Abstract
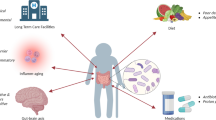
Alterations in the composition and function of the gut microbiome have been implicated in a range of conditions and diseases. Culture-dependent and culture-independent studies both showed that older people harbour a gut microbiome that differs in composition from that of younger adults. Detailed analyses have identified discrete microbiota subtypes that characterize intermediates between a high diversity microbiota found in healthy community-dwelling subjects and a low diversity microbiota typical for elderly living in long-term residential care. There are also alterations in the microbiome composition associated with biological age, independent of health status. Even after adjusting for confounding factors such as age and medication, trends in microbiota composition correlate with gradients in clinical metadata particularly frailty and inflammatory status. There are few known mechanisms by which these associations might be causative rather than consequential, and this is a subject of intensive research. The strongest candidate effectors are microbial metabolites that could impact host energy balance, act as signalling molecules to modulate host metabolism or inflammation, and potentially also impact on the gut–brain axis.

Similar content being viewed by others
References
Bonsall MB (2006) Longevity and ageing: appraising the evolutionary consequences of growing old. Philos Trans R Soc Lond B Biol Sci 361(1465):119–135
Browne HP et al (2016) Culturing of ‘unculturable’ human microbiota reveals novel taxa and extensive sporulation. Nature 533(7604):543–546
Qin J et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464(7285):59–65
Consortium, H.M.P. (2012) A framework for human microbiome research. Nature 486(7402):215–221
Falony G et al (2016) Population-level analysis of gut microbiome variation. Science 352(6285):560–564
Zhernakova A et al (2016) Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352(6285):565–569
Xie H et al (2016) Shotgun metagenomics of 250 adult twins reveals genetic and environmental impacts on the gut microbiome. Cell Syst 3(6):572 e3–584 e3
Faith JJ et al (2013) The long-term stability of the human gut microbiota. Science 341(6141):1237439
Human Microbiome Jumpstart Reference Strains C et al (2010) A catalog of reference genomes from the human microbiome. Science 328(5981):994–999
Franceschi C et al (2000) Inflamm-aging. An evolutionary perspective on immunosenescence. Ann NY Acad Sci 908:244–254
O’Toole PW, Claesson MJ (2010) Gut microbiota: changes throughout the lifespan from infancy to elderly. Int Dairy J 20:281–291
Mitsuoka T (1978) Intestinal bacteria and health. Harcourt Brace Jovanovich, Tokyo
Woodmansey EJ et al (2004) Comparison of compositions and metabolic activities of fecal microbiotas in young adults and in antibiotic-treated and non-antibiotic-treated elderly subjects. Appl Environ Microbiol 70(10):6113–6122
Fraher MH, O’Toole PW, Quigley EM (2012) Techniques used to characterize the gut microbiota: a guide for the clinician. Nat Rev Gastroenterol Hepatol 9(6):312–322
Hayashi H et al (2003) Molecular analysis of fecal microbiota in elderly individuals using 16S rDNA library and T-RFLP. Microbiol Immunol 47(8):557–570
Mueller S et al (2006) Differences in fecal microbiota in different European study populations in relation to age, gender, and country: a cross-sectional study. Appl Environ Microbiol 72(2):1027–1033
Mariat D et al (2009) The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol 9:123
Zwielehner J et al (2009) Combined PCR-DGGE fingerprinting and quantitative-PCR indicates shifts in fecal population sizes and diversity of Bacteroides, bifidobacteria and Clostridium cluster IV in institutionalized elderly. Exp Gerontol 44(6–7):440–446
Bartosch S et al (2004) Characterization of bacterial communities in feces from healthy elderly volunteers and hospitalized elderly patients by using real-time PCR and effects of antibiotic treatment on the fecal microbiota. Appl Environ Microbiol 70(6):3575–3581
Sullivan A, Edlund C, Nord CE (2001) Effect of antimicrobial agents on the ecological balance of human microflora. Lancet Infect Dis 1(2):101–114
Blaser MJ, Falkow S (2009) What are the consequences of the disappearing human microbiota? Nat Rev Microbiol 7(12):887–894
Dethlefsen L et al (2008) The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biol 6(11):e280
van Tongeren SP et al (2005) Fecal microbiota composition and frailty. Appl Environ Microbiol 71(10):6438–6442
Claesson MJ et al (2011) Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc Natl Acad Sci USA 108(Suppl 1):4586–4591
Consortium, H.M.P., The Human Microbiome Project Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486(7402):207–214
Cusack S, O’Toole PW, Consortium E (2013) Challenges and implications for biomedical research and intervention studies in older populations: insights from the ELDERMET study. Gerontology 59(2):114–121
Arumugam M et al (2011) Enterotypes of the human gut microbiome. Nature 473(7346):174–180
Wu GD et al (2011) Linking long-term dietary patterns with gut microbial enterotypes. Science 334:105–108
Flint HJ et al (2012) The role of the gut microbiota in nutrition and health. Nat Rev Gastroenterol Hepatol 9(10):577–589
Sokol H et al (2008) Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA 105(43):16731–16736
Claesson MJ et al (2012) Gut microbiota composition correlates with diet and health in the elderly. Nature 488:178–185
Drescher LS, Thiele S, Mensink GB (2007) A new index to measure healthy food diversity better reflects a healthy diet than traditional measures. J Nutr 137(3):647–651
Pryde SE et al (2002) The microbiology of butyrate formation in the human colon. FEMS Microbiol Lett 217(2):133–139
Jeffery IB et al (2012) Categorization of the gut microbiota: enterotypes or gradients? Nat Rev Microbiol 10:591–592
Biagi E et al (2010) Through ageing, and beyond: gut microbiota and inflammatory status in seniors and centenarians. PLoS ONE 5(5):e10667
Odamaki T et al (2016) Age-related changes in gut microbiota composition from newborn to centenarian: a cross-sectional study. BMC Microbiol 16:90
Kong F et al (2016) Gut microbiota signatures of longevity. Curr Biol 26(18):R832–R833
LeBlanc JG et al (2013) Bacteria as vitamin suppliers to their host: a gut microbiota perspective. Curr Opin Biotechnol 24(2):160–168
Bashan A et al (2016) Universality of human microbial dynamics. Nature 534(7606):259–262
Holmes I, Harris K, Quince C (2012) Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS ONE 7(2):e30126
Jeffery IB, Lynch DB, O’Toole PW (2016) Composition and temporal stability of the gut microbiota in older persons. ISME J 10(1):170–182
O’Toole PW, Jeffery IB (2015) Gut microbiota and aging. Science 350(6265):1214–1215
Rampelli S et al (2013) Functional metagenomic profiling of intestinal microbiome in extreme ageing. Aging 5(12):902–912
da Costa Maranduba CM et al (2015) Intestinal microbiota as modulators of the immune system and neuroimmune system: impact on the host health and homeostasis. J Immunol Res. doi:10.1155/2015/931574
Kim KA et al (2016) Gut microbiota lipopolysaccharide accelerates inflamm-aging in mice. BMC Microbiol 16:9
Jackson MA et al (2016) Signatures of early frailty in the gut microbiota. Genome Med 8(1):8
Miquel S et al (2015) Identification of metabolic signatures linked to anti-inflammatory effects of Faecalibacterium prausnitzii. MBio 6:1–10
Manzanares W et al (2016) Probiotic and synbiotic therapy in critical illness: a systematic review and meta-analysis. Crit Care 19:262
Whorwell PJ et al (2006) Efficacy of an encapsulated probiotic Bifidobacterium infantis 35624 in women with irritable bowel syndrome. Am J Gastroenterol 101(7):1581–1590
Luyer MD et al (2005) Strain-specific effects of probiotics on gut barrier integrity following hemorrhagic shock. Infect Immun 73(6):3686–3692
Riggs JE, Schochet SS Jr (1989) Osmotic stress, osmotic myelinolysis, and oligodendrocyte topography. Arch Pathol Lab Med 113(12):1386–1388
Pigneur B, Sokol H (2016) Fecal microbiota transplantation in inflammatory bowel disease: the quest for the holy grail. Mucosal Immunol 9(6):1360–1365
Hughes V (2012) Microbiome: cultural differences. Nature 492(7427):S14–S15
de la Cuesta-Zuluaga J et al (2017) Metformin is associated with higher relative abundance of mucin-degrading Akkermansia muciniphila and several short-chain fatty acid-producing microbiota in the gut. Diabetes Care 40(1):54–62
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
O’Toole, P.W., Jeffery, I.B. Microbiome–health interactions in older people. Cell. Mol. Life Sci. 75, 119–128 (2018). https://doi.org/10.1007/s00018-017-2673-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2673-z