF-κB. Similarly, to other viral products, such as HIV-1 Vpu, VV A49, or Epstein–Barr virus LMP-1, RV NSP1 associates with β-TrCP through mimicry of the phosphorylated IκB that requires degradation [155,156,157,158]. RV NSP1, however, presents the particularity of not only blocking the interaction of IκB with the E3 ligase complex but also of targeting β-TrCP for degradation.
Viruses can also impair IRF3/7 activity downstream of their activation by phosphorylation. The vIRF1 encoded by KSHV can block IRF3 activity downstream of IRF3 activation by impairing the recruitment of the CBP-p300 coactivators to the IRF3 complex [159]. This can also be achieved by blocking the transport of activated transcription factors to the nucleus. For instance, some paramyxoviruses use their V protein to impair IRF3 translocation to the nucleus [160]. JEV NS5 blocks IRF3 and p65 subunit from NF-κB transport to the nucleus by interacting with nuclear transport proteins [161]. The NSs from the emerging bunyavirus severe fever with thrombocytopenia syndrome virus have also been described to sequester the TBK1–IKKε–IRF3 complex in viral inclusion bodies to prevent the trafficking of IRF3 to the nucleus [162].
Some viruses also code for proteins that interfere with IFN induction in the nucleus. In many instances, the exact mechanisms of IFN antagonism by viral products in the nucleus are not fully resolved. MERS-CoV ORF4b protein impairs IFN induction in the nucleus by a mechanism yet to be elucidated [121]. BTV encodes non-structural protein 4 (NS4) that localizes in cell nucleoli and possess IFN antagonist activity [163]. MeV C protein can interfere with IFN-β promoter activation in the nucleus and this property has been linked to the virus pathogenicity [164]. The NSs protein from the Phlebovirus Sandfly fever Sicilian virus can prevent IRF3 activity by interacting with the DNA binding domain of IRF3 [165]. The protein ORF36 from the murine gamma-herpes virus 68 was shown to bind to IRF3 and to prevent its association with the co-transcriptional activator CBP, thus impairing IRF3 binding to the IFN-β promoter [166].
Viruses have developed strategies to antagonize IFN induction at multiple stages of the signaling cascade. This includes limiting the recognition of their genetic material, impairing the activation of PRRs or their signaling partners, promoting the degradation of key components of the signaling cascade, sequestering signaling complexes away from their site of action, or impairing DNA binding of the transcription factors involved in IFN induction. In viruses, multiple mechanisms have also often evolved to target several elements in these pathways, and hence augment their capacity to evade innate immunity.
Viral evasion strategies: blockade of IFN signaling
Intracellular blockade of IFN signaling pathways
Viruses can suppress the IFN signaling at different levels (Table 4). In this section, we will discuss some of the mechanisms that viruses use to counteract the action of this signaling cascade.
One of the first steps in the signaling cascade is the phosphorylation of JAK1 and TYK2, relevant for initiating the JAK-STAT signaling. By directly promoting the dephosphorylation of the JAK/STAT pathway, viruses counteract the IFN response. For instance, Sendai virus (Respirovirus) C protein inhibits the phosphorylation of receptor-associated kinases JAK1 and TYK2 by binding to the IFN receptor subunit IFN-α/β [167]. The NS5 protein of the JEV blocks the tyrosine phosphorylation of TYK2 and STAT1 [168]. STAT1 phosphorylation is targeted by several viruses, using different mechanisms. The Paramyxovirus family, that includes MeV, peste des petits ruminants virus (PPRV) and mumps, express V and P proteins that interact directly with STAT1, inhibiting phosphorylation [169, 170]. The immediate-early protein ICP27 of HSV-1 downregulate STAT1 phosphorylation and prevent the accumulation of STAT1 in the nucleus [171]. The DENV proteins NS2A, NS4A, NS4B block the phosphorylation of STAT1 [172]. The NS5 protein of HCV interacts with STAT1, interfering with its phosphorylation [173]. Among the Reoviridae family, the NSP1 protein encoded by rotavirus block the phosphorylation of STAT1 [174], and the NS3 protein encoded by BTV blocks STAT1 phosphorylation [175, 176].
Many viruses target STAT2 to antagonize IFN signaling. Thus, MeV V protein binds to STAT2 [177, 178]. Yellow fever virus NS5 protein also binds STAT2 but this interaction requires STAT2 activation by IFN [179]. Other example is DENV NS5 protein acts as a bridge between UBR4 and STAT2, driving STAT2 to degradation through the proteasome [180]. The nsp11 protein of PPRS degrades STAT2 via proteasome [181]. The proteasome is not the only catabolic cellular machinery that viruses can highjack to degrade signaling components of the IFN signaling pathway. BTV was recently shown to use ubiquitination of its NS3 protein to drive STAT2 degradation by an autophagy-dependent mechanism [175].
Another strategy used by viruses includes the interference with IFN signal transduction by modification of the constitutive or basal levels of molecules involved in the JAK/STAT pathway that viruses members of the Rubulavirus genus like Simian virus 5, mumps virus or human parainfluenza virus type 2, use [182, 183]. The human Papillomavirus-16 (HPV-16) expresses the viral E7 protein that binds the p48 protein blocking its translocation to the nucleus, impeding the association of IRF-9 with the STAT-1/STAT-2 heterodimer (ISGF3), and thereby inhibiting the induction of IFN-I inducible genes [184].
Viruses involved in persistent infections, such as cytomegaloviruses (CMV), polyomaviruses, HCV [185], or HSV-1 [186] use similar strategies. CMV affects the expression levels of JAK1 and IRF-9 [187], and the viral large T antigen of murine polyomavirus binds to JAK1 inactivating the transduction signal through IFN receptors [188].
Blockade of IFN induced antiviral activities
Viruses employ mechanisms that impair ISG activity to enhance their evasion from the IFN system. In this section, we will discuss some of these mechanisms. As previously mentioned, some ISG products enhance the recognition of viral PAMPs and provide the cell with effector mechanisms that block viral replication. Thus, FMDV Lpro cleaves G3BP1, an RNA-binding protein essential to stress granules (SG) assembly, to impair the formation of these structures [189]. MeV C protein has been involved in SG inhibition through blockade of PKR-induced SG by the activity of the adenosine deaminase acting on RNA 1 (ADAR1) [190]. DENV sfRNA can bind to various components involved in SG assembly. This property of DENV sfRNA also led to impaired ISG mRNA translation, thus dampening IFN responses [191].
To overcome recognition by ISG products, some viral mechanisms are discussed. The reovirus σ3 protein was shown to inhibit PKR activity probably through its ability to bind dsRNA [192, 193]. NSs protein from bunyavirus can promote PKR degradation [194]. Poxvirus D9 and D10 decapping enzyme promote dsRNA degradation, thus preventing PKR recognition [81]. Viruses can also highjack regulatory ISG pathways to evade ISG product action. For instance, ADAR1 is an ISG involved in RLR regulation. ADAR1 has an important physiological function as it edits adenosines to inosines in RNA, a feature that destabilize the structure of complementary dsRNA strands, thus preventing RLR or PKR recognition. This is an important mechanism in the prevention of autoimmunity, as it limits the recognition of cellular dsRNA. Viruses such as MeV, VSV or HIV-1 have been shown to use ADAR1 function to block PKR activation and thus evade translation shutdown [195,196,197].
Viruses can also interfere with the OAS-RNase L pathway. Rotavirus VP3 protein blocks RNaseL activation [198] by cleaving the 2′,5′-oligoadenylates produced by OAS. The NS2 protein from coronavirus murine hepatitis virus has been shown to act similarly [199].
The IFIT is another ISG that contribute to viral RNA recognition. Some viruses have developed 2′-O methyltransferase activities on their gene products to prevent translation blockade of their RNA by IFIT. This has been described for WNV, coronaviruses, RV and poxvirus [200,201,202,203,204].
The IFN-induced transmembrane proteins (IFITM) expression is greatly enhanced upon IFN activation, but these proteins are also expressed ubiquitously in the absence of IFN. The family of IFITM proteins has been shown to block IAV, WNV and DENV cell entry, a mechanism that probably involves viral hemagglutinin recognition [205]. HCoV-OC43 has been shown to highjack IFITM2 and IFITM3 for cell entry. This mechanism could be important for virus entry in lower respiratory tract under inflammatory conditions induced by IFN [206]. In HIV-1, Vpu and Env proteins can mutate to increase infectivity and overcome IFITM1-mediated restriction of replication [207]. Mutations to overcome the activity of the ISG product MxGTPase have also been described for IAV, indicating that multiple ISG products probably exert a selective pressure on viruses. For instance, pandemic avian IAV strains appear to adapt to human through evasion of the NP recognition by MxA GTPase [208].
Viruses have evolved strategies to dampen ISG15 effects on IFN signaling. For instance, the PLP from HCoV-NL63, SARS-CoV and MERS-CoV have been shown to not only act as a deubiquitinase but also as a deconjugating protease for ISG15 chains [94, 96, 97]. FMDV Lpro has also been associated with cleavage of ISG15, but instead of targeting the isopeptide bond used in ISG15 conjugation, it hydrolyzes the peptide bond preceding the ISGylation motif [209]. Human cytomegalovirus (HCMV) tegument pUL26 protein prevents ISGylation [210]. pUL26 activity appears to be supported by two other tegument proteins pUL25 and pp65 [211]. HCMV also uses another tegument protein pUL50 to affect ISGylation by targeting UBE1L, an important ligase responsible for ISG15 linkage, for proteasomal degradation [212]. Since ISGylation also regulates the activation of IFN-related pathways, some viruses have harnessed ISGylation to favor their replication. HCV has also been described to use ISGylation to favor its replication and develop persistent infections [213, 214]. Recently, ISGylation was also associated with increased replication in HBV infections [215].
Multifaceted strategies have evolved in viruses to circumvent ISG product activity and thus enhance their replication and spreading. These range from directly impairing the activity of ISG products involved in host cellular defense to highjack the IFN modulating activity of some ISG products. Understanding these mechanisms of evasion will undoubtedly shed light on some of the pathogenic processes induced by viral infections.
Viral evasion strategies: secreted IFN binding proteins
A unique strategy to inhibit the activity of IFN was described in 1995 with the identification of a poxviral secreted IFN type I binding protein (IFN-I BP) encoded by the B18R gene of Vaccinia virus (VV) [216] a well characterized member of the Orthopoxvirus genus that contains the strains used for the efficacious worldwide vaccination campaigns against smallpox. The protein was found to be a secreted glycoprotein of about 60 kDa expressed early during infection. While its sequence is unrelated to either of the two subunits of the cellular IFN-I receptor, IFNAR1 or IFNAR2, the IFN-I BP was found to bind with high affinity (KD = 175 pM for hIFNα2) to several subtypes of human IFN-Is, inhibiting their binding to the receptor and thus abrogating their biological activity [217]. In stark contrast to the high species specificity observed for the cellular receptor, the viral protein is able to bind and inhibit the activity of IFN from different species, including mouse, rat, bovine and rabbit ligands, suggesting different interaction modes. Currently, all human IFN-I molecules tested including 8 (out of 13) IFNα, IFNβ, IFNϖ as well as the more divergent IFNκ and IFNε are known to be bound and inhibited by B18 [218,219,220], although with varying affinities. Interestingly, murine IFNα, but not murine IFNβ are inhibited by the poxviral IFN-I BP in spite of being bound with high affinity, as assessed by surface plasmon resonance [219, 221]. Competition studies using anti IFN monoclonal antibodies (mAbs) showed that the binding interface of IFNs with B18 is larger than the one with their cellular receptor, which could probably account for its broad species specificity and inhibitory capacity over a wide range of affinities [222].
Further examinations substantiated an additional property of the poxviral IFN-I BP which is its saturable binding to the cell surface after secretion, where the protein is active and can inhibit IFN as efficiently as the secreted one [221]. This suggested that the main site of action is the cell surface, providing local tissue protection by protecting neighboring, still uninfected cells from entering into an IFN mediated antiviral state. Examination of a truncated version of B18 expressed by the attenuated Wyeth VACCV strain lacking its third, C-terminal immunoglobulin (Ig) domain showed that cell binding capacity is mediated by the N-terminal regions of the protein [221]. Additional transfection analyses with different constructs suggested that cell binding activity is mediated by Ig domain 1, while IFN blocking activity requires Ig domains 2 and 3 [223] Site directed mutagenesis assays identified stretches of basic residues at the N terminus of B18 to mediate high affinity binding to cell surface sulfated glycosaminoglycans, preferentially heparan sulfate [224] and showed that mutants lacking GAG binding activity could still bind and inhibit IFN efficiently.
The IFN-I BP protein has been found to be conserved in other orthopoxviruses including cowpox virus and ectromelia virus (ECTV), a natural mouse pathogen, as well as the two viruses causing significant disease in humans, monkeypox virus and variola virus, the causative agent of smallpox [219]. Interestingly, the human viruses show an enhanced affinity for the human ligands, possibly reflecting the host adaptation of the virus, as occurs with other secreted cytokine binding receptors in this family. While possible orthologues can be readily found in virus species from several other poxvirus genera, these are frequently more distantly related, and their properties have not been extensively studied. The single exception to this is protein Y136 of the Tanapoxvirus Yaba-like disease virus (YLDV), a primate virus causing infection restricted to the skin. This protein, which shares only 27% aminoacid identity to the VACV B18, can bind and inhibit both human (and monkey) IFN- I as well as the more recently described family of type III IFNs [218]. The latter are a specialized group of IFNs mediating antiviral response specifically at mucosal sites without compromising barrier integrity of the epithelia and promoting long-lasting humoral and cellular responses which signal through a distinct, specific heterodimeric cellular receptor (reviewed by [225]). The authors have proposed that inhibition of these IFNs might be related to the specific tissue tropism of YLDV, although information on its role in vivo has not yet been provided.
Insights into the biological role of poxviral IFN-I BPs comes from murine infection models using VACCV and ECTV, the latter naturally causing fatal mousepox in susceptible mouse strains. Early reports showed that deletion of IFN-I BP gene from VACV attenuated the virus in vivo both in intranasal [216] as well as intracranial [217] infection models. In ECTV, absence of IFN-I BP resulted in a completely attenuated phenotype upon footpad inoculation (LD50 reduction at least 107-fold) with severely impaired dissemination of virus to its secondary replication sites, liver and spleen, as well as enhanced NK cell recruitment and both CD4+ and CD8+ T cell activation [226] Crucially, these effects were shown to be dependent on IFNAR signaling by the use of knockout mice. The IFN-I BP was found to bind to uninfected cells around infection foci in the liver and spleen protecting these tissues locally from IFN induced antiviral activity [227]. The biological relevance of tissue retention of this inhibitory protein was shown using recombinant ECTV that express a mutated IFN-I BP unable to bind to the cell surface but still able to inhibit IFN-I efficiently. Infection with these recombinants resulted in non-lethal infection as in the case of the virus lacking IFN-I BP altogether [228]. Interestingly, it was found that immunization of mice with recombinant IFN-I BP could prevent the development of mousepox upon challenge [226], probably through the development of antibodies capable of impairing its interaction with its ligands [227] and also pointing to a novel therapeutic target for the treatment of poxviral infections in humans.
The structure of the complex of ECTV IFN-I BP with murine IFNα-5 has been solved to high resolution (PDB entry 3OQ3, deposited by Fremont and Lee, results to be published). Comparisons with the ternary complexes of different IFN-I with their cellular receptor [229, 230] will be crucial to disentangle the structure function relationships in the interaction and inhibition of the biological activities of IFN-I ligands by the poxviral IFN-I BPs.
The particular properties of the poxviral IFN-I BP, especially its broad species and type specificity as well its high affinity have been instrumental to its use as a biotechnological tool. Thus, B18 has been used to determine the implication of IFN-I in diverse processes, such as the monocyte-derived macrophage-mediated inhibition of human cytomegalovirus (HCMV) spread [231]. In addition, recombinant oncolytic herpes simplex viruses expressing B18 have been developed to improve their infectivity in the face of antiviral responses [232]. Finally, B18 has been used to block IFNα mediated HIV associated encephalitis in a murine model [233] or to inhibit the detrimental IFN mediated effects produced by mRNA exposure in induced pluripotent stem (iPS) cell reprogramming [234].
Secreted IFN-I BPs have been exclusively found in poxviruses to date. A recent report described the murine norovirus NS1 protein, which is secreted by an unconventional caspase-3 mediated pathway, to be essential for tuft cell infection in gastrointestinal tissue through blockade of IFN type III signaling [235]. While a direct inhibition of IFN type III could not be demonstrated in the reporter assay used, the molecular mechanism employed by this protein remains unsolved and raises the question as to whether additional and different soluble IFN-I or IFN-III BPs might be identified in other virus species.
Concluding remarks
IFN responses are a complex and important component of the innate immune system. This is reflected in the vastness and complexity of ISGs roles, not only involved in antiviral responses but also in several immunomodulatory functions. Viruses can disrupt IFN responses leading to the antiviral state to promote their successful replication. Indeed, viruses often interfere with multiple pathways involved in the IFN response to evade innate immunity. The importance of the IFN system in host antiviral responses is highlighted by the fact that viruses dedicate some of their genetic material to encode for IFN antagonists. The viral mechanisms of IFN evasion can be mediated directly by viral gene products. Viruses also often usurp components of the cellular machinery to carry out their IFN antagonistic activity. There is no doubt that understanding how viruses evade the IFN system will shed some light on the pathogenicity and allow for a better design of therapeutic approaches. The interaction of these pathogens with the IFN system can also shed some light on some of the regulatory cellular mechanisms that control the IFN response. Studying the interaction of viral components with the IFN system remains essential to understand the pathogenesis of emergent viruses that threatened global health.
References
Wu J, Chen ZJ (2014) Innate immune sensing and signaling of cytosolic nucleic acids. Annu Rev Immunol 32:461–488. https://doi.org/10.1146/annurev-immunol-032713-120156
Levraud JP, Boudinot P, Colin I, Benmansour A, Peyrieras N, Herbomel P, Lutfalla G (2007) Identification of the zebrafish IFN receptor: implications for the origin of the vertebrate IFN system. J Immunol 178(7):4385–4394. https://doi.org/10.4049/jimmunol.178.7.4385
Stark GR, Kerr IM, Williams BR, Silverman RH, Schreiber RD (1998) How cells respond to interferons. Annu Rev Biochem 67:227–264. https://doi.org/10.1146/annurev.biochem.67.1.227
Isaacs A, Lindenmann J (1957) Virus interference. I. The interferon. Proc R Soc Lond B Biol Sci 147 (927):258–267. doi:https://doi.org/10.1098/rspb.1957.0048
Krause CD, Pestka S (2015) Cut, copy, move, delete: The study of human interferon genes reveal multiple mechanisms underlying their evolution in amniotes. Cytokine 76(2):480–495. https://doi.org/10.1016/j.cyto.2015.07.019
Pestka S (2000) The human interferon alpha species and receptors. Biopolymers 55(4):254–287. https://doi.org/10.1002/1097-0282(2000)55:4%3c254::AID-BIP1001%3e3.0.CO;2-1
Cull VS, Tilbrook PA, Bartlett EJ, Brekalo NL, James CM (2003) Type I interferon differential therapy for erythroleukemia: specificity of STAT activation. Blood 101(7):2727–2735. https://doi.org/10.1182/blood-2002-05-1521
Moraga I, Harari D, Schreiber G, Uze G, Pellegrini S (2009) Receptor density is key to the alpha2/beta interferon differential activities. Mol Cell Biol 29(17):4778–4787. https://doi.org/10.1128/MCB.01808-08
Lee AJ, Ashkar AA (2018) The dual nature of Type I and Type II interferons. Front Immunol 9:2061. https://doi.org/10.3389/fimmu.2018.02061
Vivier E, Tomasello E, Baratin M, Walzer T, Ugolini S (2008) Functions of natural killer cells. Nat Immunol 9(5):503–510. https://doi.org/10.1038/ni1582
Kotenko SV, Gallagher G, Baurin VV, Lewis-Antes A, Shen M, Shah NK, Langer JA, Sheikh F, Dickensheets H, Donnelly RP (2003) IFN-lambdas mediate antiviral protection through a distinct class II cytokine receptor complex. Nat Immunol 4(1):69–77. https://doi.org/10.1038/ni875
Sheppard P, Kindsvogel W, Xu W, Henderson K, Schlutsmeyer S, Whitmore TE, Kuestner R, Garrigues U, Birks C, Roraback J, Ostrander C, Dong D, Shin J, Presnell S, Fox B, Haldeman B, Cooper E, Taft D, Gilbert T, Grant FJ, Tackett M, Krivan W, McKnight G, Clegg C, Foster D, Klucher KM (2003) IL-28, IL-29 and their class II cytokine receptor IL-28R. Nat Immunol 4(1):63–68. https://doi.org/10.1038/ni873
Akira S, Uematsu S, Takeuchi O (2006) Pathogen recognition and innate immunity. Cell 124(4):783–801. https://doi.org/10.1016/j.cell.2006.02.015
Medzhitov R (2007) TLR-mediated innate immune recognition. Semin Immunol 19(1):1–2. https://doi.org/10.1016/j.smim.2007.02.001
Finberg RW, Wang JP, Kurt-Jones EA (2007) Toll like receptors and viruses. Rev Med Virol 17(1):35–43. https://doi.org/10.1002/rmv.525
Kato H, Takeuchi O, Sato S, Yoneyama M, Yamamoto M, Matsui K, Uematsu S, Jung A, Kawai T, Ishii KJ, Yamaguchi O, Otsu K, Tsujimura T, Koh CS, Reisesousa C, Matsuura Y, Fujita T, Akira S (2006) Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441(7089):101–105. https://doi.org/10.1038/nature04734
Ichinohe T, Lee HK, Ogura Y, Flavell R, Iwasaki A (2009) Inflammasome recognition of influenza virus is essential for adaptive immune responses. J Exp Med 206(1):79–87. https://doi.org/10.1084/jem.20081667
Ma Z, Ni G, Damania B (2018) Innate sensing of DNA virus genomes. Annu Rev Virol 5(1):341–362. https://doi.org/10.1146/annurev-virology-092917-043244
Heil F, Hemmi H, Hochrein H, Ampenberger F, Kirschning C, Akira S, Lipford G, Wagner H, Bauer S (2004) Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. Science 303(5663):1526–1529. https://doi.org/10.1126/science.1093620
Boehme KW, Compton T (2004) Innate sensing of viruses by toll-like receptors. J Virol 78(15):7867–7873. https://doi.org/10.1128/JVI.78.15.7867-7873.2004
Kawai T, Takahashi K, Sato S, Coban C, Kumar H, Kato H, Ishii KJ, Takeuchi O, Akira S (2005) IPS-1, an adaptor triggering RIG-I- and Mda5-mediated type I interferon induction. Nat Immunol 6(10):981–988. https://doi.org/10.1038/ni1243
Levy DE, Garcia-Sastre A (2001) The virus battles: IFN induction of the antiviral state and mechanisms of viral evasion. Cytokine Growth Factor Rev 12(2–3):143–156. https://doi.org/10.1016/s1359-6101(00)00027-7
Honda K, Taniguchi T (2006) IRFs: master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat Rev Immunol 6(9):644–658. https://doi.org/10.1038/nri1900
Osterlund PI, Pietila TE, Veckman V, Kotenko SV, Julkunen I (2007) IFN regulatory factor family members differentially regulate the expression of type III IFN (IFN-lambda) genes. J Immunol 179(6):3434–3442. https://doi.org/10.4049/jimmunol.179.6.3434
Au WC, Moore PA, LaFleur DW, Tombal B, Pitha PM (1998) Characterization of the interferon regulatory factor-7 and its potential role in the transcription activation of interferon A genes. J Biol Chem 273(44):29210–29217. https://doi.org/10.1074/jbc.273.44.29210
Marie I, Durbin JE, Levy DE (1998) Differential viral induction of distinct interferon-alpha genes by positive feedback through interferon regulatory factor-7. EMBO J 17(22):6660–6669. https://doi.org/10.1093/emboj/17.22.6660
Brazda V, Coufal J, Liao JC, Arrowsmith CH (2012) Preferential binding of IFI16 protein to cruciform structure and superhelical DNA. Biochem Biophys Res Commun 422(4):716–720. https://doi.org/10.1016/j.bbrc.2012.05.065
Murat P, Zhong J, Lekieffre L, Cowieson NP, Clancy JL, Preiss T, Balasubramanian S, Khanna R, Tellam J (2014) G-quadruplexes regulate Epstein–Barr virus-encoded nuclear antigen 1 mRNA translation. Nat Chem Biol 10(5):358–364. https://doi.org/10.1038/nchembio.1479
Civril F, Deimling T, de Oliveira Mann CC, Ablasser A, Moldt M, Witte G, Hornung V, Hopfner KP (2013) Structural mechanism of cytosolic DNA sensing by cGAS. Nature 498(7454):332–337. https://doi.org/10.1038/nature12305
Li XD, Wu J, Gao D, Wang H, Sun L, Chen ZJ (2013) Pivotal roles of cGAS-cGAMP signaling in antiviral defense and immune adjuvant effects. Science 341(6152):1390–1394. https://doi.org/10.1126/science.1244040
Burdette DL, Monroe KM, Sotelo-Troha K, Iwig JS, Eckert B, Hyodo M, Hayakawa Y, Vance RE (2011) STING is a direct innate immune sensor of cyclic di-GMP. Nature 478(7370):515–518. https://doi.org/10.1038/nature10429
Liu S, Cai X, Wu J, Cong Q, Chen X, Li T, Du F, Ren J, Wu YT, Grishin NV, Chen ZJ (2015) Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347 (6227):aaa2630. doi:https://doi.org/10.1126/science.aaa2630
Unterholzner L, Keating SE, Baran M, Horan KA, Jensen SB, Sharma S, Sirois CM, Jin T, Latz E, Xiao TS, Fitzgerald KA, Paludan SR, Bowie AG (2010) IFI16 is an innate immune sensor for intracellular DNA. Nat Immunol 11(11):997–1004. https://doi.org/10.1038/ni.1932
Pattyn E, Van Ostade X, Schauvliege L, Verhee A, Kalai M, Vandekerckhove J, Tavernier J (1999) Dimerization of the interferon type I receptor IFNaR2-2 is sufficient for induction of interferon effector genes but not for full antiviral activity. J Biol Chems 274(49):34838–34845. https://doi.org/10.1074/jbc.274.49.34838
Schindler C, Levy DE, Decker T (2007) JAK-STAT signaling: from interferons to cytokines. J Biol Chem 282(28):20059–20063. https://doi.org/10.1074/jbc.R700016200
Kessler DS, Veals SA, Fu XY, Levy DE (1990) Interferon-alpha regulates nuclear translocation and DNA-binding affinity of ISGF3, a multimeric transcriptional activator. Genes Dev 4(10):1753–1765. https://doi.org/10.1101/gad.4.10.1753
Schoggins JW, Rice CM (2011) Interferon-stimulated genes and their antiviral effector functions. Curr Opin Virol 1(6):519–525. https://doi.org/10.1016/j.coviro.2011.10.008
Schneider WM, Chevillotte MD, Rice CM (2014) Interferon-stimulated genes: a complex web of host defenses. Annu Rev Immunol 32:513–545. https://doi.org/10.1146/annurev-immunol-032713-120231
Verhelst J, Hulpiau P, Saelens X (2013) Mx proteins: antiviral gatekeepers that restrain the uninvited. Microbiol Mol Biol Rev 77(4):551–566. https://doi.org/10.1128/MMBR.00024-13
Dauber B, Wolff T (2009) Activation of the antiviral kinase PKR and viral countermeasures. Viruses 1(3):523–544. https://doi.org/10.3390/v1030523
Zhou A, Paranjape JM, Der SD, Williams BR, Silverman RH (1999) Interferon action in triply deficient mice reveals the existence of alternative antiviral pathways. Virology 258(2):435–440. https://doi.org/10.1006/viro.1999.9738
Haller O, Kochs G (2011) Human MxA protein: an interferon-induced dynamin-like GTPase with broad antiviral activity. J interferon Cytokine Res 31(1):79–87. https://doi.org/10.1089/jir.2010.0076
Staeheli P, Haller O (2018) Human MX2/MxB: a Potent Interferon-Induced Postentry Inhibitor of Herpesviruses and HIV-1. J Virol. https://doi.org/10.1128/JVI.00709-18
Gusho E, Baskar D, Banerjee S (2020) New advances in our understanding of the “unique” RNase L in host pathogen interaction and immune signaling. Cytokine 133:153847. https://doi.org/10.1016/j.cyto.2016.08.009
Kerr IM, Brown RE (1978) pppA2’p5’A2’p5’A: an inhibitor of protein synthesis synthesized with an enzyme fraction from interferon-treated cells. Proc Natl Acad Sci USA 75(1):256–260. https://doi.org/10.1073/pnas.75.1.256
Dong B, Xu L, Zhou A, Hassel BA, Lee X, Torrence PF, Silverman RH (1994) Intrinsic molecular activities of the interferon-induced 2-5A-dependent RNase. J Biol Chem 269(19):14153–14158
Chakrabarti A, Jha BK, Silverman RH (2011) New insights into the role of RNase L in innate immunity. J Interferon Cytokine Res 31(1):49–57. https://doi.org/10.1089/jir.2010.0120
Williams BR (1999) PKR:a sentinel kinase for cellular stress. Oncogene 18(45):6112–6120. https://doi.org/10.1038/sj.onc.1203127
Gil J, Esteban M (2000) The interferon-induced protein kinase (PKR), triggers apoptosis through FADD-mediated activation of caspase 8 in a manner independent of Fas and TNF-alpha receptors. Oncogene 19(32):3665–3674. https://doi.org/10.1038/sj.onc.1203710
Chu WM, Ostertag D, Li ZW, Chang L, Chen Y, Hu Y, Williams B, Perrault J, Karin M (1999) JNK2 and IKKbeta are required for activating the innate response to viral infection. Immunity 11(6):721–731. https://doi.org/10.1016/s1074-7613(00)80146-6
Kirchhoff S, Koromilas AE, Schaper F, Grashoff M, Sonenberg N, Hauser H (1995) IRF-1 induced cell growth inhibition and interferon induction requires the activity of the protein kinase PKR. Oncogene 11(3):439–445
Kumar A, Yang YL, Flati V, Der S, Kadereit S, Deb A, Haque J, Reis L, Weissmann C, Williams BR (1997) Deficient cytokine signaling in mouse embryo fibroblasts with a targeted deletion in the PKR gene: role of IRF-1 and NF-kappaB. EMBO J 16(2):406–416. https://doi.org/10.1093/emboj/16.2.406
Wong AH, Tam NW, Yang YL, Cuddihy AR, Li S, Kirchhoff S, Hauser H, Decker T, Koromilas AE (1997) Physical association between STAT1 and the interferon-inducible protein kinase PKR and implications for interferon and double-stranded RNA signaling pathways. EMBO J 16(6):1291–1304. https://doi.org/10.1093/emboj/16.6.1291
Vladimer GI, Gorna MW, Superti-Furga G (2014) IFITs: emerging roles as key anti-viral proteins. Front Immunol 5:94. https://doi.org/10.3389/fimmu.2014.00094
Habjan M, Hubel P, Lacerda L, Benda C, Holze C, Eberl CH, Mann A, Kindler E, Gil-Cruz C, Ziebuhr J, Thiel V, Pichlmair A (2013) Sequestration by IFIT1 impairs translation of 2׳O-unmethylated capped RNA. PLoS Pathog 9(10):e1003663. https://doi.org/10.1371/journal.ppat.1003663
Abbas YM, Pichlmair A, Górna MW, Superti-Furga G, Nagar B (2013) Structural basis for viral 5’-PPP-RNA recognition by human IFIT proteins. Nature 494(7435):60–64. https://doi.org/10.1038/nature11783
Albert M, Becares M, Falqui M, Fernandez-Lozano C, Guerra S (2018) ISG15, a small molecule with huge implications: regulation of mitochondrial homeostasis. Viruses. https://doi.org/10.3390/v10110629
Zhang D, Zhang DE (2011) Interferon-stimulated gene 15 and the protein ISGylation system. J Interferon Cytokine Res 31(1):119–130. https://doi.org/10.1089/jir.2010.0110
Malakhov MP, Kim KI, Malakhova OA, Jacobs BS, Borden EC, Zhang DE (2003) High-throughput immunoblotting Ubiquitiin-like protein ISG15 modifies key regulators of signal transduction. J Biol Chem 278(19):16608–16613. https://doi.org/10.1074/jbc.M208435200
Shi HX, Yang K, Liu X, Liu XY, Wei B, Shan YF, Zhu LH, Wang C (2010) Positive regulation of interferon regulatory factor 3 activation by Herc5 via ISG15 modification. Mol Cell Biol 30(10):2424–2436. https://doi.org/10.1128/mcb.01466-09
Ganesan M, Poluektova LY, Tuma DJ, Kharbanda KK, Osna NA (2016) Acetaldehyde disrupts interferon alpha signaling in hepatitis C virus-infected liver cells by up-regulating USP18. Alcohol Clin Exp Res 40(11):2329–2338. https://doi.org/10.1111/acer.13226
Fan JB, Arimoto K, Motamedchaboki K, Yan M, Wolf DA, Zhang DE (2015) Identification and characterization of a novel ISG15-ubiquitin mixed chain and its role in regulating protein homeostasis. Sci Rep 5:12704. https://doi.org/10.1038/srep12704
Kim MJ, Hwang SY, Imaizumi T, Yoo JY (2008) Negative feedback regulation of RIG-I-mediated antiviral signaling by interferon-induced ISG15 conjugation. J Virol 82(3):1474–1483. https://doi.org/10.1128/jvi.01650-07
Okumura F, Zou W, Zhang DE (2007) ISG15 modification of the eIF4E cognate 4EHP enhances cap structure-binding activity of 4EHP. Genes Dev 21(3):255–260. https://doi.org/10.1101/gad.1521607
D’Cunha J, Knight E Jr, Haas AL, Truitt RL, Borden EC (1996) Immunoregulatory properties of ISG15, an interferon-induced cytokine. Proc Natl Acad Sci USA 93(1):211–215. https://doi.org/10.1073/pnas.93.1.211
Perng YC, Lenschow DJ (2018) ISG15 in antiviral immunity and beyond. Nat Rev Microbiol 16(7):423–439. https://doi.org/10.1038/s41579-018-0020-5
Uchida L, Espada-Murao LA, Takamatsu Y, Okamoto K, Hayasaka D, Yu F, Nabeshima T, Buerano CC, Morita K (2014) The dengue virus conceals double-stranded RNA in the intracellular membrane to escape from an interferon response. Sci Rep 4:7395. https://doi.org/10.1038/srep07395
Westaway EG, Mackenzie JM, Kenney MT, Jones MK, Khromykh AA (1997) Ultrastructure of Kunjin virus-infected cells: colocalization of NS1 and NS3 with double-stranded RNA, and of NS2B with NS3, in virus-induced membrane structures. J Virol 71(9):6650–6661
Overby AK, Popov VL, Niedrig M, Weber F (2010) Tick-borne encephalitis virus delays interferon induction and hides its double-stranded RNA in intracellular membrane vesicles. J Virol 84(17):8470–8483. https://doi.org/10.1128/JVI.00176-10
Dou D, Revol R, Ostbye H, Wang H, Daniels R (2018) Influenza A virus cell entry, replication, virion assembly and movement. Front Immunol 9:1581. https://doi.org/10.3389/fimmu.2018.01581
Tolonen N, Doglio L, Schleich S, Krijnse Locker J (2001) Vaccinia virus DNA replication occurs in endoplasmic reticulum-enclosed cytoplasmic mini-nuclei. Mol Biol Cell 12(7):2031–2046. https://doi.org/10.1091/mbc.12.7.2031
Desselberger U (2014) Rotaviruses. Virus Res 190:75–96. https://doi.org/10.1016/j.virusres.2014.06.016
Yue Z, Shatkin AJ (1996) Regulated, stable expression and nuclear presence of reovirus double-stranded RNA-binding protein sigma3 in HeLa cells. J Virol 70(6):3497–3501
Di Palma F, Daino GL, Ramaswamy VK, Corona A, Frau A, Fanunza E, Vargiu AV, Tramontano E, Ruggerone P (2019) Relevance of Ebola virus VP35 homo-dimerization on the type I interferon cascade inhibition. Antivir Chem Chemother 27:2040206619889220. https://doi.org/10.1177/2040206619889220
Weber M, Sediri H, Felgenhauer U, Binzen I, Bänfer S, Jacob R, Brunotte L, García-Sastre A, Schmid-Burgk JL, Schmidt T, Hornung V, Kochs G, Schwemmle M, Klenk HD, Weber F (2015) Influenza virus adaptation PB2-627K modulates nucleocapsid inhibition by the pathogen sensor RIG-I. Cell Host Microbe 17(3):309–319. https://doi.org/10.1016/j.chom.2015.01.005
Donelan NR, Basler CF, Garcia-Sastre A (2003) A recombinant influenza A virus expressing an RNA-binding-defective NS1 protein induces high levels of beta interferon and is attenuated in mice. J Virol 77(24):13257–13266. https://doi.org/10.1128/jvi.77.24.13257-13266.2003
Goodfellow I (2011) The genome-linked protein VPg of vertebrate viruses—a multifaceted protein. Curr Opin Virol 1(5):355–362. https://doi.org/10.1016/j.coviro.2011.09.003
Qi X, Lan S, Wang W, Schelde LM, Dong H, Wallat GD, Ly H, Liang Y, Dong C (2010) Cap binding and immune evasion revealed by Lassa nucleoprotein structure. Nature 468(7325):779–783. https://doi.org/10.1038/nature09605
Hastie KM, Kimberlin CR, Zandonatti MA, MacRae IJ, Saphire EO (2011) Structure of the Lassa virus nucleoprotein reveals a dsRNA-specific3׳ to5׳ eٴٴxonuclease activity essential for immune suppression. Proc Natl Acad Sci USA 108(6):2396–2401. https://doi.org/10.1073/pnas.1016404108
Habjan M, Andersson I, Klingstrom J, Schumann M, Martin A, Zimmermann P, Wagner V, Pichlmair A, Schneider U, Muhlberger E, Mirazimi A, Weber F (2008) Processing of genome 5׳ termini as a strategy of negative-strand RNA viruses to avoid RIG-I-dependent interferon induction. PLoS ONE 3(4):e2032. https://doi.org/10.1371/journal.pone.0002032
Liu SW, Katsafanas GC, Liu R, Wyatt LS, Moss B (2015) Poxvirus decapping enzymes enhance virulence by preventing the accumulation of dsRNA and the induction of innate antiviral responses. Cell Host Microbe 17(3):320–331. https://doi.org/10.1016/j.chom.2015.02.002
Nakatsu Y, Takeda M, Ohno S, Shirogane Y, Iwasaki M, Yanagi Y (2008) Measles virus circumvents the host interferon response by different actions of the C and V proteins. J Virol 82(17):8296–8306. https://doi.org/10.1128/jvi.00108-08
Pfaller CK, Bloyet LM, Donohue RC, Huff AL, Bartemes WP, Yousaf I, Urzua E, Clavière M, Zachary M, de Masson DV, Carson S, Schieferecke AJ, Meyer AJ, Gerlier D, Cattaneo R (2020) The C protein is recruited to measles virus ribonucleocapsids by the phosphoprotein. J Virol. https://doi.org/10.1128/jvi.01733-19
Wu JJ, Li W, Shao Y, Avey D, Fu B, Gillen J, Hand T, Ma S, Liu X, Miley W, Konrad A, Neipel F, Sturzl M, Whitby D, Li H, Zhu F (2015) Inhibition of cGAS DNA sensing by a herpesvirus virion protein. Cell Host Microbe 18(3):333–344. https://doi.org/10.1016/j.chom.2015.07.015
Huang J, You H, Su C, Li Y, Chen S, Zheng C (2018) Herpes simplex virus 1 tegument protein VP22 abrogates cGAS/STING-mediated antiviral innate immunity. J Virol. https://doi.org/10.1128/JVI.00841-18
Fan L, Briese T, Lipkin WI (2010) Z proteins of New World arenaviruses bind RIG-I and interfere with type I interferon induction. J Virol 84(4):1785–1791. https://doi.org/10.1128/jvi.01362-09
Siu KL, Kok KH, Ng MH, Poon VK, Yuen KY, Zheng BJ, Jin DY (2009) Severe acute respiratory syndrome coronavirus M protein inhibits type I interferon production by impeding the formation of TRAF3.TANK.TBK1/IKKepsilon complex. J Biol Chem 284(24):16202–16209. https://doi.org/10.1074/jbc.M109.008227
Zhu Z, Wang G, Yang F, Cao W, Mao R, Du X, Zhang X, Li C, Li D, Zhang K, Shu H, Liu X, Zheng H (2016) Foot-and-mouth disease virus viroporin 2B antagonizes RIG-I-mediated antiviral effects by inhibition of its protein expression. J Virol 90(24):11106–11121. https://doi.org/10.1128/JVI.01310-16
Barral PM, Sarkar D, Fisher PB, Racaniello VR (2009) RIG-I is cleaved during picornavirus infection. Virology 391(2):171–176. https://doi.org/10.1016/j.virol.2009.06.045
Feng Q, Langereis MA, Lork M, Nguyen M, Hato SV, Lanke K, Emdad L, Bhoopathi P, Fisher PB, Lloyd RE, van Kuppeveld FJ (2014) Enterovirus 2Apro targets MDA5 and MAVS in infected cells. J Virol 88(6):3369–3378. https://doi.org/10.1128/jvi.02712-13
Orzalli MH, DeLuca NA, Knipe DM (2012) Nuclear IFI16 induction of IRF-3 signaling during herpesviral infection and degradation of IFI16 by the viral ICP0 protein. Proc Natl Acad Sci USA 109(44):E3008-3017. https://doi.org/10.1073/pnas.1211302109
Gori Savellini G, Anichini G, Gandolfo C, Prathyumnan S, Cusi MG (2019) Toscana virus non-structural protein NSs acts as E3 ubiquitin ligase promoting RIG-I degradation. PLoS Pathog 15(12):e1008186. https://doi.org/10.1371/journal.ppat.1008186
Aguirre S, Luthra P, Sanchez-Aparicio MT, Maestre AM, Patel J, Lamothe F, Fredericks AC, Tripathi S, Zhu T, Pintado-Silva J, Webb LG, Bernal-Rubio D, Solovyov A, Greenbaum B, Simon V, Basler CF, Mulder LC, Garcia-Sastre A, Fernandez-Sesma A (2017) Dengue virus NS2B protein targets cGAS for degradation and prevents mitochondrial DNA sensing during infection. Nat Microbiol 2:17037. https://doi.org/10.1038/nmicrobiol.2017.37
Clementz MA, Chen Z, Banach BS, Wang Y, Sun L, Ratia K, Baez-Santos YM, Wang J, Takayama J, Ghosh AK, Li K, Mesecar AD, Baker SC (2010) Deubiquitinating and interferon antagonism activities of coronavirus papain-like proteases. J Virol 84(9):4619–4629. https://doi.org/10.1128/JVI.02406-09
Deng X, Chen Y, Mielech AM, Hackbart M, Kesely KR, Mettelman RC, O’Brien A, Chapman ME, Mesecar AD, Baker SC (2020) Structure-guided mutagenesis alters deubiquitinating activity and attenuates pathogenesis of a murine coronavirus. J Virol. https://doi.org/10.1128/JVI.01734-19
Ratia K, Kilianski A, Baez-Santos YM, Baker SC, Mesecar A (2014) Structural basis for the ubiquitin-linkage specificity and deISGylating activity of SARS-CoV papain-like protease. PLoS Pathog 10(5):e1004113. https://doi.org/10.1371/journal.ppat.1004113
Mielech AM, Kilianski A, Baez-Santos YM, Mesecar AD, Baker SC (2014) MERS-CoV papain-like protease has deISGylating and deubiquitinating activities. Virology 450–451:64–70. https://doi.org/10.1016/j.virol.2013.11.040
Parisien JP, Bamming D, Komuro A, Ramachandran A, Rodriguez JJ, Barber G, Wojahn RD, Horvath CM (2009) A shared interface mediates paramyxovirus interference with antiviral RNA helicases MDA5 and LGP2. J Virol 83(14):7252–7260. https://doi.org/10.1128/jvi.00153-09
Davis ME, Wang MK, Rennick LJ, Full F, Gableske S, Mesman AW, Gringhuis SI, Geijtenbeek TB, Duprex WP, Gack MU (2014) Antagonism of the phosphatase PP1 by the measles virus V protein is required for innate immune escape of MDA5. Cell Host Microbe 16(1):19–30. https://doi.org/10.1016/j.chom.2014.06.007
Mesman AW, Zijlstra-Willems EM, Kaptein TM, de Swart RL, Davis ME, Ludlow M, Duprex WP, Gack MU, Gringhuis SI, Geijtenbeek TB (2014) Measles virus suppresses RIG-I-like receptor activation in dendritic cells via DC-SIGN-mediated inhibition of PP1 phosphatases. Cell Host Microbe 16(1):31–42. https://doi.org/10.1016/j.chom.2014.06.008
Kok KH, Lui PY, Ng MH, Siu KL, Au SW, Jin DY (2011) The double-stranded RNA-binding protein PACT functions as a cellular activator of RIG-I to facilitate innate antiviral response. Cell Host Microbe 9(4):299–309. https://doi.org/10.1016/j.chom.2011.03.007
Lui PY, Wong LR, Ho TH, Au SWN, Chan CP, Kok KH, Jin DY (2017) PACT facilitates RNA-induced activation of MDA5 by promoting MDA5 oligomerization. J Immunol 199(5):1846–1855. https://doi.org/10.4049/jimmunol.1601493
Siu KL, Yeung ML, Kok KH, Yuen KS, Kew C, Lui PY, Chan CP, Tse H, Woo PC, Yuen KY, Jin DY (2014) Middle east respiratory syndrome coronavirus 4a protein is a double-stranded RNA-binding protein that suppresses PACT-induced activation of RIG-I and MDA5 in the innate antiviral response. J Virol 88(9):4866–4876. https://doi.org/10.1128/jvi.03649-13
Luthra P, Ramanan P, Mire CE, Weisend C, Tsuda Y, Yen B, Liu G, Leung DW, Geisbert TW, Ebihara H, Amarasinghe GK, Basler CF (2013) Mutual antagonism between the Ebola virus VP35 protein and the RIG-I activator PACT determines infection outcome. Cell Host Microbe 14(1):74–84. https://doi.org/10.1016/j.chom.2013.06.010
Shao J, Huang Q, Liu X, Di D, Liang Y, Ly H (2018) Arenaviral nucleoproteins suppress PACT-induced augmentation of RIG-I function to inhibit type I interferon production. J Virol. https://doi.org/10.1128/jvi.00482-18
Oshiumi H, Miyashita M, Matsumoto M, Seya T (2013) A distinct role of Riplet-mediated K63-Linked polyubiquitination of the RIG-I repressor domain in human antiviral innate immune responses. PLoS Pathog 9(8):e1003533. https://doi.org/10.1371/journal.ppat.1003533
Gack MU, Shin YC, Joo CH, Urano T, Liang C, Sun L, Takeuchi O, Akira S, Chen Z, Inoue S, Jung JU (2007) TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 446(7138):916–920. https://doi.org/10.1038/nature05732
Gack MU, Albrecht RA, Urano T, Inn KS, Huang IC, Carnero E, Farzan M, Inoue S, Jung JU, Garcia-Sastre A (2009) Influenza A virus NS1 targets the ubiquitin ligase TRIM25 to evade recognition by the host viral RNA sensor RIG-I. Cell Host Microbe 5(5):439–449. https://doi.org/10.1016/j.chom.2009.04.006
Koliopoulos MG, Lethier M, van der Veen AG, Haubrich K, Hennig J, Kowalinski E, Stevens RV, Martin SR, Reis e Sousa C, Cusack S, Rittinger K, (2018) Molecular mechanism of influenza A NS1-mediated TRIM25 recognition and inhibition. Nat Commun 9(1):1820. https://doi.org/10.1038/s41467-018-04214-8
Manokaran G, Finol E, Wang C, Gunaratne J, Bahl J, Ong EZ, Tan HC, Sessions OM, Ward AM, Gubler DJ, Harris E, Garcia-Blanco MA, Ooi EE (2015) Dengue subgenomic RNA binds TRIM25 to inhibit interferon expression for epidemiological fitness. Science 350(6257):217–221. https://doi.org/10.1126/science.aab3369
Chan YK, Gack MU (2016) A phosphomimetic-based mechanism of dengue virus to antagonize innate immunity. Nat Immunol 17(5):523–530. https://doi.org/10.1038/ni.3393
Chen X, Yang X, Zheng Y, Yang Y, Xing Y, Chen Z (2014) SARS coronavirus papain-like protease inhibits the type I interferon signaling pathway through interaction with the STING–TRAF3–TBK1 complex. Protein Cell 5(5):369–381. https://doi.org/10.1007/s13238-014-0026-3
Sun L, Xing Y, Chen X, Zheng Y, Yang Y, Nichols DB, Clementz MA, Banach BS, Li K, Baker SC, Chen Z (2012) Coronavirus papain-like proteases negatively regulate antiviral innate immune response through disruption of STING-mediated signaling. PLoS ONE 7(2):e30802. https://doi.org/10.1371/journal.pone.0030802
Ni G, Konno H, Barber GN (2017) Ubiquitination of STING at lysine 224 controls IRF3 activation. Sci Immunol. https://doi.org/10.1126/sciimmunol.aah7119
Li SW, Wang CY, Jou YJ, Huang SH, Hsiao LH, Wan L, Lin YJ, Kung SH, Lin CW (2016) SARS Coronavirus papain-like protease inhibits the TLR7 signaling pathway through removing Lys63-linked polyubiquitination of TRAF3 and TRAF6. Int J Mol Sci. https://doi.org/10.3390/ijms17050678
Pourcelot M, Zemirli N, Silva Da Costa L, Loyant R, Garcin D, Vitour D, Munitic I, Vazquez A, Arnoult D (2016) The Golgi apparatus acts as a platform for TBK1 activation after viral RNA sensing. BMC Biol 14:69. https://doi.org/10.1186/s12915-016-0292-z
Eaglesham JB, Pan Y, Kupper TS, Kranzusch PJ (2019) Viral and metazoan poxins are cGAMP-specific nucleases that restrict cGAS-STING signalling. Nature 566(7743):259–263. https://doi.org/10.1038/s41586-019-0928-6
Lau L, Gray EE, Brunette RL, Stetson DB (2015) DNA tumor virus oncogenes antagonize the cGAS-STING DNA-sensing pathway. Science 350(6260):568–571. https://doi.org/10.1126/science.aab3291
Lui PY, Wong LY, Fung CL, Siu KL, Yeung ML, Yuen KS, Chan CP, Woo PC, Yuen KY, Jin DY (2016) Middle East respiratory syndrome coronavirus M protein suppresses type I interferon expression through the inhibition of TBK1-dependent phosphorylation of IRF3. Emerg Microbes Infect 5:e39. https://doi.org/10.1038/emi.2016.33
Siu KL, Chan CP, Kok KH, Chiu-Yat Woo P, Jin DY (2014) Suppression of innate antiviral response by severe acute respiratory syndrome coronavirus M protein is mediated through the first transmembrane domain. Cell Mol Immunol 11(2):141–149. https://doi.org/10.1038/cmi.2013.61
Yang Y, Ye F, Zhu N, Wang W, Deng Y, Zhao Z, Tan W (2015) Middle East respiratory syndrome coronavirus ORF4b protein inhibits type I interferon production through both cytoplasmic and nuclear targets. Sci Rep 5:17554. https://doi.org/10.1038/srep17554
Ma Z, Jacobs SR, West JA, Stopford C, Zhang Z, Davis Z, Barber GN, Glaunsinger BA, Dittmer DP, Damania B (2015) Modulation of the cGAS-STING DNA sensing pathway by gammaherpesviruses. Proc Natl Acad Sci USA 112(31):E4306-4315. https://doi.org/10.1073/pnas.1503831112
Christensen MH, Jensen SB, Miettinen JJ, Luecke S, Prabakaran T, Reinert LS, Mettenleiter T, Chen ZJ, Knipe DM, Sandri-Goldin RM, Enquist LW, Hartmann R, Mogensen TH, Rice SA, Nyman TA, Matikainen S, Paludan SR (2016) HSV-1 ICP27 targets the TBK1-activated STING signalsome to inhibit virus-induced type I IFN expression. EMBO J 35(13):1385–1399. https://doi.org/10.15252/embj.201593458
Ning YJ, Feng K, Min YQ, Deng F, Hu Z, Wang H (2017) Heartland virus NSs protein disrupts host defenses by blocking the TBK1 kinase-IRF3 transcription factor interaction and signaling required for interferon induction. J Biol Chem 292(40):16722–16733. https://doi.org/10.1074/jbc.M117.805127
Ding Q, Gaska JM, Douam F, Wei L, Kim D, Balev M, Heller B, Ploss A (2018) Species-specific disruption of STING-dependent antiviral cellular defenses by the Zika virus NS2B3 protease. Proc Natl Acad Sci USA 115(27):E6310–E6318. https://doi.org/10.1073/pnas.1803406115
Aguirre S, Maestre AM, Pagni S, Patel JR, Savage T, Gutman D, Maringer K, Bernal-Rubio D, Shabman RS, Simon V, Rodriguez-Madoz JR, Mulder LC, Barber GN, Fernandez-Sesma A (2012) DENV inhibits type I IFN production in infected cells by cleaving human STING. PLoS Pathog 8(10):e1002934. https://doi.org/10.1371/journal.ppat.1002934
Yu CY, Chang TH, Liang JJ, Chiang RL, Lee YL, Liao CL, Lin YL (2012) Dengue virus targets the adaptor protein MITA to subvert host innate immunity. PLoS Pathog 8(6):e1002780. https://doi.org/10.1371/journal.ppat.1002780
Qian S, Fan W, Liu T, Wu M, Zhang H, Cui X, Zhou Y, Hu J, Wei S, Chen H, Li X, Qian P (2017) Seneca valley virus suppresses host type I interferon production by targeting adaptor proteins MAVS, TRIF, and TANK for cleavage. J Virol. https://doi.org/10.1128/jvi.00823-17
Mukherjee A, Morosky SA, Delorme-Axford E, Dybdahl-Sissoko N, Oberste MS, Wang T, Coyne CB (2011) The coxsackievirus B 3C protease cleaves MAVS and TRIF to attenuate host type I interferon and apoptotic signaling. PLoS Pathog 7(3):e1001311. https://doi.org/10.1371/journal.ppat.1001311
Dong J, Xu S, Wang J, Luo R, Wang D, Xiao S, Fang L, Chen H, Jiang Y (2015) Porcine reproductive and respiratory syndrome virus 3C protease cleaves the mitochondrial antiviral signalling complex to antagonize IFN-beta expression. J General Virol 96(10):3049–3058. https://doi.org/10.1099/jgv.0.000257
Li XD, Sun L, Seth RB, Pineda G, Chen ZJ (2005) Hepatitis C virus protease NS3/4A cleaves mitochondrial antiviral signaling protein off the mitochondria to evade innate immunity. Proc Natl Acad Sci USA 102(49):17717–17722. https://doi.org/10.1073/pnas.0508531102
Nandi S, Chanda S, Bagchi P, Nayak MK, Bhowmick R, Chawla-Sarkar M (2014) MAVS protein is attenuated by rotavirus nonstructural protein 1. PLoS ONE 9(3):e92126. https://doi.org/10.1371/journal.pone.0092126
Ding S, Zhu S, Ren L, Feng N, Song Y, Ge X, Li B, Flavell RA, Greenberg HB (2018) Rotavirus VP3 targets MAVS for degradation to inhibit type III interferon expression in intestinal epithelial cells. Elife. https://doi.org/10.7554/eLife.39494
Shi CS, Qi HY, Boularan C, Huang NN, Abu-Asab M, Shelhamer JH, Kehrl JH (2014) SARS-coronavirus open reading frame-9b suppresses innate immunity by targeting mitochondria and the MAVS/TRAF3/TRAF6 signalosome. J Immunol 193(6):3080–3089. https://doi.org/10.4049/jimmunol.1303196
You F, Sun H, Zhou X, Sun W, Liang S, Zhai Z, Jiang Z (2009) PCBP2 mediates degradation of the adaptor MAVS via the HECT ubiquitin ligase AIP4. Nat Immunol 10(12):1300–1308. https://doi.org/10.1038/ni.1815
Yu CY, Liang JJ, Li JK, Lee YL, Chang BL, Su CI, Huang WJ, Lai MM, Lin YL (2015) Dengue virus impairs mitochondrial fusion by cleaving mitofusins. PLoS Pathog 11(12):e1005350. https://doi.org/10.1371/journal.ppat.1005350
Xia M, Meng G, Jiang A, Chen A, Dahlhaus M, Gonzalez P, Beltinger C, Wei J (2014) Mitophagy switches cell death from apoptosis to necrosis in NSCLC cells treated with oncolytic measles virus. Oncotarget 5(11):3907–3918. https://doi.org/10.18632/oncotarget.2028
Meng G, Xia M, Wang D, Chen A, Wang Y, Wang H, Yu D, Wei J (2014) Mitophagy promotes replication of oncolytic Newcastle disease virus by blocking intrinsic apoptosis in lung cancer cells. Oncotarget 5(15):6365–6374. https://doi.org/10.18632/oncotarget.2219
Kim SJ, Khan M, Quan J, Till A, Subramani S, Siddiqui A (2013) Hepatitis B virus disrupts mitochondrial dynamics: induces fissi on and mitophagy to attenuate apoptosis. PLoS Pathog 9(12):e1003722. https://doi.org/10.1371/journal.ppat.1003722
Ding B, Zhang L, Li Z, Zhong Y, Tang Q, Qin Y, Chen M (2017) The matrix protein of human parainfluenza virus type 3 induces mitophagy that suppresses interferon responses. Cell Host Microbe 21(4):538–547. https://doi.org/10.1016/j.chom.2017.03.004
Wang R, Zhu Y, Ren C, Yang S, Tian S, Chen H, Jin M, Zhou H (2020) Influenza A virus protein PB1-F2 impairs innate immunity by inducing mitophagy. Autophagy. https://doi.org/10.1080/15548627.2020.1725375
Yamaguchi M, Kitagawa Y, Zhou M, Itoh M, Gotoh B (2014) An anti-interferon activity shared by paramyxovirus C proteins: inhibition of Toll-like receptor 7/9-dependent alpha interferon induction. FEBS Lett 588(1):28–34. https://doi.org/10.1016/j.febslet.2013.11.015
Anglero-Rodriguez YI, Pantoja P, Sariol CA (2014) Dengue virus subverts the interferon induction pathway via NS2B/3 protease-IkappaB kinase epsilon interaction. Clin Vaccine Immunol 21(1):29–38. https://doi.org/10.1128/CVI.00500-13
Pythoud C, Rodrigo WW, Pasqual G, Rothenberger S, Martinez-Sobrido L, de la Torre JC, Kunz S (2012) Arenavirus nucleoprotein targets interferon regulatory factor-activating kinase IKKepsilon. J Virol 86(15):7728–7738. https://doi.org/10.1128/JVI.00187-12
Martinez-Sobrido L, Zuniga EI, Rosario D, Garcia-Sastre A, de la Torre JC (2006) Inhibition of the type I interferon response by the nucleoprotein of the prototypic arenavirus lymphocytic choriomeningitis virus. J Virol 80(18):9192–9199. https://doi.org/10.1128/JVI.00555-06
Hiscott J (2007) Convergence of the NF-kappaB and IRF pathways in the regulation of the innate antiviral response. Cytokine Growth Factor Rev 18(5–6):483–490. https://doi.org/10.1016/j.cytogfr.2007.06.002
Rodrigo WW, Ortiz-Riano E, Pythoud C, Kunz S, de la Torre JC, Martinez-Sobrido L (2012) Arenavirus nucleoproteins prevent activation of nuclear factor kappa B. J Virol 86(15):8185–8197. https://doi.org/10.1128/JVI.07240-11
Wang D, Fang L, Wei D, Zhang H, Luo R, Chen H, Li K, Xiao S (2014) Hepatitis A virus 3C protease cleaves NEMO to impair induction of beta interferon. J Virol 88(17):10252–10258. https://doi.org/10.1128/JVI.00869-14
Wang D, Fang L, Li K, Zhong H, Fan J, Ouyang C, Zhang H, Duan E, Luo R, Zhang Z, Liu X, Chen H, Xiao S (2012) Foot-and-mouth disease virus 3C protease cleaves NEMO to impair innate immune signaling. J Virol 86(17):9311–9322. https://doi.org/10.1128/jvi.00722-12
Wang D, Fang L, Shi Y, Zhang H, Gao L, Peng G, Chen H, Li K, Xiao S (2016) Porcine epidemic diarrhea virus 3C-like protease regulates its interferon antagonism by cleaving NEMO. J Virol 90(4):2090–2101. https://doi.org/10.1128/JVI.02514-15
Chen J, Wang D, Sun Z, Gao L, Zhu X, Guo J, Xu S, Fang L, Li K, Xiao S (2019) Arterivirus nsp4 antagonizes interferon beta production by proteolytically cleaving NEMO at multiple sites. J Virol. https://doi.org/10.1128/JVI.00385-19
Arnold MM, Patton JT (2011) Diversity of interferon antagonist activities mediated by NSP1 proteins of different rotavirus strains. J Virol 85(5):1970–1979. https://doi.org/10.1128/JVI.01801-10
Arnold MM, Barro M, Patton JT (2013) Rotavirus NSP1 mediates degradation of interferon regulatory factors through targeting of the dimerization domain. J Virol 87(17):9813–9821. https://doi.org/10.1128/jvi.01146-13
Graff JW, Ettayebi K, Hardy ME (2009) Rotavirus NSP1 inhibits NFkappaB activation by inducing proteasome-dependent degradation of beta-TrCP: a novel mechanism of IFN antagonism. PLoS Pathog 5(1):e1000280. https://doi.org/10.1371/journal.ppat.1000280
Morelli M, Dennis AF, Patton JT (2015) Putative E3 ubiquitin ligase of human rotavirus inhibits NF-kappaB activation by using molecular mimicry to target beta-TrCP. mBio. https://doi.org/10.1128/mBio.02490-14
Bour S, Perrin C, Akari H, Strebel K (2001) The human immunodeficiency virus type 1 Vpu protein inhibits NF-kappa B activation by interfering with beta TrCP-mediated degradation of Ikappa B. J Biol Chem 276(19):15920–15928. https://doi.org/10.1074/jbc.M010533200
Tang W, Pavlish OA, Spiegelman VS, Parkhitko AA, Fuchs SY (2003) Interaction of Epstein–Barr virus latent membrane protein 1 with SCFHOS/beta-TrCP E3 ubiquitin ligase regulates extent of NF-kappaB activation. J Biol Chem 278(49):48942–48949. https://doi.org/10.1074/jbc.M307962200
Mansur DS, Maluquer de Motes C, Unterholzner L, Sumner RP, Ferguson BJ, Ren H, Strnadova P, Bowie AG, Smith GL (2013) Poxvirus targeting of E3 ligase beta-TrCP by molecular mimicry: a mechanism to inhibit NF-kappaB activation and promote immune evasion and virulence. PLoS Pathog 9(2):e1003183. https://doi.org/10.1371/journal.ppat.1003183
Lin R, Genin P, Mamane Y, Sgarbanti M, Battistini A, Harrington WJ Jr, Barber GN, Hiscott J (2001) HHV-8 encoded vIRF-1 represses the interferon antiviral response by blocking IRF-3 recruitment of the CBP/p300 coactivators. Oncogene 20(7):800–811. https://doi.org/10.1038/sj.onc.1204163
Irie T, Kiyotani K, Igarashi T, Yoshida A, Sakaguchi T (2012) Inhibition of interferon regulatory factor 3 activation by paramyxovirus V protein. J Virol 86(13):7136–7145. https://doi.org/10.1128/JVI.06705-11
Ye J, Chen Z, Li Y, Zhao Z, He W, Zohaib A, Song Y, Deng C, Zhang B, Chen H, Cao S (2017) Japanese encephalitis virus NS5 inhibits type I interferon (IFN) production by blocking the nuclear translocation of IFN regulatory factor 3 and NF-kappaB. J Virol. https://doi.org/10.1128/JVI.00039-17
Wu X, Qi X, Qu B, Zhang Z, Liang M, Li C, Cardona CJ, Li D, Xing Z (2014) Evasion of antiviral immunity through sequestering of TBK1/IKKepsilon/IRF3 into viral inclusion bodies. J Virol 88(6):3067–3076. https://doi.org/10.1128/jvi.03510-13
Ratinier M, Shaw AE, Barry G, Gu Q, Di Gialleonardo L, Janowicz A, Varela M, Randall RE, Caporale M, Palmarini M (2016) Bluetongue virus NS4 protein is an interferon antagonist and a determinant of virus virulence. J Virol 90(11):5427–5439. https://doi.org/10.1128/jvi.00422-16
Sparrer KM, Pfaller CK, Conzelmann KK (2012) Measles virus C protein interferes with Beta interferon transcription in the nucleus. J Virol 86(2):796–805. https://doi.org/10.1128/jvi.05899-11
Wuerth JD, Habjan M, Wulle J, Superti-Furga G, Pichlmair A, Weber F (2018) NSs protein of sandfly fever sicilian phlebovirus counteracts interferon (IFN) induction by masking the DNA-binding domain of IFN regulatory factor 3. J Virol. https://doi.org/10.1128/jvi.01202-18
Hwang S, Kim KS, Flano E, Wu TT, Tong LM, Park AN, Song MJ, Sanchez DJ, O’Connell RM, Cheng G, Sun R (2009) Conserved herpesviral kinase promotes viral persistence by inhibiting the IRF-3-mediated type I interferon response. Cell Host Microbe 5(2):166–178. https://doi.org/10.1016/j.chom.2008.12.013
Kitagawa Y, Yamaguchi M, Kohno M, Sakai M, Itoh M, Gotoh B (2020) Respirovirus C protein inhibits activation of type I interferon receptor-associated kinases to block JAK–STAT signaling. FEBS Lett 594(5):864–877. https://doi.org/10.1002/1873-3468.13670
Lin RJ, Chang BL, Yu HP, Liao CL, Lin YL (2006) Blocking of interferon-induced Jak–Stat signaling by Japanese encephalitis virus NS5 through a protein tyrosine phosphatase-mediated mechanism. J Virol 80(12):5908–5918. https://doi.org/10.1128/JVI.02714-05
Sanchez-Aparicio MT, Feinman LJ, Garcia-Sastre A, Shaw ML (2018) Paramyxovirus V proteins interact with the RIG-I/TRIM25 regulatory complex and inhibit RIG-I signaling. J Virol. https://doi.org/10.1128/JVI.01960-17
Li P, Zhu Z, Zhang X, Dang W, Li L, Du X, Zhang M, Wu C, Xue Q, Liu X, Zheng H, Nan Y (2019) The nucleoprotein and phosphoprotein of peste des petits ruminants virus inhibit interferons signaling by blocking the JAK-STAT pathway. Viruses. https://doi.org/10.3390/v11070629
Johnson KE, Song B, Knipe DM (2008) Role for herpes simplex virus 1 ICP27 in the inhibition of type I interferon signaling. Virology 374(2):487–494. https://doi.org/10.1016/j.virol.2008.01.001
Munoz-Jordan JL, Laurent-Rolle M, Ashour J, Martinez-Sobrido L, Ashok M, Lipkin WI, Garcia-Sastre A (2005) Inhibition of alpha/beta interferon signaling by the NS4B protein of flaviviruses. J Virol 79(13):8004–8013. https://doi.org/10.1128/JVI.79.13.8004-8013.2005
Kumthip K, Chusri P, Jilg N, Zhao L, Fusco DN, Zhao H, Goto K, Cheng D, Schaefer EA, Zhang L, Pantip C, Thongsawat S, O’Brien A, Peng LF, Maneekarn N, Chung RT, Lin W (2012) Hepatitis C virus NS5A disrupts STAT1 phosphorylation and suppresses type I interferon signaling. J Virol 86(16):8581–8591. https://doi.org/10.1128/JVI.00533-12
Sen A, Rott L, Phan N, Mukherjee G, Greenberg HB (2014) Rotavirus NSP1 protein inhibits interferon-mediated STAT1 activation. J Virol 88(1):41–53. https://doi.org/10.1128/JVI.01501-13
Avia M, Rojas JM, Miorin L, Pascual E, Van Rijn PA, Martin V, Garcia-Sastre A, Sevilla N (2019) Virus-induced autophagic degradation of STAT2 as a mechanism for interferon signaling blockade. EMBO Rep 20(11):e48766. https://doi.org/10.15252/embr.201948766
Vitour D, Doceul V, Ruscanu S, Chauveau E, Schwartz-Cornil I, Zientara S (2014) Induction and control of the type I interferon pathway by Bluetongue virus. Virus Res 182:59–70. https://doi.org/10.1016/j.virusres.2013.10.027
Nagano Y, Sugiyama A, Kimoto M, Wakahara T, Noguchi Y, Jiang X, Saijo S, Shimizu N, Yabuno N, Yao M, Gooley PR, Moseley GW, Tadokoro T, Maenaka K, Ose T (2020) The measles virus V protein binding site to STAT2 overlaps with that of IRF9. J Virol. https://doi.org/10.1128/JVI.01169-20
Ramachandran A, Parisien JP, Horvath CM (2008) STAT2 is a primary target for measles virus V protein-mediated alpha/beta interferon signaling inhibition. J Virol 82(17):8330–8338. https://doi.org/10.1128/JVI.00831-08
Laurent-Rolle M, Morrison J, Rajsbaum R, Macleod JML, Pisanelli G, Pham A, Ayllon J, Miorin L, Martinez C, tenOever BR, Garcia-Sastre A (2014) The interferon signaling antagonist function of yellow fever virus NS5 protein is activated by type I interferon. Cell Host Microbe 16(3):314–327. https://doi.org/10.1016/j.chom.2014.07.015
Morrison J, Laurent-Rolle M, Maestre AM, Rajsbaum R, Pisanelli G, Simon V, Mulder LC, Fernandez-Sesma A, Garcia-Sastre A (2013) Dengue virus co-opts UBR4 to degrade STAT2 and antagonize type I interferon signaling. PLoS Pathog 9(3):e1003265. https://doi.org/10.1371/journal.ppat.1003265
Yang L, He J, Wang R, Zhang X, Lin S, Ma Z, Zhang Y (2019) Nonstructural protein 11 of porcine reproductive and respiratory syndrome virus induces STAT2 degradation to inhibit interferon signaling. J Virol. https://doi.org/10.1128/JVI.01352-19
Didcock L, Young DF, Goodbourn S, Randall RE (1999) The V protein of simian virus 5 inhibits interferon signalling by targeting STAT1 for proteasome-mediated degradation. J Virol 73(12):9928–9933
Kubota T, Yokosawa N, Yokota S, Fujii N (2002) Association of mumps virus V protein with RACK1 results in dissociation of STAT-1 from the alpha interferon receptor complex. J Virol 76(24):12676–12682. https://doi.org/10.1128/jvi.76.24.12676-12682.2002
Antonsson A, Payne E, Hengst K, McMillan NA (2006) The human papillomavirus type 16 E7 protein binds human interferon regulatory factor-9 via a novel PEST domain required for transformation. J Interferon Cytokine Res 26(7):455–461. https://doi.org/10.1089/jir.2006.26.455
Francois C, DuverlieRebouillat GD, Khorsi H, Castelain S, Blum HE, Gatignol A, Wychowski C, Moradpour D, Meurs EF (2000) Expression of hepatitis C virus proteins interferes with the antiviral action of interferon independently of PKR-mediated control of protein synthesis. J Virol 74(12):5587–5596. https://doi.org/10.1128/jvi.74.12.5587-5596.2000
Chee AV, Roizman B (2004) Herpes simplex virus 1 gene products occlude the interferon signaling pathway at multiple sites. J Virol 78(8):4185–4196. https://doi.org/10.1128/jvi.78.8.4185-4196.2004
Miller DM, Zhang Y, Rahill BM, Waldman WJ, Sedmak DD (1999) Human cytomegalovirus inhibits IFN-alpha-stimulated antiviral and immunoregulatory responses by blocking multiple levels of IFN-alpha signal transduction. J Immunol 162(10):6107–6113
Weihua X, Ramanujam S, Lindner DJ, Kudaravalli RD, Freund R, Kalvakolanu DV (1998) The polyoma virus T antigen interferes with interferon-inducible gene expression. Proc Natl Acad Sci USA 95(3):1085–1090. https://doi.org/10.1073/pnas.95.3.1085
Visser LJ, Medina GN, Rabouw HH, de Groot RJ, Langereis MA, de Los Santos T, van Kuppeveld FJM (2019) Foot-and-mouth disease virus leader protease cleaves G3BP1 and G3BP2 and inhibits stress granule formation. J Virol. https://doi.org/10.1128/JVI.00922-18
Okonski KM, Samuel CE (2013) Stress granule formation induced by measles virus is protein kinase PKR dependent and impaired by RNA adenosine deaminase ADAR1. J Virol 87(2):756–766. https://doi.org/10.1128/JVI.02270-12
Bidet K, Dadlani D, Garcia-Blanco MA (2014) G3BP1, G3BP2 and CAPRIN1 are required for translation of interferon stimulated mRNAs and are targeted by a dengue virus non-coding RNA. PLoS Pathog 10(7):e1004242. https://doi.org/10.1371/journal.ppat.1004242
Imani F, Jacobs BL (1988) Inhibitory activity for the interferon-induced protein kinase is associated with the reovirus serotype 1 sigma 3 protein. Proc Natl Acad Sci USA 85(21):7887–7891. https://doi.org/10.1073/pnas.85.21.7887
Denzler KL, Jacobs BL (1994) Site-directed mutagenic analysis of reovirus sigma 3 protein binding to dsRNA. Virology 204(1):190–199. https://doi.org/10.1006/viro.1994.1523
Ly HJ, Ikegami T (2016) Rift Valley fever virus NSs protein functions and the similarity to other bunyavirus NSs proteins. Virology 13:118. https://doi.org/10.1186/s12985-016-0573-8
Toth AM, Li Z, Cattaneo R, Samuel CE (2009) RNA-specific adenosine deaminase ADAR1 suppresses measles virus-induced apoptosis and activation of protein kinase PKR. J Biol Chem 284(43):29350–29356. https://doi.org/10.1074/jbc.M109.045146
Nie Y, Hammond GL, Yang JH (2007) Double-stranded RNA deaminase ADAR1 increases host susceptibility to virus infection. J Virol 81(2):917–923. https://doi.org/10.1128/jvi.01527-06
Clerzius G, Gélinas JF, Daher A, Bonnet M, Meurs EF, Gatignol A (2009) ADAR1 interacts with PKR during human immunodeficiency virus infection of lymphocytes and contributes to viral replication. J Virol 83(19):10119–10128. https://doi.org/10.1128/jvi.02457-08
Morelli M, Ogden KM, Patton JT (2015) Silencing the alarms: Innate immune antagonism by rotavirus NSP1 and VP3. Virology 479–480:75–84. https://doi.org/10.1016/j.virol.2015.01.006
Zhang R, Jha BK, Ogden KM, Dong B, Zhao L, Elliott R, Patton JT, Silverman RH, Weiss SR (2013) Homologous 2’,5’-phosphodiesterases from disparate RNA viruses antagonize antiviral innate immunity. Proc Natl Acad Sci USA 110(32):13114–13119. https://doi.org/10.1073/pnas.1306917110
Chen D, Luongo CL, Nibert ML, Patton JT (1999) Rotavirus open cores catalyze 5′-capping and methylation of exogenous RNA: evidence that VP3 is a methyltransferase. Virology 265(1):120–130. https://doi.org/10.1006/viro.1999.0029
Daffis S, Szretter KJ, Schriewer J, Li J, Youn S, Errett J, Lin T-Y, Schneller S, Zust R, Dong H, Thiel V, Sen GC, Fensterl V, Klimstra WB, Pierson TC, Buller RM, Gale M Jr, Shi P-Y, Diamond MS (2010) 2′-O methylation of the viral mRNA cap evades host restriction by IFIT family members. Nature 468(7322):452–456. https://doi.org/10.1038/nature09489
Menachery VD, Yount BL Jr, Josset L, Gralinski LE, Scobey T, Agnihothram S, Katze MG, Baric RS (2014) Attenuation and restoration of severe acute respiratory syndrome coronavirus mutant lacking 2’-o-methyltransferase activity. J Virol 88(8):4251–4264. https://doi.org/10.1128/jvi.03571-13
Chen Y, Su C, Ke M, Jin X, Xu L, Zhang Z, Wu A, Sun Y, Yang Z, Tien P, Ahola T, Liang Y, Liu X, Guo D (2011) Biochemical and structural insights into the mechanisms of SARS coronavirus RNA ribose 2’-O-methylation by nsp16/nsp10 protein complex. PLoS Pathog 7(10):e1002294. https://doi.org/10.1371/journal.ppat.1002294
Menachery VD, Gralinski LE, Mitchell HD, Dinnon KH 3rd, Leist SR, Yount BL Jr, Graham RL, McAnarney ET, Stratton KG, Cockrell AS, Debbink K, Sims AC, Waters KM, Baric RS (2017) Middle East Respiratory Syndrome Coronavirus Nonstructural Protein 16 Is Necessary for Interferon Resistance and Viral Pathogenesis. mSphere. https://doi.org/10.1128/mSphere.00346-17
Brass AL, Huang IC, Benita Y, John SP, Krishnan MN, Feeley EM, Ryan BJ, Weyer JL, van der Weyden L, Fikrig E, Adams DJ, Xavier RJ, Farzan M, Elledge SJ (2009) The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 139(7):1243–1254. https://doi.org/10.1016/j.cell.2009.12.017
Zhao X, Guo F, Liu F, Cuconati A, Chang J, Block TM, Guo JT (2014) Interferon induction of IFITM proteins promotes infection by human coronavirus OC43. Proc Natl Acad Sci USA 111(18):6756–6761. https://doi.org/10.1073/pnas.1320856111
Ding S, Pan Q, Liu SL, Liang C (2014) HIV-1 mutates to evade IFITM1 restriction. Virology 454–455:11–24. https://doi.org/10.1016/j.virol.2014.01.020
Deeg CM, Hassan E, Mutz P, Rheinemann L, Gotz V, Magar L, Schilling M, Kallfass C, Nurnberger C, Soubies S, Kochs G, Haller O, Schwemmle M, Staeheli P (2017) In vivo evasion of MxA by avian influenza viruses requires human signature in the viral nucleoprotein. J Exp Med 214(5):1239–1248. https://doi.org/10.1084/jem.20161033
Swatek KN, Aumayr M, Pruneda JN, Visser LJ, Berryman S, Kueck AF, Geurink PP, Ovaa H, van Kuppeveld FJM, Tuthill TJ, Skern T, Komander D (2018) Irreversible inactivation of ISG15 by a viral leader protease enables alternative infection detection strategies. Proc Natl Acad Sci USA 115(10):2371–2376. https://doi.org/10.1073/pnas.1710617115
Kim YJ, Kim ET, Kim YE, Lee MK, Kwon KM, Kim KI, Stamminger T, Ahn JH (2016) Consecutive inhibition of ISG15 expression and ISGylation by cytomegalovirus regulators. PLoS Pathog 12(8):e1005850. https://doi.org/10.1371/journal.ppat.1005850
Zimmermann C, Buscher N, Krauter S, Kramer N, Wolfrum U, Sehn E, Tenzer S, Plachter B (2018) The abundant tegument protein pUL25 of human cytomegalovirus prevents proteasomal degradation of pUL26 and supports its suppression of ISGylation. J Virol. https://doi.org/10.1128/JVI.01180-18
Lee MK, Kim YJ, Kim YE, Han TH, Milbradt J, Marschall M, Ahn JH (2018) Transmembrane protein pUL50 of human cytomegalovirus inhibits ISGylation by downregulating UBE1L. J Virol. https://doi.org/10.1128/JVI.00462-18
Chen L, Sun J, Meng L, Heathcote J, Edwards AM, McGilvray ID (2010) ISG15, a ubiquitin-like interferon-stimulated gene, promotes hepatitis C virus production in vitro: implications for chronic infection and response to treatment. J Gen Virol 91(Pt 2):382–388. https://doi.org/10.1099/vir.0.015388-0
Broering R, Zhang X, Kottilil S, Trippler M, Jiang M, Lu M, Gerken G, Schlaak JF (2010) The interferon stimulated gene 15 functions as a proviral factor for the hepatitis C virus and as a regulator of the IFN response. Gut 59(8):1111–1119. https://doi.org/10.1136/gut.2009.195545
Li Y, Li S, Duan X, Chen Y, Jiao B, Ye H, Yao M, Chen L (2016) Interferon-stimulated gene 15 conjugation stimulates hepatitis B virus production independent of type I interferon signaling pathway in vitro. Mediat Inflamm 2016:7417648. https://doi.org/10.1155/2016/7417648
Symons JA, Alcami A, Smith GL (1995) Vaccinia virus encodes a soluble type I interferon receptor of novel structure and broad species specificity. Cell 81(4):551–560. https://doi.org/10.1016/0092-8674(95)90076-4
Colamonici OR, Domanski P, Sweitzer SM, Larner A, Buller RM (1995) Vaccinia virus B18R gene encodes a type I interferon-binding protein that blocks interferon alpha transmembrane signaling. J Biol Chem 270(27):15974–15978. https://doi.org/10.1074/jbc.270.27.15974
Huang J, Smirnov SV, Lewis-Antes A, Balan M, Li W, Tang S, Silke GV, Putz MM, Smith GL, Kotenko SV (2007) Inhibition of type I and type III interferons by a secreted glycoprotein from Yaba-like disease virus. Proc Natl Acad Sci USA 104(23):9822–9827. https://doi.org/10.1073/pnas.0610352104
de Marco F, Mdel M, Alejo A, Hudson P, Damon IK, Alcami A (2010) The highly virulent variola and monkeypox viruses express secreted inhibitors of type I interferon. FASEB J 24(5):1479–1488. https://doi.org/10.1096/fj.09-144733
Harris BD, Schreiter J, Chevrier M, Jordan JL, Walter MR (2018) Human interferon- and interferon-kappa exhibit low potency and low affinity for cell-surface IFNAR and the poxvirus antagonist B18R. J Biol Chem 293(41):16057–16068. https://doi.org/10.1074/jbc.RA118.003617
Alcami A, Symons JA, Smith GL (2000) The vaccinia virus soluble alpha/beta interferon (IFN) receptor binds to the cell surface and protects cells from the antiviral effects of IFN. J Virol 74(23):11230–11239. https://doi.org/10.1128/jvi.74.23.11230-11239.2000
Liptakova H, Kontsekova E, Alcami A, Smith GL, Kontsek P (1997) Analysis of an interaction between the soluble vaccinia virus-coded type I interferon (IFN)-receptor and human IFN-alpha1 and IFN-alpha2. Virology 232(1):86–90. https://doi.org/10.1006/viro.1997.8527
Golden JW, Hooper JW (2010) Evaluating the orthopoxvirus type I interferon-binding molecule as a vaccine target in the vaccinia virus intranasal murine challenge model. Clin Vaccine Immunol 17(11):1656–1665. https://doi.org/10.1128/CVI.00235-10
Montanuy I, Alejo A, Alcami A (2011) Glycosaminoglycans mediate retention of the poxvirus type I interferon binding protein at the cell surface to locally block interferon antiviral responses. FASEB J 25(6):1960–1971. https://doi.org/10.1096/fj.10-177188
Broggi A, Granucci F, Zanoni I (2020) Type III interferons: Balancing tissue tolerance and resistance to pathogen invasion. J Exp Med. https://doi.org/10.1084/jem.20190295
Xu RH, Cohen M, Tang Y, Lazear E, Whitbeck JC, Eisenberg RJ, Cohen GH, Sigal LJ (2008) The orthopoxvirus type I IFN binding protein is essential for virulence and an effective target for vaccination. J Exp Med 205(4):981–992. https://doi.org/10.1084/jem.20071854
Xu RH, Rubio D, Roscoe F, Krouse TE, Truckenmiller ME, Norbury CC, Hudson PN, Damon IK, Alcami A, Sigal LJ (2012) Antibody inhibition of a viral type 1 interferon decoy receptor cures a viral disease by restoring interferon signaling in the liver. PLoS Pathog 8(1):e1002475. https://doi.org/10.1371/journal.ppat.1002475
Hernaez B, Alonso-Lobo JM, Montanuy I, Fischer C, Sauer S, Sigal L, Sevilla N, Alcami A (2018) A virus-encoded type I interferon decoy receptor enables evasion of host immunity through cell-surface binding. Nat Commun 9(1):5440. https://doi.org/10.1038/s41467-018-07772-z
Thomas C, Moraga I, Levin D, Krutzik PO, Podoplelova Y, Trejo A, Lee C, Yarden G, Vleck SE, Glenn JS, Nolan GP, Piehler J, Schreiber G, Garcia KC (2011) Structural linkage between ligand discrimination and receptor activation by type I interferons. Cell 146(4):621–632. https://doi.org/10.1016/j.cell.2011.06.048
de Weerd NA, Vivian JP, Nguyen TK, Mangan NE, Gould JA, Braniff SJ, Zaker-Tabrizi L, Fung KY, Forster SC, Beddoe T, Reid HH, Rossjohn J, Hertzog PJ (2013) Structural basis of a unique interferon-beta signaling axis mediated via the receptor IFNAR1. Nat Immunol 14(9):901–907. https://doi.org/10.1038/ni.2667
Becker J, Kinast V, Doring M, Lipps C, Duran V, Spanier J, Tegtmeyer PK, Wirth D, Cicin-Sain L, Alcami A, Kalinke U (2018) Human monocyte-derived macrophages inhibit HCMV spread independent of classical antiviral cytokines. Virulence 9(1):1669–1684. https://doi.org/10.1080/21505594.2018.1535785
Fu X, Rivera A, Tao L, Zhang X (2012) Incorporation of the B18R gene of vaccinia virus into an oncolytic herpes simplex virus improves antitumor activity. Mol Therapy 20(10):1871–1881. https://doi.org/10.1038/mt.2012.113
Fritz-French C, Shawahna R, Ward JE, Maroun LE, Tyor WR (2014) The recombinant vaccinia virus gene product, B18R, neutralizes interferon alpha and alleviates histopathological complications in an HIV encephalitis mouse model. J Interferon Cytokine Res 34(7):510–517. https://doi.org/10.1089/jir.2013.0072
Warren L, Manos PD, Ahfeldt T, Loh YH, Li H, Lau F, Ebina W, Mandal PK, Smith ZD, Meissner A, Daley GQ, Brack AS, Collins JJ, Cowan C, Schlaeger TM, Rossi DJ (2010) Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7(5):618–630. https://doi.org/10.1016/j.stem.2010.08.012
Lee S, Liu H, Wilen CB, Sychev ZE, Desai C, Hykes BL Jr, Orchard RC, McCune BT, Kim KW, Nice TJ, Handley SA, Baldridge MT, Amarasinghe GK, Virgin HW (2019) A secreted viral nonstructural protein determines intestinal norovirus pathogenesis. Cell Host Microbe 25(6):845–857. https://doi.org/10.1016/j.chom.2019.04.005
Acknowledgements
This work was supported by grant RTI2018-094616-B-100 from the Spanish Ministerio de Ciencia e Innovación; grant S2018/BAA-4370-PLATESA2 from the Comunidad de Madrid (Fondo Europeo de Desarrollo Regional, FEDER) and VetBionet INFRAIA-731014 from the Euopean Union H2020.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rojas, J.M., Alejo, A., Martín, V. et al. Viral pathogen-induced mechanisms to antagonize mammalian interferon (IFN) signaling pathway. Cell. Mol. Life Sci. 78, 1423–1444 (2021). https://doi.org/10.1007/s00018-020-03671-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03671-z