Abstract
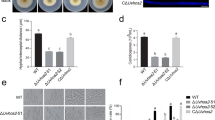
Histone acetylation/deacetylation represent a general and efficient epigenetic mechanism through which fungal cells control gene expression. Here we report developmental requirement of MoHOS2-mediated histone deacetylation (HDAC) for the rice blast fungus, Magnaporthe oryzae. Structural similarity and nuclear localization indicated that MoHOS2 is an ortholog of Saccharomyces cerevisiae Hos2, which is a member of class I histone deacetylases and subunit of Set3 complex. Deletion of MoHOS2 led to 25% reduction in HDAC activity, compared to the wild-type, confirming that it is a bona-fide HDAC. Lack of MoHOS2 caused decrease in radial growth and impinged dramatically on asexual sporulation. Such reduction in HDAC activity and phenotypic defects of ΔMohos2 were recapitulated by a single amino acid change in conserved motif that is known to be important for HDAC activity. Expression analysis revealed up-regulation of MoHOS2 and concomitant down-regulation of some of the key genes involved in asexual reproduction under sporulation-promoting condition. In addition, the deletion mutant exhibited defect in appressorium formation from both germ tube tip and hyphae. As a result, ΔMohos2 was not able to cause disease symptoms. Wound-inoculation showed that the mutant is compromised in its ability to grow inside host plants as well. We found that some of ROS detoxifying genes and known effector genes are de-regulated in the mutant. Taken together, our data suggest that MoHOS2-dependent histone deacetylation is pivotal for proper timing and induction of transcription of the genes that coordinate developmental changes and host infection in M. oryzae.
Similar content being viewed by others
References
Adachi, N., Kimura, A., and Horikoshi, M. 2002. A conserved motif common to the histone acetyltransferase Esa1 and the histone deacetylase Rpd3. J. Biol. Chem.277, 35688–35695.
Bannister, A.J. and Kouzarides, T. 2011. Regulation of chromatin by histone modifications. Cell Res.21, 381–395.
Brosch, G., Loidl, P., and Graessle, S. 2008. Histone modifications and chromatin dynamics: A focus on filamentous fungi. FEMS Microbiol. Rev.32, 409–439.
Chi, M.H., Park, S.Y., and Lee, Y.H. 2009. A quick and safe method for fungal DNA extraction. Plant Pathol.J. 25, 108–111.
Dean, R., Van Kan, J.A., Pretorius, Z.A., Hammond-Kosack, K.E., Di Pietro, A., Spanu, P.D., Rudd, J.J., Dickman, M., Kahmann, R., Ellis, J., et al. 2012. The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol.13, 414–430.
Ding, S.L., Liu, W., Iliuk, A., Ribot, C., Vallet, J., Tao, A., Wang, Y., Lebrun, M.H., and Xu, J.R. 2010. The Tig1 histone deacetylase complex regulates infectious growth in the rice blast fungus Magnaporthe oryzae. Plant Cell22, 2495–2508.
Elias-Villalobos, A., Fernandez-Alvarez, A., Moreno-Sanchez, I., Helmlinger, D., and Ibeas, J.I. 2015. The Hos2 histone deacetylase controls ustilago maydis virulence through direct regulation of mating-type genes. PLoS Pathog.11, e1005134.
Fernandez, J. and Orth, K. 2018. Rise of a cereal killer: The biology of Magnaporthe oryzae biotrophic growth. Trends Microbiol.26, 582–597.
Grewal, C., Hickmott, J., Rentas, S., and Karagiannis, J. 2012. A conserved histone deacetylase with a role in the regulation of cytokinesis in Schizosaccharomyces pombe. Cell Div.7, 13.
Howard, R.J. and Valent, B. 1996. Breaking and entering: Host penetration by the fungal rice blast pathogen Magnaporthe grisea. Annu. Rev. Microbiol.50, 491–512.
Huh, A., Dubey, A., Kim, S., Jeon, J., and Lee, Y.H. 2017. MoJMJ1, encoding a histone demethylase containing JmjC domain, is required for pathogenic development of the rice blast fungus, Magnaporthe oryzae. Plant Pathol. J.33, 193–205.
Huh, A., Dubey, A., Kim, S., Jeon, J., and Lee, Y.H. 2014. Histone acetylation in fungal pathogens of plants. Plant Pathol. J.30, 1–9.
Huh, A., Dubey, A., Kim, S., Jeon, J., and Lee, Y.H. 2018. Mitogen-activated protein kinase signaling in plant pathogenic fungi. PLoS Pathog.14, e1006875.
Kim, S., Park, S.Y., Kim, K.S., Rho, H.S., Chi, M.H., Choi, J., Park, J., Kong, S., Park, J., Goh, J., et al., Y.H. 2009. Homeobox transcription factors are required for conidiation and appressorium development in the rice blast fungus Magnaporthe oryzae. PLoS Genet.5, e1000757.
Kwon, S., Lee, J., Jeon, J., Kim, S., Park, S.Y., Jeon, J., and Lee, Y.H. 2018. Role of the histone acetyltransferase Rtt109 in development and pathogenicity of the rice blast fungus. Mol. Plant Microbe Interact.31, 1200–1210.
Lau, G.W. and Hamer, J.E. 1998. Acropetal: A genetic locus required for conidiophore architecture and pathogenicity in the rice blast fungus. Fungal Genet. Biol.24, 228–239.
Lee, K.K. and Workman, J.L. 2007. Histone acetyltransferase complexes: One size doesn't fit all. Nat. Rev. Mol. Cell. Biol.8, 284–295.
Lengeler, K.B., Davidson, R.C., D'Souza, C., Harashima, T., Shen, W.C., Wang, P., Pan, X., Waugh, M., and Heitman, J. 2000. Signal transduction cascades regulating fungal development and virulence. Microbiol. Mol. Biol. Rev.64, 746–785.
Li, Y., Wang, C., Liu, W., Wang, G., Kang, Z., Kistler, H.C., and Xu, J.R. 2011. The HDF1 histone deacetylase gene is important for conidiation, sexual reproduction, and pathogenesis in Fusarium graminearum. Mol. Plant Microbe Interact.24, 487–496.
Matheis, S., Yemelin, A., Scheps, D., Andresen, K., Jacob, S., Thines, E., and Foster, A.J. 2017. Functions of the Magnaporthe oryzae Flb3p and Flb4p transcription factors in the regulation of conidiation. Microbiol. Res.196, 106–117.
Millar, C.B., Kurdistani, S.K., and Grunstein, M. 2004. Acetylation of yeast histone H4 lysine 16: A switch for protein interactions in heterochromatin and euchromatin. Cold Spring Harb. Symp. Quant. Biol.69, 193–200.
Mulder, N.J. and Apweiler, R. 2008. The InterPro database and tools for protein domain analysis. Curr. Protoc. Bioinformatics Chapter 2, Unit 2.7.
Orbach, M.J., Farrall, L., Sweigard, J.A., Chumley, F.G., and Valent, B. 2000. A telomeric avirulence gene determines efficacy for the rice blast resistance gene Pi-ta. Plant Cell12, 2019–2032.
Park, J., Kim, S., Kwon, S., and Lee, Y.H. 2014. A quick and accurate screening method for fungal gene-deletion mutants by direct, priority-based, and inverse PCRs. J. Microbiol. Methods105, 39–41.
Pham, K.T., Inoue, Y., Vu, B.V., Nguyen, H.H., Nakayashiki, T., Ikeda, K., and Nakayashiki, H. 2015. MoSet1 (histone H3K4 methyltransferase in Magnaporthe oryzae) regulates global gene expression during infection-related morphogenesis. PLoS Genet.11, e1005385.
Pidroni, A., Faber, B., Brosch, G., Bauer, I., and Graessle, S. 2018. A class 1 histone deacetylase as major regulator of secondary metabolite production in Aspergillus nidulans. Front. Microbiol.9, 2212.
Pijnappel, W.W., Schaft, D., Roguev, A., Shevchenko, A., Tekotte, H., Wilm, M., Rigaut, G., Seraphin, B., Aasland, R., and Stewart, A.F. 2001. The S. cerevisiae Set3 complex includes two histone deacetylases, Hos2 and Hst1, and is a meiotic-specific repressor of the sporulation gene program. Genes Dev.15, 2991–3004.
Rundlett, S.E., Carmen, A.A., Kobayashi, R., Bavykin, S., Turner, B.M., and Grunstein, M. 1996. Hda1 and Rpd3 are members of distinct yeast histone deacetylase complexes that regulate silencing and transcription. Proc. Natl. Acad. Sci. USA93, 14503–14508.
Ryder, L.S. and Talbot, N.J. 2015. Regulation of appressorium development in pathogenic fungi. Curr. Opin. Plant Biol.26, 8–13.
Selin, C., de Kievit, T.R., Belmonte, M.F., and Fernando, W.G. 2016. Elucidating the role of effectors in plant-fungal interactions: Progress and challenges. Front. Microbiol.7, 600.
Seto, E. and Yoshida, M. 2014. Erasers of histone acetylation: The histone deacetylase enzymes. Cold Spring Harb. Perspect. Biol.6, a018713.
Sievers, F. and Higgins, D.G. 2018. Clustal omega for making accurate alignments of many protein sequences. Protein Sci.27, 135–145.
Soyer, J.L., El Ghalid, M., Glaser, N., Ollivier, B., Linglin, J., Grandaubert, J., Balesdent, M.H., Connolly, L.R., Freitag, M., Rouxel, T., et al. 2014. Epigenetic control of effector gene expression in the plant pathogenic fungus Leptosphaeria maculans. PLoS Genet.10, e1004227.
Talbot, N.J. 2003. On the trail of a cereal killer: Exploring the biology of Magnaporthe grisea. Annu. Rev. Microbiol.57, 177–202.
Tessarz, P. and Kouzarides, T. 2014. Histone core modifications regulating nucleosome structure and dynamics. Nat. Rev. Mol. Cell. Biol.15, 703–708.
Wang, A., Kurdistani, S.K., and Grunstein, M. 2002. Requirement of Hos2 histone deacetylase for gene activity in yeast. Science298, 1412–1414.
Wilson, R.A. and Talbot, N.J. 2009. Under pressure: Investigating the biology of plant infection by Magnaporthe oryzae. Nat. Rev. Microbiol.7, 185–195.
Wiren, M., Silverstein, R.A., Sinha, I., Walfridsson, J., Lee, H.M., Laurenson, P., Pillus, L., Robyr, D., Grunstein, M., and Ekwall, K. 2005. Genome-wide analysis of nucleosome density histone acetylation and hdac function in fission yeast. EMBO J.24, 2906–2918.
Zhang, S. and Xu, J.R. 2014. Effectors and effector delivery in Magnaporthe oryzae. PLoS Pathog.10, e1003826.
Acknowledgements
This work was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (NRF-2017R1D1A3B03033932).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Supporting Information
Rights and permissions
About this article
Cite this article
Lee, J., Lee, JJ. & Jeon, J. A histone deacetylase, MoHOS2 regulates asexual development and virulence in the rice blast fungus. J Microbiol. 57, 1115–1125 (2019). https://doi.org/10.1007/s12275-019-9363-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-019-9363-5