Abstract
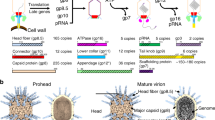
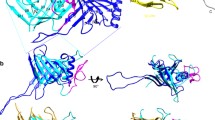
The tail of bacteriophage T4 undergoes large structural changes upon infection while delivering the phage genome into the host cell. The baseplate is located at the distal end of the contractile tail and plays a central role in transmitting the signal to the tail sheath that the tailfibers have been adsorbed by a host bacterium. This then triggers the sheath contraction. In order to understand the mechanism of assembly and conformational changes of the baseplate upon infection, we have determined the structure of an in vitro assembled baseplate through the three-dimensional reconstruction of cryo-electron microscopy images to a resolution of 3.8 Å from electron micrographs. The atomic structure was fitted to the baseplate structure before and after sheath contraction in order to elucidate the conformational changes that occur after bacteriophage T4 has attached itself to a cell surface. The structure was also used to investigate the protease digestion of the assembly intermediates and the mutation sites of the tail genes, resulting in a number of phenotypes.











Similar content being viewed by others
References
Abuladze NK, Gingery M, Tsai I, Eiserling FA (1994) Tail length determination in bacteriophage T4. Virology 199:301–310
Akhter T, Zhao L, Kohda A, Mio K, Kanamaru S, Arisaka F (2007) The neck of bacteriophage T4 is a ring-like structure formed by a hetero-oligomer of gp13 and gp14. Biochim Biophys Acta 1774(8):1036–1043
Aksyuk AA, Leiman PG, Kurochkina LP, Schneider MM, Kostyuchenko VA, Mesyanzhinov VV, Rossmann MG (2009) The tail sheath structure of bacteriophage T4: a molecular machine for infecting bacteria. EMBO J 28:821–829. doi:10.1038/emboj.2009.3
Arisaka F, Tschopp J, van Driel R, Engel J (1979) Reassembly of the bacteriophage T4 tail from the corebaseplate and the monomeric sheath protein P18: a cooperative association process. J Mol Biol 132:369–386
Arisaka F, Engel J, Klump J (1981) Contraction and dissociation of the bacteriophage T4 tail sheath induced by heat and urea. In: DuBow M (ed) Bacteriophage assembly. Alan R. Liss, New York, pp 365–379
Brenner S, Horne RW (1959) A negative staining method for high resolution electron microscopy of viruses. Biochim Biophys Acta 34:103–110
Caspar DLD (1980) Movement and self-control in protein assemblies. Quasi-equivalence revisited. Biophys J 32:103135
Coombs DH, Arisaka F (1994) T4 tail structure and function. In: Karam JD (ed) Molecular biology of bacteriophage T4. American Society for Microbiology, Washington DC, pp 259–281
Coombs DH, Eiserling FA (1977) Studies on the structure, protein composition and assembly of the neck of bacteriophage T4. J Mol Biol 116(3):375–405.
Crowther RA, Lenk EV, Kikuchi Y, King J (1977) Molecular reorganization in the hexagon to star transition of the baseplate of bacteriophage T4. J Mol Biol 116:489–523
Crowther RA (1980) Mutants of bacteriophage T4 that produce infective fibreless particles. J Mol Biol 137(2):159–74
Driedonks RA, Caldentey J (1983) Gene 20 product of bacteriophage T4. II. Its structural organization in prehead and bacteriophage. J Mol Biol 166:341–360
El Omari K, Ren J, Bird LE, Bona MK, Klarmann G, LeGrice SF, Stammers DK (2006) Molecular architecture and ligand recognition determinants for T4 RNA ligase. J Biol Chem 281(3):1573–1579
Ferguson PL, Coombs DH (2000) Pulse-chase analysis of the in vivo assembly of the Bacteriophage T4 tail. J Mol Biol 297:99–117
Fokine A, Zhang Z, Kanamaru S, Bowman VD, Aksyuk AA, Arisaka F, Rao VB, Rossmann MG (2013) The molecular architecture of the bacteriophage T4 neck. J Mol Biol 425:1731–1744. doi:10.1016/j.jmb.2013.02.012
Follansbee SE, Vanderslice RW, Chavez LG, Yegian CD (1974) A new set of adsorption mutants of bacteriophage T4D: identification of a new gene. Virology 58:180–199
Furukawa H, Mizushima S (1982) Roles of cell surface components of Escherichia coli K-12 in bacteriophage T4 infection: interaction of tail core with phospholipids. J Bacteriol 150:916–924
Furukawa H, Yamada H, Mizushima S (1979) Interaction of bacteriophage T4 with reconstituted cell envelopes of Escherichia coliK-12. J Bacteriol 140:1071–1080
Furukawa H, Kuroiwa H, Mizushima S (1983) DNA injection during bacteriophage T4 infection of Escherichia coli. J Bacteriol 154:938–945
Hall DH, Sargent RG, Trofatter KF, Russell DL (1980) Suppressors of mutations in the bacteriophage T4 gene coding for both RNA ligase and tail fiber attachment activities. J Virol 36:103–108
Hu B, Margolin W, Molineux IJ, Liu J (2015) Structural remodeling of bacteriophage T4 and host membranes during infection initiation. Proc Natl Acad Sci USA 112:E4919–E4928. doi:10.1073/pnas.1501064112
Kanamaru S, Gassner NC, Ye N, Takeda S, Arisaka F (1999) C-terminal fragment of the precursor tail lysozyme of bacteriophage T4 stays as a strucutral component of the baseplate after cleavage. J Bacteriol 181:2739–2744
Kanamaru S, Leiman PG, Kostychenco VA, Chipman PR, Mesyanzhinov VV, Arisaka F, Rossmann MG (2002) Structure of the cell-puncturing device of bacteriophage T4. Nature 415:553–557
Kao S-H, McClain WH (1980) Roles of bacteriophage T4 gene 5 and gene s products in cell lysi. J Virol 34:104–107
Kikuchi Y, King J (1975) Genetic control of bacteriophage T4 baseplate morphogenesis. l. Sequential assembly of the major precursor, in vivo and in vitro. J Mol Biol 99:645--672
King J (1971) Bacteriophage T4 tail assembly: four steps in core formation. J Mol Biol 58(3):693-709
Kostyuchenko VA, Navruzbekov GA, Kurochkina LP, Strelkov SV, Mesyanzhinov W., Rossmann MG (1999) The structure of bacteriophage T4 gene product 9: the trigger for tail contraction. Structure 7(10):1213–22
Kostyuchenko VA, Leiman PG, Chipman PR, Kanamaru S, van Raaij M, Arisaka F, Mesyanzhinov VV, Rossmann MG (2003) Three-dimensional structure of bacteriophage T4 baseplate. Nat Struct Biol 10:688–693. doi:10.1038/nsb970
Kostyuchenko VA, Chipman PR, Leiman PG, Arisaka F, Mesyanzhinov VV, Rossmann MG (2005) The tail structure of bacteriophage T4 and its mechanism of contraction. Nat Struct Mol Biol 12:810–813
Labedan B, Heller KB, Jasaitis AA, Wilson TH, Goldberg EB (1980) A membrane potential threshold for phage T4 DNA injection. Biochem Biophys Res Commun 93:625–630
Leiman PG, Kostychenko VA, Shneider MM, Kurochkina LP, Mesyanzhinov W, Rossmann MG (2000) Structure of bacteriophage T4 gene product 11, the interrace between the baseplate and short tail fibers. J Mol Biol 301(4):975–85
Leiman PG, Shneider MM, Kostyuchenko VA, Chipman PR, Mesyanzhinov W, Rossmann MG. (2003) J Mol Biol 328(4):821–33
Leiman PG, Chipman PR, Kostyuchenko VA, Mesyanzhinov VV, Rossmann MG (2004) Three-dimensional rearrangement of proteins in the tail of bacteriophage T4 on infection of its host. Cell 118:419–429. doi:10.1016/j.cell.2004.07.022
Leiman PG, Arisaka F, van Raaij MJ, Kostyuchenko VA, Aksyuk AA, Kanamaru S, Rossmann MG (2010) Morphogenesis of the T4 tail and tail fibers. Virol J 7:355–382
Moody MF (1973) Sheath of bacteriophage T4. III. Contraction mechanism deduced from partially contracted sheaths. J Mol Biol 80:613–635
Nieradko J, Koszałka P (1999) Evidence of interactions between Gp27 and Gp28 constituents of the central part of bacteriophage T4 baseplate. Acta Microbiol Pol 48:233–242
Nieradko J, Koszałka P, Krzywicka A (1998) Characteristics of gene 28 product, the constituent of the central part of bacteriophage T4 baseplate. Acta Microbiol Pol 47:243–252
Nivinskas R, Raudonikiene A, Vaishkunaite R (1990) Bacteriophage T4 gene 26. Nucleic Acids Res 18:1913
Nivinskas R, Vaiskunaite R, Dagyte R, Raudonikiene A, Klausa V (1992) Cloning, sequence, and overexpression of bacteriophage T4 gene 51. Virology 188:887–889
Scotti PD (1968) A new class of temperature conditional lethal mutants of bacteriophage T4D. Mutant Res 6:1–14
Shneider MM, Buth SA, Ho BT, Basler M, Mekalanos JJ, Leiman PG (2013) PAAR-repeat proteins sharpen and diversify the type VI secretion system spike. Nature 500:350–353
Snopek TJ, Wood WB, Conley MP, Chen P, Cozzarelli NR (1977) Bacteriophage T4 RNA ligase is gene 63 product, the protein that promotes tail fiber attachment to the baseplate. Proc Natl Acad Sci U S A 74:3355–3359
Snustad DP (1968) Dominance interactions in Escherichia coli cells mixedly infected with T4D wild-type and amber mutants and their possible implications as to type of gene-product function: catalytic vs. stoichiometric. Virology 35:550–563
Sun L, Zhang X, Gao S, Rao PA, Padilla-Sanchez V, Chen Z, Sun S, Xiang Y, Subramaniam S, Rao VB, Rossmann MG (2015) Cryo-EM structure of the bacteriophage T4 portal protein assembly at near-atomic resolution. Nat Commun 6:7548. doi:10.1038/ncomms8548
Takeda S, Hoshida K, Arisaka F (1998) Mapping of functional sites on the primary structure of the tail lysozyme of bacteriophage T4 by mutational analysis. Biochim Biophys Acta. 1384(2):243–52
Takeda Y, Suzuki M, Yamada T, Kageyama F, Arisaka F (2004) Mapping of functional sites on the primary structure of the contractile tail sheath protein of bacteriophage T4 by mutation analysis. Biochim Biophys Acta 1699:163–171
Taylor NM, Prokhorov NS, Guerrero-Ferreira RC, Shneider MM, Browning C, Goldie KN, Stahlberg H, Leiman PG (2016) Structure of the T4 baseplate and its function in triggering sheath contraction. Nature 533(7603):346–352
Tschopp J, Arisaka F, van Driel R, Engel J (1977) Purification, characterization and reassembly of the bacteriophage T4D tail sheath protein, P18. J Mol Biol 128:247–258
Yamamoto Y, Uchida H (1975) Organization and function of the tail of bacteriophage T4. II. Structural control of the tail contraction. J Mol Biol 92:207–223
Yap ML, Mio K, Leiman PG, Kanamaru S, Arisaka F (2010) The baseplate wedges of bacteriophage T4 spontaneously assemble into hubless baseplate-like structure in vitro. J Mol Biol 395:349–360
Yap ML, Klose T, Plevka P, Aksyuk A, Zhang X, Arisaka F, Rossmann MG (2014) Structure of the 3.3MDa, in vitro assembled, hubless bacteriophage T4 baseplate. J Struct Biol 187:95–102
Yap ML, Klose T, Arisaka F, Speirc JA, Veeslerc D, Fokine A, Rossmann MG (2016) The role of bacteriophage T4 baseplate in regulating assembly and infection. Proc Natl Acad Sci USA 113(10):2654–2659
Acknowledgment
The authors would like to acknowledge Professor Donald J. Winzor for his long-standing contribution to analytical ultracentrifugation on the occasion of his 80th birthday. This work was supported by JSPS KAKENHI Grant JP16087204 to FA and NIH grant AI081726 to MGR.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Fumio Arisaka declares that none of the authors have any conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by the author.
Additional information
This article is part of a Special Issue on ‘Analytical Quantitative Relations in Biochemistry’ edited by Damien Hall and Stephen Harding.
Rights and permissions
About this article
Cite this article
Arisaka, F., Yap, M.L., Kanamaru, S. et al. Molecular assembly and structure of the bacteriophage T4 tail. Biophys Rev 8, 385–396 (2016). https://doi.org/10.1007/s12551-016-0230-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-016-0230-x