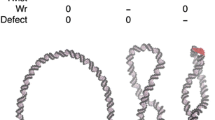
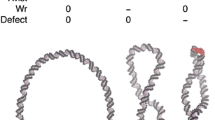
Abstract
Genomic DNA is vastly longer than the space allotted to it in a cell. The molecule must fold with a level of organization that satisfies the imposed spatial constraints as well as allow for the processing of genetic information. Key players in this organization include the negative supercoiling of DNA, which facilitates the unwinding of the double-helical molecule, and the associations of DNA with proteins, which partition the DNA into isolated loops, or domains. In order to gain insight into the principles of genome organization and to visualize the folding of spatially constrained DNA, we have developed new computational methods to identify the preferred three-dimensional pathways of protein-mediated DNA loops and to characterize the topological properties of these structures. Here, we focus on the levels of supercoiling and the spatial arrangements of DNA in model nucleoprotein systems with two topological domains. We construct these systems by anchoring DNA loops in opposing orientations on a common protein–DNA assembly, namely the Lac repressor protein with two bound DNA operators. The linked pieces of DNA form a covalently closed circle such that the protein attaches to two widely spaced sites along the DNA. We examine the effects of operator spacing, loop orientation, and long-range contacts on overall chain configuration and topology, and discuss our findings in the context of classic experiments on the effects of supercoiling and operator spacing on Lac repressor-mediated looping and recent work on the role of proteins as barriers that divide genomes into independent topological domains.







Similar content being viewed by others
Notes
If the 5′-ends of the DNA are anchored to a common operator, the former pair of loops initially points toward the inside of the repressor assembly and the latter pair toward the outside.
Concatenation of opposing loops entails removal of a common base pair from each of the bound operators, thereby reducing the number of base-pair steps in the resultant minicircle by two compared to the total number of steps in the two loops.
The choice of loop lengths is based on the difficulty of anchoring very short (<73 bp) loops to the assumed V-shaped Lac repressor model (Czapla et al. 2013) and the restraints of the minicircle on maximum loop size, i.e., 182 + 2 = 73 + 111.
Estimation of the twist from the rigid-body parameter of the same name exaggerates the unwinding at the CG step and incorrectly suggests that the repressor unwinds DNA by ~15°. Such treatment ignores the contribution to supercoiling from the large shear of base pairs at the CG step (Britton et al. 2009).
Our work to date has focused on the energies and topological properties of short DNA loops of 73–143 bp, where one can omit consideration of the thermal fluctuations of the closed structures and the accompanying effects of these changes on long-range inter-loop interactions.
References
Becker NA, Kahn JD, Maher LJ 3rd (2005) Bacterial repression loops require enhanced DNA flexibility. J Mol Biol 349:716–730
Bell CE, Lewis M (2000) A closer view of the conformation of the Lac repressor bound to operator. Nat Struct Biol 7(3):209–214
Bondarenko VA, Jiang YI, Studitsky VM (2003) Rationally designed insulator-like elements can block enhancer action in vitro. EMBO J 22(18):4728–4737
Borowiec JA, Zhang L, Sasse-Dwight S, Gralla JD (1987) DNA supercoiling promotes formation of a bent repression loop in lac DNA. J Mol Biol 196(1):101–111
Britton LA, Olson WK, Tobias I (2009) Two perspectives on the twist of DNA. J Chem Phys 131:245101
Călugăreanu G (1961) Sur les classes d’isotopie des noeuds tridimensionnels et leurs invariants. Czech Math J 11(4):588–625
Cavalli G, Misteli T (2013) Functional implications of genome topology. Nat Struct Mol Biol 20(3):290–299
Clauvelin N, Olson WK (2014) The synergy between protein positioning and DNA elasticity: energy minimization of protein-decorated DNA minicircles. Available online at: http://arxiv.org/abs/1405.7638.
Clauvelin N, Olson WK, Tobias I (2012) Characterization of the geometry and topology of DNA pictured as a discrete collection of atoms. J Chem Theory Comp 8(3):1092–1107
Clauvelin N, Lo P, Kulaeva OI, Nizovtseva EV, Diaz-Montes J, Zola J, Parashar M, Studitsky VM, Olson WK (2015) Nucleosome positioning and composition modulate in silico chromatin flexibility. J Phys Condens Matter 27:064112
Colasanti AV, Grosner MA, Perez PJ, Clauvelin N, Lu X-J, Olson WK (2013) Weak operator binding enhances simulated Lac repressor-mediated DNA looping. Biopolymers 99(12):1070–1081
Czapla L, Swigon D, Olson WK (2006) Sequence-dependent effects in the cyclization of short DNA. J Chem Theory Comp 2(3):685–695
Czapla L, Swigon D, Olson WK (2008) Effects of the nucleoid protein HU on the structure, flexibility, and ring-closure properties of DNA deduced from Monte Carlo simulations. J Mol Biol 382(2):353–370
Czapla L, Peters JP, Rueter EM, Olson WK, Maher LJ 3rd (2011) Understanding apparent DNA flexibility enhancement by HU and HMGB architectural proteins: experiment and simulation. J Mol Biol 409(2):278–289
Czapla L, Grosner MA, Swigon D, Olson WK (2013) Interplay of protein and DNA structure revealed in simulations of the lac operon. PLoS One 8(2):e56548
Dekker J, Mirny L (2016) The 3D genome as moderator of chromosomal communication. Cell 164(6):1110–1121
Dekker J, Misteli T (2015) Long-range chromatin interactions. Cold Spring Harb Perspect Biol 7(10):a019356
Dillon SC, Dorman CJ (2010) Bacterial nucleoid-associated proteins, nucleoid structure and gene expression. Nat Rev Microbiol 8(3):185–195
Eismann ER, Müller-Hill B (1990) lac Repressor forms stable loops in vitro with supercoiled wild-type lac DNA containing all three natural lac operators. J Mol Biol 213(4):763–775
Geanacopoulos M, Vasmatzis G, Zhurkin VB, Adhya S (2001) Gal repressosome contains an antiparallel DNA loop. Nat Struct Biol 8:432–436
Han L, Garcia HG, Blumberg S, Towles KB, Beausang JF, Nelson PC, Phillips R (2009) Concentration and length dependence of DNA looping in transcriptional regulation. PLoS One 4(5):e5621
Hirsh AD, Lillian TD, Lionberger TA, Perkins NC (2011) DNA modeling reveals an extended Lac repressor conformation in classic in vitro binding assays. Biophys J 101(3):718–726
Huang J, Schlick T, Vologodskii A (2001) Dynamics of site juxtaposition in supercoiled DNA. Proc Natl Acad Sci U S A 98(3):968–973
Johnson S, Lindén M, Phillips R (2012) Sequence dependence of transcription factor-mediated DNA looping. Nucleic Acids Res 40(16):7728–7738
Krämer H, Niemöller M, Amouyal M, Revêt B, von Wilcken-Bergmann B, Müller-Hill B (1987) lac repressor forms loops with linear DNA carrying two suitably spaced lac operators. EMBO J 6:1481–1491
Krämer H, Amouyal M, Nordheim A, Müller-Hill B (1988) DNA supercoiling changes the spacing requirement of two lac operators for DNA loop formation with lac repressor. EMBO J 7(2):547–556
Leng F, Chen B, Dunlap DD (2011) Dividing a supercoiled DNA molecule into two independent topological domains. Proc Natl Acad Sci U S A 108(50):19973–19978
Lewis M, Chang G, Horton NC, Kercher MA, Pace HC, Schumacher MA, Brennan RG, Lu P (1996) Crystal structure of the lactose operon repressor and its complexes with DNA and inducer. Science 271:1247–1254
Liu Z, Deibler RW, Chan HS, Zechiedrich L (2009) The why and how of DNA unlinking. Nucleic Acids Res 37(3):661–671
Lu X-J, Olson WK (2003) 3DNA: a software package for the analysis, rebuilding and visualization of three-dimensional nucleic acid structures. Nucleic Acids Res 31(17):5108–5121
Lu X-J, Olson WK (2008) 3DNA: a versatile, integrated software system for the analysis, rebuilding and visualization of three-dimensional nucleic-acid structures. Nat Protoc 3(7):1213–1227
McKay DB, Pickover CA, Steitz TA (1982) Escherichia coli lac repressor is elongated with its operator DNA binding domains located at both ends. J Mol Biol 156(1):175–183
Normanno D, Vanzi F, Pavone FS (2008) Single-molecule manipulation reveals supercoiling-dependent modulation of Lac repressor-mediated DNA looping. Nucleic Acids Res 36(8):2505–2513
Olson WK, Gorin AA, Lu X-J, Hock LM, Zhurkin VB (1998) DNA sequence-dependent deformability deduced from protein–DNA crystal complexes. Proc Natl Acad Sci U S A 95:11163–11168
Peck LJ, Wang JC (1981) Sequence dependence of the helical repeat of DNA in solution. Nature 292(5821):375–378
Perez PJ, Clauvelin N, Grosner MA, Colasanti AV, Olson WK (2014) What controls DNA looping? Int J Mol Sci 15(9):15090–15108
Postow L, Hardy CD, Arsuaga J, Cozzarelli NR (2004) Topological domain structure of the Escherichia coli chromosome. Genes Dev 18(14):1766–1779
Rhodes D, Klug A (1981) Sequence-dependent helical periodicity of DNA. Nature 292(5821):378–380
Ruben GC, Roos TB (1997) Conformation of Lac repressor tetramer in solution, bound and unbound to operator DNA. Microsc Res Tech 36:400–416
Spronk CAEM, Folkers GE, Noordman A-MGW, Wechselberger R, van den Brink N, Boelens R, Kaptein R (1999) Hinge–helix formation and DNA bending in various Lac repressor–operator complexes. EMBO J 18:6472–6480
Steitz TA, Richmond TJ, Wise D, Engelman D (1974) The lac repressor protein: molecular shape, subunit structure, and proposed model for operator interaction based on structural studies of microcrystals. Proc Natl Acad Sci U S A 71(3):593–597
Swigon D, Olson WK (2008) Mesoscale modeling of multi-protein-DNA assemblies: the role of the catabolic activator protein in Lac-repressor-mediated looping. Int J Non Linear Mech 43:1082–1093
Swigon D, Coleman BD, Olson WK (2006) Modeling the Lac repressor–operator assembly: the influence of DNA looping on Lac repressor conformation. Proc Natl Acad Sci U S A 103(26):9879–9884
Taraban M, Zhan H, Whitten AE, Langley DB, Matthews KS, Swint-Kruse L, Trewhella J (2008) Ligand-induced conformational changes and conformational dynamics in the solution structure of the lactose repressor protein. J Mol Biol 376(2):466–481
Tobias I, Coleman BD, Olson WK (1994) The dependence of DNA tertiary structure on end conditions: theory and implications for topological transitions. J Chem Phys 101(12):10990–10996
Wang JC (1979) Helical repeat of DNA in solution. Proc Natl Acad Sci U S A 76(1):200–203
White JH (1969) Self-linking and the Gauss integral in higher dimensions. Am J Math 91(3):693–728
White JH, Bauer WR (1987) Superhelical DNA with local substructures. A generalization of the topological constraint in terms of the intersection number and the ladder-like correspondence surface. J Mol Biol 195(1):205–213
Whitson PA, Hsieh W-T, Wells RD, Matthews KS (1987) Supercoiling facilitates lac operator–repressor–pseudooperator interactions. J Biol Chem 262(11):4943–4946
Acknowledgments
This work was generously supported by the U.S. Public Health Service under research grant GM34809. P.J.P. gratefully acknowledges support from a U.S. Department of Education Graduate Assistance in Areas of National Need Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
Pamela J. Perez declares that she has no conflict of interest.
Wilma K. Olson declares that she has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed byany of the authors.
Additional information
This article is part of a Special Issue on “DNA supercoiling, protein interactions and genetic function” edited by Laura Finzi and Wilma Olson.
Rights and permissions
About this article
Cite this article
Perez, P.J., Olson, W.K. Insights into genome architecture deduced from the properties of short Lac repressor-mediated DNA loops. Biophys Rev 8 (Suppl 1), 135–144 (2016). https://doi.org/10.1007/s12551-016-0209-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-016-0209-7