Abstract
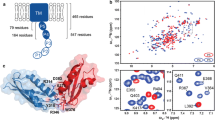
We report the 1H, 13C and 15N backbone chemical shift assignments and secondary structure of the Escherichia coli protein BamC, a 32-kDa protein subunit that forms part of the BAM (Omp85) complex, the β-barrel assembly machinery present in all Gram-negative bacteria and which is essential for viability.


Similar content being viewed by others
References
Bos MP, Robert V, Tommassen J (2007) Biogenesis of the gram-negative bacterial outer membrane. Annu Rev Microbiol 61:191–214. doi:10.1146/annurev.micro.61.080706.093245
Breyton C, Haase W, Rapoport TA, Kuhlbrandt W, Collinson I (2002) Three-dimensional structure of the bacterial protein-translocation complex SecYEG. Nature 418:662–665. doi:10.1038/nature00827
Cornilescu G, Delaglio F, Bax A (1999) Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J Biomol NMR 13:289–302. doi:10.1023/A:1008392405740
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. doi:10.1007/BF00197809
Driessen AJ, Nouwen N (2008) Protein translocation across the bacterial cytoplasmic membrane. Annu Rev Biochem 77:643–667. doi:10.1146/annurev.biochem.77.061606.160747
Goddard TD, Kneller DG, SPARKY 3. University of California, San Francisco
Knowles TJ, Jeeves M, Bobat S, Dancea F, McClelland D et al (2008) Fold and function of polypeptide transport-associated domains responsible for delivering unfolded proteins to membranes. Mol Microbiol 68:1216–1227. doi:10.1111/j.1365-2958.2008.06225.x
Metzler WJ, Constantine KL, Friedrichs MS, Bell AJ, Ernst EG et al (1993) Characterization of the three-dimensional solution structure of human profilin: 1H, 13C, and 15N NMR assignments and global folding pattern. Biochemistry 32:13818–13829. doi:10.1021/bi00213a010
Misra R (2007) First glimpse of the crystal structure of YaeT’s POTRA domains. ACS Chem Biol 2:649–651. doi:10.1021/cb700212p
Muhandiram DR, Kay LE (1994) Gradient-enhanced triple-resonance three-dimensional NMR experiments with improved sensitivity. J Magn Reson B 103:14. doi:10.1006/jmrb.1994.1032
Onufryk C, Crouch ML, Fang FC, Gross CA (2005) Characterization of six lipoproteins in the sigmaE regulon. J Bacteriol 187:4552–4561. doi:10.1128/JB.187.13.4552-4561.2005
Rizzitello AE, Harper JR, Silhavy TJ (2001) Genetic evidence for parallel pathways of chaperone activity in the periplasm of Escherichia coli. J Bacteriol 183:6794–6800. doi:10.1128/JB.183.23.6794-6800.2001
Sklar JG, Wu T, Gronenberg LS, Malinverni JC, Kahne D, Silhavy TJ (2007a) Lipoprotein SmpA is a component of the YaeT complex that assembles outer membrane proteins in Escherichia coli. Proc Natl Acad Sci USA 104:6400–6405. doi:10.1073/pnas.0701579104
Sklar JG, Wu T, Kahne D, Silhavy TJ (2007b) Defining the roles of the periplasmic chaperones SurA, Skp, and DegP in Escherichia coli. Genes Dev 21:2473–2484. doi:10.1101/gad.1581007
Van den Berg B, Clemons WM Jr, Collinson I, Modis Y, Hartmann E et al (2004) X-ray structure of a protein-conducting channel. Nature 427:36–44. doi:10.1038/nature02218
Vuong P, Bennion D, Mantei J, Frost D, Misra R (2008) Analysis of YfgL and YaeT interactions through bioinformatics, mutagenesis, and biochemistry. J Bacteriol 190:1507–1517. doi:10.1128/JB.01477-07
Wimley WC (2003) The versatile beta-barrel membrane protein. Curr Opin Struct Biol 13:404–411. doi:10.1016/S0959-440X(03)00099-X
Wishart DS, Sykes BD (1994) The 13C chemical-shift index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J Biomol NMR 4:171–180. doi:10.1007/BF00175245
Wu T, Malinverni J, Ruiz N, Kim S, Silhavy TJ, Kahne D (2005) Identification of a multicomponent complex required for outer membrane biogenesis in Escherichia coli. Cell 121:235–245. doi:10.1016/j.cell.2005.02.015
Acknowledgments
We thank Christian Ludwig, Sara Whittaker and the other staff of The Henry Wellcome Building for Biomolecular NMR Spectroscopy, which is funded by the Wellcome Trust. This work was supported by the BBSRC and EU PRISM project [T.J.K and M.O.].
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Knowles, T.J., McClelland, D.M., Rajesh, S. et al. Secondary structure and 1H, 13C and 15N backbone resonance assignments of BamC, a component of the outer membrane protein assembly machinery in Escherichia coli . Biomol NMR Assign 3, 203–206 (2009). https://doi.org/10.1007/s12104-009-9175-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-009-9175-3