Abstract
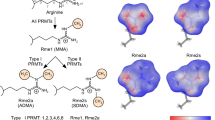
Arginine methylation of histones is one mechanism of epigenetic regulation in eukaryotic cells. Methylarginines can also be found in non-histone proteins involved in various different processes in a cell. An enzyme family of nine protein arginine methyltransferases catalyses the addition of methyl groups on arginines of histone and non-histone proteins, resulting in either mono- or dimethylated-arginine residues. The reversibility of histone modifications is an essential feature of epigenetic regulation to respond to changes in environmental factors, signalling events, or metabolic alterations. Prominent histone modifications like lysine acetylation and lysine methylation are reversible. Enzyme family pairs have been identified, with each pair of lysine acetyltransferases/deacetylases and lysine methyltransferases/demethylases operating complementarily to generate or erase lysine modifications. Several analyses also indicate a reversible nature of arginine methylation, but the enzymes facilitating direct removal of methyl moieties from arginine residues in proteins have been discussed controversially. Differing reports have been seen for initially characterized putative candidates, like peptidyl arginine deiminase 4 or Jumonji-domain containing protein 6. Here, we review the most recent cellular, biochemical, and mass spectrometry work on arginine methylation and its reversible nature with a special focus on putative arginine demethylases, including the enzyme superfamily of Fe(II) and 2-oxoglutarate-dependent oxygenases.



Similar content being viewed by others
References
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293(5532):1074–1080
Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705
Carlson SM, Gozani O (2016) Nonhistone lysine methylation in the regulation of cancer pathways. Cold Spring Harb Perspect Med 6(11):pii a026435. doi:10.1101/cshperspect.a026435
Zhang X, Huang Y, Shi X (2015) Emerging roles of lysine methylation on non-histone proteins. Cell Mol Life Sci CMLS 72(22):4257–4272
Zhang T, Cooper S, Brockdorff N (2015) The interplay of histone modifications—writers that read. EMBO Rep 16(11):1467–1481
Allis CD, Berger SL, Cote J, Dent S, Jenuwien T, Kouzarides T, Pillus L, Reinberg D, Shi Y, Shiekhattar R et al (2007) New nomenclature for chromatin-modifying enzymes. Cell 131(4):633–636
Mozzetta C, Boyarchuk E, Pontis J, Ait-Si-Ali S (2015) Sound of silence: the properties and functions of repressive Lys methyltransferases. Nat Rev Mol Cell Biol 16(8):499–513
Kooistra SM, Helin K (2012) Molecular mechanisms and potential functions of histone demethylases. Nat Rev Mol Cell Biol 13:297–311
Larsen SC, Sylvestersen KB, Mund A, Lyon D, Mullari M, Madsen MV, Daniel JA, Jensen LJ, Nielsen ML (2016) Proteome-wide analysis of arginine monomethylation reveals widespread occurrence in human cells. Sci Signal 9(443):rs9
Morales Y, Caceres T, May K, Hevel JM (2016) Biochemistry and regulation of the protein arginine methyltransferases (PRMTs). Arch Biochem Biophys 590:138–152
Chang B, Chen Y, Zhao Y, Bruick RK (2007) JMJD6 is a histone arginine demethylase. Science 318(5849):444–447
Boeckel JN, Guarani V, Koyanagi M, Roexe T, Lengeling A, Schermuly RT, Gellert P, Braun T, Zeiher A, Dimmeler S (2011) Jumonji domain-containing protein 6 (Jmjd6) is required for angiogenic sprouting and regulates splicing of VEGF-receptor 1. Proc Natl Acad Sci USA 108(8):3276–3281
Mantri M, Webby CJ, Loik ND, Hamed RB, Nielsen ML, McDonough MA, McCullagh JSO, Bottger A, Schofield CJ, Wolf A (2012) Self-hydroxylation of the splicing factor lysyl hydroxylase, JMJD6. MedChemComm 3(1):80–85
Unoki M, Masuda A, Dohmae N, Arita K, Yoshimatsu M, Iwai Y, Fukui Y, Ueda K, Hamamoto R, Shirakawa M et al (2013) Lysyl 5-hydroxylation, a novel histone modification, by jumonji domain containing 6 (JMJD6). J Biol Chem 288(9):6053–6062
Wang F, He L, Huangyang P, Liang J, Si W, Yan R, Han X, Liu S, Gui B, Li W et al (2014) JMJD6 promotes colon carcinogenesis through negative regulation of p53 by hydroxylation. PLoS Biol 12(3):e1001819
Webby CJ, Wolf A, Gromak N, Dreger M, Kramer H, Kessler B, Nielsen ML, Schmitz C, Butler DS, Yates JR 3rd et al (2009) Jmjd6 catalyses lysyl-hydroxylation of U2AF65, a protein associated with RNA splicing. Science 325(5936):90–93
Walport LJ, Hopkinson RJ, Chowdhury R, Schiller R, Ge W, Kawamura A, Schofield CJ (2016) Arginine demethylation is catalysed by a subset of JmjC histone lysine demethylases. Nat Commun 7:11974
Fisk JC, Read LK (2011) Protein arginine methylation in parasitic protozoa. Eukaryot Cell 10(8):1013–1022
Bachand F (2007) Protein arginine methyltransferases: from unicellular eukaryotes to humans. Eukaryot Cell 6(6):889–898
Zobel-Thropp P, Gary JD, Clarke S (1998) delta-N-methylarginine is a novel posttranslational modification of arginine residues in yeast proteins. J Biol Chem 273(45):29283–29286
Martens-Lobenhoffer J, Bode-Boger SM, Clement B (2016) First detection and quantification of N(delta)-monomethylarginine, a structural isomer of N(G)-monomethylarginine, in humans using MS(3). Anal Biochem 493:14–20
Wang YC, Li C (2012) Evolutionarily conserved protein arginine methyltransferases in non-mammalian animal systems. FEBS J 279(6):932–945
Low JK, Wilkins MR (2012) Protein arginine methylation in Saccharomyces cerevisiae. FEBS J 279(24):4423–4443
Wei H, Mundade R, Lange KC, Lu T (2014) Protein arginine methylation of non-histone proteins and its role in diseases. Cell Cycle 13(1):32–41
Di Lorenzo A, Bedford MT (2011) Histone arginine methylation. FEBS Lett 585(13):2024–2031
Kirmizis A, Santos-Rosa H, Penkett CJ, Singer MA, Green RD, Kouzarides T (2009) Distinct transcriptional outputs associated with mono- and dimethylated histone H3 arginine 2. Nat Struct Mol Biol 16(4):449–451
Migliori V, Muller J, Phalke S, Low D, Bezzi M, Mok WC, Sahu SK, Gunaratne J, Capasso P, Bassi C et al (2012) Symmetric dimethylation of H3R2 is a newly identified histone mark that supports euchromatin maintenance. Nat Struct Mol Biol 19(2):136–144
Olsen JV, Mann M (2013) Status of large-scale analysis of post-translational modifications by mass spectrometry. Mol Cell Proteom MCP 12(12):3444–3452
Carlson SM, Gozani O (2014) Emerging technologies to map the protein methylome. J Mol Biol 426(20):3350–3362
Doll S, Burlingame AL (2015) Mass spectrometry-based detection and assignment of protein posttranslational modifications. ACS Chem Biol 10(1):63–71
Boisvert FM, Cote J, Boulanger MC, Richard S (2003) A proteomic analysis of arginine-methylated protein complexes. Mol Cell Proteom MCP 2(12):1319–1330
Bremang M, Cuomo A, Agresta AM, Stugiewicz M, Spadotto V, Bonaldi T (2013) Mass spectrometry-based identification and characterisation of lysine and arginine methylation in the human proteome. Mol Biosyst 9(9):2231–2247
Fisk JC, Li J, Wang H, Aletta JM, Qu J, Read LK (2013) Proteomic analysis reveals diverse classes of arginine methylproteins in mitochondria of trypanosomes. Mol Cell Proteom MCP 12(2):302–311
Geoghegan V, Guo A, Trudgian D, Thomas B, Acuto O (2015) Comprehensive identification of arginine methylation in primary T cells reveals regulatory roles in cell signalling. Nat Commun 6:6758
Guo A, Gu H, Zhou J, Mulhern D, Wang Y, Lee KA, Yang V, Aguiar M, Kornhauser J, Jia X et al (2014) Immunoaffinity enrichment and mass spectrometry analysis of protein methylation. Mol Cell Proteom MCP 13(1):372–387
Lott K, Li J, Fisk JC, Wang H, Aletta JM, Qu J, Read LK (2013) Global proteomic analysis in trypanosomes reveals unique proteins and conserved cellular processes impacted by arginine methylation. J Proteom 91:210–225
Low JK, Hart-Smith G, Erce MA, Wilkins MR (2013) Analysis of the proteome of Saccharomyces cerevisiae for methylarginine. J Proteome Res 12(9):3884–3899
Ong SE, Mittler G, Mann M (2004) Identifying and quantifying in vivo methylation sites by heavy methyl SILAC. Nat Methods 1(2):119–126
Plank M, Fischer R, Geoghegan V, Charles PD, Konietzny R, Acuto O, Pears C, Schofield CJ, Kessler BM (2015) Expanding the yeast protein arginine methylome. Proteomics 15(18):3232–3243
Sylvestersen KB, Horn H, Jungmichel S, Jensen LJ, Nielsen ML (2014) Proteomic analysis of arginine methylation sites in human cells reveals dynamic regulation during transcriptional arrest. Mol Cell Proteom MCP 13(8):2072–2088
Wang K, Zhou YJ, Liu H, Cheng K, Mao J, Wang F, Liu W, Ye M, Zhao ZK, Zou H (2015) Proteomic analysis of protein methylation in the yeast Saccharomyces cerevisiae. J Proteom 114:226–233
Uhlmann T, Geoghegan VL, Thomas B, Ridlova G, Trudgian DC, Acuto O (2012) A method for large-scale identification of protein arginine methylation. Mol Cell Proteom MCP 11(11):1489–1499
Lakowski TM, Pak ML, Szeitz A, Thomas D, Vhuiyan MI, Clement B, Frankel A (2015) Arginine methylation in yeast proteins during stationary-phase growth and heat shock. Amino Acids 47(12):2561–2571
Pang CN, Gasteiger E, Wilkins MR (2010) Identification of arginine- and lysine-methylation in the proteome of Saccharomyces cerevisiae and its functional implications. BMC Genomics 11:92
Yagoub D, Hart-Smith G, Moecking J, Erce MA, Wilkins MR (2015) Yeast proteins Gar1p, Nop1p, Npl3p, Nsr1p, and Rps2p are natively methylated and are substrates of the arginine methyltransferase Hmt1p. Proteomics 15(18):3209–3218
Bedford MT, Clarke SG (2009) Protein arginine methylation in mammals: who, what, and why. Mol Cell 33(1):1–13
Le Romancer M, Treilleux I, Leconte N, Robin-Lespinasse Y, Sentis S, Bouchekioua-Bouzaghou K, Goddard S, Gobert-Gosse S, Corbo L (2008) Regulation of estrogen rapid signaling through arginine methylation by PRMT1. Mol Cell 31(2):212–221
Metivier R, Penot G, Hubner MR, Reid G, Brand H, Kos M, Gannon F (2003) Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell 115(6):751–763
Sakabe K, Hart GW (2010) O-GlcNAc transferase regulates mitotic chromatin dynamics. J Biol Chem 285(45):34460–34468
Tsai WC, Gayatri S, Reineke LC, Sbardella G, Bedford MT, Lloyd RE (2016) Arginine demethylation of G3BP1 promotes stress granule assembly. J Biol Chem 291:22671–22685. doi:10.1074/jbc.M116.739573
Wang S, Wang Y (2013) Peptidylarginine deiminases in citrullination, gene regulation, health and pathogenesis. Biochim Biophys Acta 1829(10):1126–1135
Wang Y, Wysocka J, Sayegh J, Lee YH, Perlin JR, Leonelli L, Sonbuchner LS, McDonald CH, Cook RG, Dou Y et al (2004) Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 306(5694):279–283
Raijmakers R, Zendman AJ, Egberts WV, Vossenaar ER, Raats J, Soede-Huijbregts C, Rutjes FP, van Veelen PA, Drijfhout JW, Pruijn GJ (2007) Methylation of arginine residues interferes with citrullination by peptidylarginine deiminases in vitro. J Mol Biol 367(4):1118–1129
Hidaka Y, Hagiwara T, Yamada M (2005) Methylation of the guanidino group of arginine residues prevents citrullination by peptidylarginine deiminase IV. FEBS Lett 579(19):4088–4092
Kearney PL, Bhatia M, Jones NG, Yuan L, Glascock MC, Catchings KL, Yamada M, Thompson PR (2005) Kinetic characterization of protein arginine deiminase 4: a transcriptional corepressor implicated in the onset and progression of rheumatoid arthritis. BioChemistry 44(31):10570–10582
Hausinger RP (2004) FeII/alpha-ketoglutarate-dependent hydroxylases and related enzymes. Crit Rev Biochem Mol Biol 39(1):21–68
Markolovic S, Wilkins SE, Schofield CJ (2015) Protein hydroxylation catalyzed by 2-oxoglutarate-dependent oxygenases. J Biol Chem 290:20712–20722
Loenarz C, Sekirnik R, Thalhammer A, Ge W, Spivakovsky E, Mackeen MM, McDonough MA, Cockman ME, Kessler BM, Ratcliffe PJ et al (2014) Hydroxylation of the eukaryotic ribosomal decoding center affects translational accuracy. Proc Natl Acad Sci USA 111(11):4019–4024
Singleton RS, Liu-Yi P, Formenti F, Ge W, Sekirnik R, Fischer R, Adam J, Pollard PJ, Wolf A, Thalhammer A et al (2014) OGFOD1 catalyzes prolyl hydroxylation of RPS23 and is involved in translation control and stress granule formation. Proc Natl Acad Sci USA 111(11):4031–4036
Liu W, Ma Q, Wong K, Li W, Ohgi K, Zhang J, Aggarwal AK, Rosenfeld MG (2013) Brd4 and JMJD6-associated anti-pause enhancers in regulation of transcriptional pause release. Cell 155(7):1581–1595
Lawrence P, Conderino JS, Rieder E (2014) Redistribution of demethylated RNA helicase A during foot-and-mouth disease virus infection: role of Jumonji C-domain containing protein 6 in RHA demethylation. Virology 452–453:1–11
Poulard C, Rambaud J, Hussein N, Corbo L, Le Romancer M (2014) JMJD6 regulates ERalpha methylation on arginine. PLoS One 9(2):e87982
Tikhanovich I, Kuravi S, Artigues A, Villar MT, Dorko K, Nawabi A, Roberts B, Weinman SA (2015) Dynamic arginine methylation of tumor necrosis factor (TNF) receptor-associated factor 6 regulates toll-like receptor signaling. J Biol Chem 290(36):22236–22249
Gao WW, Xiao RQ, Peng BL, Xu HT, Shen HF, Huang MF, Shi TT, Yi J, Zhang WJ, Wu XN et al (2015) Arginine methylation of HSP70 regulates retinoid acid-mediated RARbeta2 gene activation. Proc Natl Acad Sci USA 112(26):E3327–E3336
Cho JN, Ryu JY, Jeong YM, Park J, Song JJ, Amasino RM, Noh B, Noh YS (2012) Control of seed germination by light-induced histone arginine demethylation activity. Dev Cell 22(4):736–748
Feng T, Yamamoto A, Wilkins SE, Sokolova E, Yates LA, Munzel M, Singh P, Hopkinson RJ, Fischer R, Cockman ME et al (2014) Optimal translational termination requires C4 lysyl hydroxylation of eRF1. Mol Cell 53(4):645–654
Mantri M, Loik ND, Hamed RB, Claridge TD, McCullagh JS, Schofield CJ (2011) The 2-oxoglutarate-dependent oxygenase JMJD6 catalyses oxidation of lysine residues to give 5 S-hydroxylysine residues. Chembiochem Eur J Chem Biol 12(4):531–534
Hahn P, Wegener I, Burrells A, Bose J, Wolf A, Erck C, Butler D, Schofield CJ, Bottger A, Lengeling A (2010) Analysis of Jmjd6 cellular localization and testing for its involvement in histone demethylation. PLoS One 5(10):e13769
Blanc RS, Richard S (2017) Arginine methylation: the coming of age. Mol Cell 65(1):8–24
Poulard C, Corbo L, Le Romancer M (2016) Protein arginine methylation/demethylation and cancer. Oncotarget 7(41):67532–67550
Aprelikova O, Chen K, El Touny LH, Brignatz-Guittard C, Han J, Qiu T, Yang HH, Lee MP, Zhu M, Green JE (2016) The epigenetic modifier JMJD6 is amplified in mammary tumors and cooperates with c-Myc to enhance cellular transformation, tumor progression, and metastasis. Clin Epigenetics 8:38
Lee CR, Lee SH, Rigas NK, Kim RH, Kang MK, Park NH, Shin KH (2016) Elevated expression of JMJD6 is associated with oral carcinogenesis and maintains cancer stemness properties. Carcinogenesis 37(2):119–128
Lee YF, Miller LD, Chan XB, Black MA, Pang B, Ong CW, Salto-Tellez M, Liu ET, Desai KV (2012) JMJD6 is a driver of cellular proliferation and motility and a marker of poor prognosis in breast cancer. Breast Cancer Res BCR 14(3):R85
Poulard C, Rambaud J, Lavergne E, Jacquemetton J, Renoir JM, Tredan O, Chabaud S, Treilleux I, Corbo L, Le Romancer M (2015) Role of JMJD6 in breast tumourigenesis. PLoS One 10(5):e0126181
Wan J, Xu W, Zhan J, Ma J, Li X, Xie Y, Wang J, Zhu WG, Luo J, Zhang H (2016) PCAF-mediated acetylation of transcriptional factor HOXB9 suppresses lung adenocarcinoma progression by targeting oncogenic protein JMJD6. Nucleic Acids Res 44(22):10662–10675
Yang Y, Bedford MT (2013) Protein arginine methyltransferases and cancer. Nat Rev Cancer 13(1):37–50
Zhang J, Ni SS, Zhao WL, Dong XC, Wang JL (2013) High expression of JMJD6 predicts unfavorable survival in lung adenocarcinoma. Tumour Biol J Int Soc Oncodev Biol Med 34(4):2397–2401
Acknowledgements
We are grateful to Angelika Böttger and Joel Schick for useful comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wesche, J., Kühn, S., Kessler, B.M. et al. Protein arginine methylation: a prominent modification and its demethylation. Cell. Mol. Life Sci. 74, 3305–3315 (2017). https://doi.org/10.1007/s00018-017-2515-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2515-z