Abstract
Major conceptual roadblocks impede the development of an HIV-1 vaccine that can stimulate a potent neutralizing antibody response. Animal models that support HIV-1 replication and allow for host genetic manipulation would be an ideal platform for testing various immunological hypotheses, but progress on this research front has been slow and disappointing. In contrast, many valuable concepts emerged from more than 50 years of studying the Friend retrovirus model. This was recently exemplified by the identification of an innate restriction gene, Apobec3, that could promote the retrovirus-specific neutralizing antibody response. Here we review both classical and recent data on humoral immunity against Friend retrovirus infection, and highlight the potential of this model for unraveling novel aspects of the retrovirus-specific antibody response that may guide HIV-1 vaccine development efforts.
Similar content being viewed by others
Introduction
A vaccine against HIV-1 is a global health priority, but remains a difficult challenge. An HIV-1 vaccine should elicit potent adaptive immune responses that include both cell-mediated and neutralizing antibody (NAb) responses. However, the best means to elicit these responses remain unclear. Recent discoveries suggested that valuable insights may be gained by understanding the earliest events following infection, when the innate arm of the immune response is at the critical juncture of shaping adaptive immunity. However, studies on HIV-infected individuals are limited by difficulties in obtaining samples at the time of transmission and the inability to directly control specific biological and genetic factors. It is therefore not surprising that many concepts that guided HIV vaccine development can be traced to basic discoveries in virus infection models.
The current gold standard for AIDS pathogenesis and vaccine studies is the SIV/rhesus macaque model [1]. SIV and HIV-1 are phylogenetically distinct, but both are lentiviruses that cause CD4+ T cell depletion, resulting in opportunistic infections, a hallmark of AIDS. Rhesus macaques can be infected with defined molecular clones of SIV, and the development of immunity in the context of viral pathogenesis can be precisely monitored. However, several factors that include high cost, elaborate infrastructure, long duration of follow-up, lack of monkey-specific reagents and inability to genetically modify the host have limited its widespread utility.
Mouse models, on the other hand, allow access to inbred strains that can be genetically modified, not to mention a plethora of reagents and protocols to dissect immune responses. Major efforts were therefore made to develop mice that are susceptible to HIV-1. One approach is to genetically “knock-in” human genes in mice to promote HIV-1 replication, but this effort proved quite difficult with the existence of multiple restriction blocks [2]. Another approach is to engraft a functional human immune system in immunocompromised mice [3]. However, the cost, variability in engraftment, outbred genetics, immaturity of the adaptive immune response and inability to completely “knock-out” specific genes currently limit their full utility for immunological studies. Thus, retroviruses that cause pathology in “normal” mice may be the most suitable starting point for basic vaccine discovery and proof-of-concept immunological studies.
Friend retrovirus infection as a model system to study retrovirus-specific humoral immunity
Friend retrovirus (FV) infection is one of the most well-characterized models of pathogenic retrovirus infection in adult, immunocompetent mice [4–6]. FV is a complex of two retroviruses: (1) a replication-competent, but nonpathogenic, Friend murine leukemia virus (F-MuLV) and (2) a replication-defective, but pathogenic, spleen focus-forming virus (SFFV). SFFV cannot produce viral particles on its own and therefore relies on F-MuLV-encoded particles to carry its genome. In contrast to HIV-1, which is a complex retrovirus encoding multiple accessory genes in addition to Gag, Pol and Env, FV is a simple retrovirus and is a member of the gammaretrovirus family. Similarities and differences between FV and HIV-1 are highlighted in Table 1.
FV infection of susceptible mice results in chronic viremia, polycythemia and splenomegaly within 2 weeks and erythroleukemia and death starting 1–2 months post-infection [4–6]. Polycythemia and splenomegaly are caused by SFFV. SFFV encodes an altered form of Env known as gp55P, a glycoprotein that associates and activates the erythropoietin receptor (EpoR) [7]. This process is dependent on Fv2, a dominant susceptibility gene that encodes a short form of the Stk kinase [8]. The signaling events orchestrated by gp55P, EpoR and Fv2 result in the rapid expansion of erythroblasts that are highly susceptible to FV infection [9, 10]. Subsequent erythroleukemia is linked to proviral integration events into the host genome near proto-oncogenes and tumor suppressor genes [11, 12].
Until 2008 [13], practically all FV studies were conducted with “classical” FV stocks containing lactate dehydrogenase elevating virus (LDV). LDV is an RNA virus endemic in wild mouse populations and belongs to the Arteriviridae family [14]. LDV does not significantly affect acute FV infection levels [13], but it could strongly stimulate Type I Interferon (IFN) responses [15, 16]. LDV has been shown to delay CD8+ T cell immunity [13], induce polyclonal activation in B cells and slow the development of NAb responses [17, 18]. Interestingly, LDV was reported in the FV stocks as early as the 1960s [19, 20], but was largely ignored. While there is growing interest to move the field toward “LDV-free” FV stocks, the results obtained with LDV-free FV should be viewed with caution if the intent is to build on results from historical data. To directly link new work with previous FV/LDV studies, the use of FV/LDV stocks are recommended initially, followed by infections with LDV-free stocks for comparison. In this review, recent studies performed with LDV-free FV stocks will be noted with a bulletpoint (•).
While HIV-1 and FV are clearly different retroviruses, the FV model continues to provide basic information on the retrovirus-specific cell-mediated immune response (reviewed in [21, 22]). Here we argue that FV infection is also a tractable model to dissect basic mechanisms underlying the retrovirus-specific humoral immune response. The humoral response to FV is relevant to HIV-1 for several reasons. First, both the HIV-1 and FV envelope (Env) glycoproteins are the primary targets of the NAb response in their respective hosts. Similar to the HIV-1 Env, the MLV Env gp85 glycoprotein protomer consists of a surface (SU; gp70) and transmembrane (TM; p15E) subunit, which form tripod-like trimers on the virus particle based on cryoelectron tomography studies [23, 24]. Second, both HIV-1 and FV propagate in immune organs and/or compartments, resulting in humoral immune dysfunction. B cell abnormalities that include polyclonal B cell activation, hypergammaglobulinemia and delayed NAb development were observed in both HIV and FV infection (Table 1). Finally, a single dominant resistance gene that promotes the FV-specific NAb response, Recovery from Friend Virus 3 (Rfv3) [25], has a direct counterpart in humans that is counteracted by HIV-1. It took 30 years to decipher the molecular identity of Rfv3 [26], but the result of these efforts has opened the door for exciting studies that compare the development of a protective versus a nonprotective humoral response in congenic mice.
Recovery from Friend retrovirus involves the humoral arm of the immune response
Since the discovery of FV by Charlotte Friend in 1957 [27], it was already apparent that host genetic factors had a tremendous impact on FV resistance and susceptibility. This led to the identification of Fv1 and Fv2 by Frank Lilly and colleagues in the 1960s [28, 29], which correspond to nonimmunological genes that control N- and B-tropic FV tropism and susceptibility to splenomegaly, respectively. In addition, immunological genes that influenced recovery from viremia and disease were identified, notably Recovery from Friend retrovirus 1 and 2 (Rfv1 and Rfv2) which both map to the major histocompatibility complex (H-2) [30–32]. The H-2 locus was identified as an important determinant for recovery when it was observed that mice with the H-2 b/b genotype recovered from splenomegaly at a higher frequency than mice with the H-2 b/d genotype in a backcross study between C57BL/6 (H-2 b/b) and DBA/2 (H-2 d/d) strains [30]. However, B1 progeny from crosses between A.BY (H-2 b/b) and BALB/c (H-2 d/d) mice all failed to recover from FV infection. This led to the discovery of a non-H-2 recovery gene, Rfv3 [25]. Rfv3 encoded a gene that suppressed viremia and lowered the frequencies of virus-producing cells via the production of NAbs [33]. An example of the genetic cross that revealed Rfv3 is shown in Fig. 1a.
Rfv3 is encoded by Apobec3. a Rfv3 genetics. A.BY mice are susceptible to FV-induced splenomegaly due to a dominant Fv2 susceptibility gene. Crossing A.BY mice to C57BL strains (B6 or B10) results in an F1 strain that should still be susceptible to splenomegaly. However, these F1 mice recover from viremia about 1 month post-infection. Recovery from viremia correlated with NAb responses and segregated as a Mendelian trait in subsequent backcrosses with A.BY mice. B6 and A.BY mice are H-2 b/b, suggesting that a single autosomal dominant, non-H-2 gene controls recovery from viremia and NAb responses. This gene was referred to as Rfv3. b F1 transcomplementation to test if Rfv3 is Apobec3. B6 Apobec3 KO mice were crossed to A.BY mice to obtain F1 strains similar to panel A. If Rfv3 ≠ Apobec3, then the Rfv3 phenotypes of recovery from viremia and NAb responses should be similar with that of (B6 × A.BY)F1 mice (panel A). However, the opposite result was observed: (B6 Apobec3 KO × A.BY)F1 strains had high levels of viremia, did not recover from splenomegaly and had weak NAb responses. Thus, the B6 Apobec3 gene was critical for conferring Rfv3 resistance. This was taken as direct proof that Rfv3 is encoded by Apobec3
Rfv3 is encoded by Apobec3
The first major clue to the molecular identity of Rfv3 came when Kim Hasenkrug and colleagues mapped Rfv3 to a 20-centimorgan region of mouse chromosome 15 in 1995 [34]. Four years later, the same group narrowed Rfv3 to a 0.83-centimorgan region [35]. In 2005, Masaaki Miyazawa and colleagues further cut the Rfv3 locus by half [36]. By this time, two major developments have occurred: (1) the near-complete sequencing of the mouse genome [37]; and (2) the identification of a potent antiretroviral restriction factor, APOBEC3G [38]. The sequencing of the mouse genome revealed that the Rfv3 locus encompass just 61 genes. The identification of APOBEC3G as the factor counteracted by the HIV-1 protein Vif provided strong support that the murine homologue, mouse Apobec3, may encode Rfv3.
To directly test whether Apobec3 encodes Rfv3, our group employed an F1 transcomplementation approach [26] (Fig. 1b). B6 wild-type (WT) and Apobec3 knockout (KO) mice were crossed to two Rfv3 susceptible strains (A.BY and BALB/c) to generate Fv2 r/s (Fv2 susceptible) F1 strains that differ only in the presence or absence of the B6 Apobec3 gene. If the B6 Apobec3 gene is the Rfv3 resistance gene, then removal of the B6 Apobec3 gene should result in Rfv3 susceptibility in the F1 strains. On the other hand, if Apobec3 is not Rfv3, then inactivation of the B6 Apobec3 gene will not influence viremia and NAb responses. Our results revealed that the B6 Apobec3 gene indeed influenced NAb responses and recovery from viremia in F1 mice, providing direct evidence that Apobec3 encodes Rfv3. This was further strengthened with additional cohorts showing the strong influence of B6 Apobec3 on survival, NAb responses and recovery from viremia, cellular infection and splenomegaly [39]. To date, the Apobec3 association with NAb responses has been reproduced in pure B6, 129/OlaHsdP2 (129/Ola), (B6 × A.BY)F1 and (B6 × BALB/c)F1 [26, 39, 40] and (B6 × A/WySn)F1 strains [41•].
Recently, a linked gene that controls B cell survival, the B cell activating factor receptor (BAFF-R), was suggested to control the Rfv3 susceptibility of A/WySn mice [41•]. However, a BAFF-R defect could not account for the Rfv3 susceptible phenotype of BALB and A.BY strains, which have normal B cell levels [39]. Given that the initial description of Rfv3 encompassed C57BL, A/WySn, A.BY and BALB strains [25], we concluded that Apobec3 encodes Rfv3.
Apobec3 properties that correlate with Rfv3 resistance and susceptibility
Significant progress has been made in determining which Apobec3 property accounts for the Rfv3 resistance and susceptible phenotype. The B6 Apobec3 gene predominantly encodes a splice variant lacking exon 5 [26, 42–44] that appears to have more potent activity [42, 43] and higher resistance to the viral protease [42] than the full-length BALB Apobec3 gene in vitro. Amino acid residues that differed between B6 and BALB strains were predicted to have undergone positive selection, suggesting a role in differential virus restriction [45]. Recently, minigene analyses revealed that the B6 Δ5 transcript was translated more efficiently than the full-length BALB transcript [46]. Thus, Apobec3 Δ5 splicing is likely important in dictating the Rfv3 phenotype.
Another striking difference between Rfv3 resistant (B6) and susceptible (A.BY, BALB, A/WySn) strains is the steady state levels of total Apobec3 transcripts, which could vary by up to 20-fold [39, 43, 47, 48]. In contrast to the antiviral properties of the Apobec3 Δ5 variant that were studied in co-transfected cells, the difference in Apobec3 mRNA expression was derived from primary cells. Thus, Apobec3 mRNA expression is likely the fundamental property that distinguishes Apobec3 between Rfv3 resistant and susceptible strains. The only exception appears to be the 129/Ola strain, which was initially classified as Rfv3 resistant [26] but upon re-analysis, had Apobec3 mRNA levels similar to that of A.BY, BALB and A/WySn strains (D. Smith and M. Santiago, unpublished). 129/Ola mice were also classified as Fv2 resistant due to high resistance to splenomegaly even with very high FV inoculum. However, the FV stock used for infections was B-tropic [26], and 129/Ola mice are in fact Fv1 nr [49, 50]. Thus, B-tropic FV likely encountered a potent Fv1 post-entry block in 129/Ola mice and served as a “live-attenuated vaccine” that elicited a potent NAb response, resulting in its erroneous designation as Rfv3 resistant (see Part III.D.). Infection of 129/Ola strains with dual-tropic (NB-tropic) FV resulted in splenomegaly, chronic viremia and weak NAb responses (K. Halemano, B. Barrett and M. Santiago, unpublished). Thus, 129/Ola mice should be classified as Fv2 s/s Rfv3 s/s Fv1 nr, linking the Rfv3 susceptible status of 129/Ola mice with its low Apobec3 mRNA expression levels.
The mA3 polymorphism(s) responsible for the difference in mA3 mRNA levels between Rfv3 resistant and susceptible strains remains unknown. Sequencing of the mA3 locus between Rfv3 resistant and susceptible strains revealed a 530-bp xenotropic MLV long-terminal repeat (X-MLV LTR) insertion just after the exon 2 splice site in B6 but not BALB or A.BY mice [39, 45]. Interestingly, we detected Apobec3 transcripts that lacked exon 2 in Rfv3 susceptible A.BY and BALB mice [26]. These Δ2 transcripts represent only a minor fraction (<10 %) of the total Apobec3 mRNA pool in the spleen [39, 47, 51], as observed for the human homologues APOBEC3G and APOBEC3F [52]. However, Apobec3 Δ2 transcripts may represent the remnants of actively degraded transcripts due to the lack of an X-MLV LTR. This hypothesis is currently being tested.
The link between Apobec3 mRNA expression levels and Rfv3 could have important implications, especially since B6 Apobec3 restriction is saturable in vivo [40]. In the recombinant CasFrKP MLV strain, an alternatively translated FV-derived gene product, Glyco-Gag, appears to function as an antagonist of Apobec3 [49], analogous to HIV-1 Vif antagonizing human APOBEC3G. Thus, Apobec3 overexpression may “swamp out” the antagonist and unleash the antiviral activity of Apobec3 in vivo. Interestingly, higher APOBEC3G mRNA expression levels correlated with HIV-1 resistance in humans and nonhuman primates [53, 54]. Thus, in addition to the discovery of compounds that can block the HIV-1 Vif–APOBEC3G interaction, upregulating APOBEC3G levels may also be a viable therapeutic strategy.
Mechanism for how Apobec3 promotes the NAb response
The identification of Rfv3 as Apobec3 immediately raised the issue of how Apobec3 promotes NAb responses. We formulated two hypotheses [26]: a direct and indirect mechanism. The direct mechanism proposes that Apobec3 may directly affect antigen-specific antibody development even in the absence of FV infection. This derives from the fact that Apobec3 is highly expressed in B cells [41, 55] and is evolutionarily related to activation-induced cytidine deaminase [56], an enzyme that catalyzes antibody affinity maturation and class switching [57]. However, B6 Apobec3 deficiency was also found to have a significant impact on infectious viremia during acute infection. This was observed not only in FV infection [26], but also in an earlier study with mouse mammary tumor virus [58], as well as Moloney MLV [59] and the CasFrKP MLV [49]. Thus, Apobec3 may enhance NAb responses through an indirect mechanism that involves limiting virus-induced humoral immune dysfunction.
To delineate which mechanism is more relevant, we utilized an immunization approach that allows for quantitative assessments of antibody affinity based on reactivity to low- versus high-molar conjugation of nitrophenyl (NP) hapten. B6 Apobec3 KO mice produced antibodies of similar affinity to NP compared with B6 WT mice. Only in the context of FV infection did the production of high-affinity antibodies to NP-hapten become impaired in B6 Apobec3 KO mice [55]. Miyazawa and colleagues also found no impact of B6 Apobec3 on antibody responses to dinitrophenyl (DNP)-Ficoll and DNP-Ovalbumin [41]. Together, these results suggest that Apobec3 modulates antibody production via an indirect mechanism.
Apobec3 may promote the NAb response by limiting FV-induced immune dysfunction. B6 Apobec3 WT F1 mice have significantly lower FV+ erythroid, myeloid, B and T cells compared with B6 Apobec3 KO F1 mice [55], suggesting that protection of these cell subsets from FV infection may limit immune dysfunction. Mice infected with FV have decreased abilities to produce antibodies against sheep red blood cells, and this suppression was dependent on the loss of IFN-γ and the increased IL-10 production [60]. Whether IL-10 dependent immunosuppression is enhanced in B6 Apobec3 KO mice remains untested. Interestingly, B6 Apobec3 WT mice displayed polyclonal B cell activation [41•] and a significant delay in hypergammaglobulinemia [55]. Thus, FV-induced humoral immune dysfunction may be directly linked to perturbations in B cell function. So far, B cell immunophenotyping studies at 3 days post-infection (dpi) revealed no Apobec3-dependent perturbations in early B cell subsets in the bone marrow (K. Halemano and M. Santiago, unpublished). In contrast, at 7 dpi, we observed significantly increased induction of germinal center (GC) B cells and plasmablasts in B6 Apobec3 deficient mice [55]. In fact, the kinetics of FV-specific IgG and NAb development was enhanced if the mice encoded the B6 Apobec3 gene [26, 41•]. Thus, in addition to preventing FV-induced immune dysfunction, Apobec3 may also drive virus-specific antibody development.
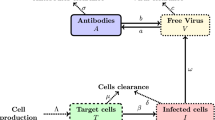
Maintenance of antibody responses is highly dependent on antigen levels [61–63]. Since most restriction factors inhibit viruses in the infected cell and decrease antigen levels, these factors may not be optimal for driving B cell responses. In contrast, Apobec3 appears to have evolved to maintain antigen levels during acute infection. In cell culture, the APOBEC3 proteins do not inhibit virus production, but inhibit the infectivity of the resulting virions [38]. This was thought to occur by incorporation of Apobec3 into nascent virions, after which, Apobec3 inhibits reverse transcription in the next target cell [64] (Fig. 2). Investigating 7 dpi plasma samples from B6 Apobec3 WT and KO F1 mice, we provided the first evidence that this phenomenon occurs in vivo [40]. Similar viral RNA loads were found in plasma of B6 WT and Apobec3-deficient mice, while the infectivity of these virions differed markedly [40]. Thus, B6 Apobec3 activity was associated with the release of noninfectious particles during acute FV infection, accounting for up to 80 % of the total virion pool relative to B6 Apobec3 KO mice. These noninfectious particles are fusion-competent and could serve as immunogens to prime the B cell response. Priming the B cell response provides a strong rationale for why mammalian hosts evolved a restriction factor that inhibits retroviruses in the next target cell (Fig. 2).
Current model for the Apobec3-dependent NAb response. (Left) In Rfv3 resistant mice, higher levels of the B6 Apobec3 gene results in the release of noninfectious particles that stimulate GC B cells via the TLR7-Myd88 pathway, resulting in a strong NAb response. (Right) In Rfv3 susceptible mice, released infectious particles result in virus replication and immune dysfunction, resulting in a weaker NAb response. Note that during acute infection, similar numbers of FV particles are released independent of B6 Apobec3 activity. Not shown are the infection of other cell types, particularly erythroblasts, myeloid cells and T cells which could all contribute to humoral immune dysfunction in Apobec3/Rfv3 susceptible mice
Innate immune recognition of retrovirus particles
Apobec3-restricted virions may be potent inducers of humoral immunity because they maintain critical immunogenic properties of infectious virions that include size, multivalency and uptake kinetics [65], but without promoting cellular dysfunction due to virus replication. In addition, virion nucleic acids have the capacity to stimulate pattern recognition receptors that may directly activate B cells. Browne et al. [66•] demonstrated that the NAb response against FV was completely abrogated in B6 MyD88 KO mice, suggesting an essential role for Toll-like receptor (TLR) signaling. B cell intrinsic MyD88, as opposed to dendritic cell intrinsic MyD88, was found to be crucial to the antibody response to FV [67•]. TLR7 was also shown to be essential to the production of NAbs and the induction of GCs [67•, 68]. Collectively, these data suggest that B cell intrinsic TLR7 signaling is essential for NAb development against retroviruses. Because TLR7 recognizes single-stranded (ss) RNA in endosomes, it is possible that when a B cell that recognizes FV Env binds to virions, endocytosis of the virion particle exposes TLR7 to the viral ssRNA, leading to a GC response (Fig. 2). In this way, Apobec3-restricted virions may be acting like “virus-like particle (VLP)” vaccines. Interestingly, we recently found that Apobec3 is the primary direct effector of IFN-α treatment against FV infection in vivo (M. Harper, B. Barrett and M. Santiago, unpublished). Thus, the TLR7-Myd88 dyad may directly activate Apobec3 through Type I IFN induction, providing a potential amplification loop. However, the Type I IFN receptor does not appear to be critical for NAb induction [68], implying the existence of alternative pathways for intrinsic priming of the B cell response.
The protective antibody response against Friend retrovirus
A global HIV-1 vaccine should elicit NAbs that can broadly neutralize different HIV-1 strains. Progress in this area is rapid and exciting, and revealed several broadly neutralizing Abs (bNAbs) and their respective epitopes in the HIV-1 Env trimer (for example, see [69]). However, designing an immunogen to elicit these bNAbs remains a major challenge and would likely require basic knowledge on how to activate B cells with a specific germline configuration, focus the GC reaction to strongly disfavored Env trimer epitopes and induce high-affinity antibodies through extensive somatic hypermutation.
In 2009, the first evidence of protection from HIV-1 acquisition was observed in the RV144 trial in Thailand [70]. Surprisingly, protection did not correlate with strong NAbs or CD8+ T cell responses. Instead, the moderate 31.2 % protection correlated with HIV-1 Env-specific binding antibodies that appear to have Fc effector functions. In addition to direct neutralization, antibodies have the capacity to counteract infected cells through (1) complement-mediated lysis, (2) opsonization for macrophage phagocytosis and (3) antibody-dependent cellular cytotoxicity (ADCC) mediated by natural killer (NK) cells. These understudied properties of antibodies have now become a major focus of HIV/AIDS research [71, 72].
Lessons from FV-specific monoclonal antibodies
While the threat of the HIV-1/AIDS pandemic was unraveling in the early 1980 s, the first attempt to characterize the antibody response against a pathogenic retrovirus using monoclonal antibodies (MAbs) was being undertaken. Bruce Chesebro and colleagues produced hundreds of hybridomas from FV-infected Rfv3 resistant mice, 17 of which were reactive to FV+ erythroleukemia cells [73]. The viral protein targets of 11 of these MAbs were identified: 8 reacted to gp70, while 3 reacted to p15E and Glyco-Gag. None of these FV-reactive MAbs neutralized F-MuLV in vitro. However, in the presence of guinea pig complement, 7 out of 8 FV gp70-reactive MAbs lysed FV virions and 4 out of 6 FV-reactive MAbs lysed infected cells in vitro [73, 74]. These four cytotoxic MAbs included MAbs 48 and 350 (IgG2a), MAb 34 (IgG2b) and MAb 307 (IgM). However, only the IgG2a MAbs conferred significant reduction in spleen infectious centers (ICs) following passive immunization in vivo [74].
The most potent MAb from this initial panel, MAb 48, was further used in in vivo studies. In contrast to the in vitro data, MAb 48 administration did not lead to clearance of FV-infected cells. Instead, MAb 48 appeared to inhibit the expansion of virus-producing cells [74]. Subsequent studies demonstrated that cell-mediated immune responses were essential to the in vivo effectiveness of MAb 48 [75, 76]. Interestingly, MAb 34, a nonneutralizing MAb specific to Glyco-Gag, enhanced the antiviral activity of MAb 48 [76].
In 1992, another MAb, MAb A9D41, was demonstrated to lyse Friend leukemia cells in vitro in a complement-dependent manner [77]. Removal of the Fc portion of MAb A9D41 completely abrogated its ability to inhibit the replication of Friend leukemia cells injected into DBA/2 mice. Moreover, passive protection mediated by MAb A9D41 was significantly reduced in NK-deficient bg/bg mice compared with bg/+ mice, suggesting a role for NK-mediated ADCC in the in vivo potency of MAb A9D41.
Together, these studies on MAbs 48 and A9D41 suggested the involvement of complement, T cells, NK cells and nonneutralizing Abs in inhibiting FV in vivo. However, these MAbs represent only a small sampling of the entire repertoire of FV-specific antibodies. Thus, these MAbs may not recapitulate the dominant mechanism(s) governing the Rfv3-dependent, FV-specific NAb response.
Studies on the FV-specific polyclonal antibody (pAb) response
Passive immunization with anti-FV serum into susceptible FV-infected mice increased survival rates [78] and dramatically lowered infection levels in splenocytes [33, 66, 79]. The combined activity of antibodies in these “bulk” antisera may better capture the most critical mechanisms governing retrovirus antibody neutralization. However, such mechanisms have yet to be studied in-depth.
The identification of Rfv3 as Apobec3 provided a unique opportunity to investigate the characteristics of a protective versus a nonprotective pAb response against retroviruses in congenic mice with or without the B6 Apobec3 gene. We are in the process of analyzing 28 dpi plasma samples from (B6 × BALB/c)F1 and (B6 × A.BY)F1 mice for isotype and IgG subclass distribution, avidity to various FV virion preparations and ultimately, effector functions through passive immunization in susceptible mice. A simple approach is to treat purified FV virions with 1 % detergent, which disrupts native trimers of HIV-1 Env [80, 81]. Using this approach, we found that 28 dpi IgM and IgG endpoint binding titers to detergent-treated FV Env were equivalent between B6 Apobec3 WT and KO F1 mice. In contrast, the IgG endpoint titers to native FV preparations were significantly higher in F1 mice with the B6 Apobec3 gene [40]. Thus, B6 Apobec3 activity (or Rfv3 resistance) is associated with the development of IgG antibodies specific to B cell epitopes on native virions, for example, functional trimers (Fig. 2).
Our ongoing analyses reveal that the protective Apobec3-dependent IgG response may be biased to the IgG2 subclass (K. Heilman and M. Santiago, unpublished). Since subclass designations are dependent on the Fc portion of IgG that interacts with Fcγ receptors, we hypothesize that antibody neutralization in vivo will have a significant Fc-mediated effector component. To test this hypothesis, we performed passive immunization studies with 28 dpi plasma from B6 Apobec3 WT and KO F1 mice with FV into susceptible BALB/c mice. Our results revealed significant inhibition of FV infection in the bone marrow with B6 Apobec3 WT F1 plasma but not B6 Apobec3 KO F1 plasma (K. Halemano, B. Barrett and M. Santiago, unpublished). Thus, an in vivo neutralization assay recapitulated the Rfv3 phenotype, and should allow for future studies on antibody effector mechanisms.
Vaccine modalities to elicit FV-specific NAb
Most vaccines work through the induction of NAbs, and FV is no exception. Thus, the FV model has been used to investigate the ability of various retroviral vaccine preparations to elicit NAb-based protection. Many of these NAb-based studies pre-dated large HIV-1 vaccine trials, while some have been designed after HIV-1 or SIV vaccine studies, particularly in the context of developing T cell vaccines. We review here some of these studies, highlighting the efficacy of various vaccine configurations and their ability to elicit NAbs.
Inactivated virus vaccines
Using formalin-fixed serum preparations, Charlotte Friend was the first to demonstrate that mice can be protected from FV infection with a vaccine [78]. However, it eventually became clear that formalin-fixed, heat-inactivated, or UV-inactivated virus were poor immunogens. Using inactivated virus without an adjuvant offers moderate protection against FV-induced splenomegaly in H-2 b/b mice while offering no protection to H-2 a/a mice, which coincided with a lack of NAbs [82, 83]. Poor protection with inactivated F-MuLV may be due to the lack of TLR stimulation, as implicated in the unfortunate formalin-inactivated respiratory syncytial virus vaccine trial in 1966 [84]. It is possible that treatments that render the virus inactive also denature or destroy TLR agonists. Supporting this contention, the addition of adjuvants to inactivated F-MuLV improved protection levels [82, 85, 86]. However, even though these immunized mice were able to produce FV Env-specific IgG, they still did not produce NAbs [82]. A pair of mutations within the “immunosuppression domain” of FV Env renders UV-inactivated virions more immunogenic, resulting in increased TM- and Gag-specific IgG and activated FV-specific T cells [87]. However, it was not shown whether these mutations resulted in a more potent NAb response.
Subunit vaccines
Purification of the FV Env in the 1970s allowed for testing a subunit vaccine approach. Early studies in STU mice demonstrated that immunization with FV Env gp70 (SU) or gp85 (SU + TM) in either complete or incomplete Freund’s adjuvant could significantly lower plasma viremia, splenomegaly and mortality in a dose-dependent manner [88, 89]. Immunization with gp85 conferred better protection against FV-induced mortality than gp70, which induced better protection than p15E (TM) [89]. Gag proteins (i.e., matrix, capsid and nucleocapsid) offered no protection from splenomegaly [82]. Hunsmann et al. [89] also demonstrated that gp70 coupled to keyhole limpet hemocyanin effectively protected mice from low-dose FV challenge. Although vaccinations with gp70 were able to produce FV-specific IgG binding titers, they were not able to produce neutralizing titers prior to challenge [82]. It remains unknown whether Fc effector mechanisms were responsible for the moderate protection levels observed with these subunit FV vaccines. The recent RV144 HIV-1 vaccine trials used a gp120 subunit boost that resulted in moderate protection levels [70], and it is currently thought that Fc effector mechanisms may have contributed to protection [71, 72].
Vaccinia and adenovirus vectors
Current HIV/SIV vaccine strategies focus on the use of vaccinia and adenovirus (Ad) vectors as delivery systems for antigens. While the intent was to elicit CD8+ cytotoxic T cell (CTL) immunity, priming of CD4+ T cell immunity may also enhance NAb responses through these vaccine vectors.
Immunization with recombinant vaccinia expressing FV Env gp85 resulted in rapid recovery from splenomegaly and spleen ICs in H-2 a/b mice but conferred very little protection in H-2 a/a mice [85, 90, 91]. Interestingly, immunization with vaccinia-FV Env seemed to protect H-2 b/b mice by T lymphocyte priming, which led to a more rapid NAb response and potent CTL response [85]. Removal of CTL epitopes from vaccinia-FV Env or depletion of CD8+ T lymphocytes from mice vaccinated with complete vaccinia-FV Env still resulted in recovery from splenomegaly for H-2 a/b mice, suggesting that rapid induction of the IgG NAb response can compensate for a missing CTL response [92]. Vaccinia-FV Gag also conferred some protection, but the recovery from splenomegaly and induction of the NAb response was slower than in mice immunized with vaccinia-FV Env [93]. Thus, the importance of the retroviral Env gene product in a vaccine was already highlighted in early FV studies, a concept that was unfortunately dismissed in the failed human Phase IIb STEP trials that did not include the HIV-1 Env [94].
Replication-defective adenovirus vectors were also tested in the FV model. In addition to Ad5-based vectors expressing FV Env and Gag, fiber chimeric vectors, in which the fiber knob and shaft domains of Ad5 were replaced with those of Ad35 (Ad5F35), were produced [95]. Heterologous prime-boost vaccination with Ad5 and Ad5F35 resulted in reduced plasma viremia and splenomegaly compared with homologous prime-boost, and this protection correlated with a significantly increased NAb response [95]. This finding agrees with assessments in the SIV model that prior immunity to Ad vectors could significantly inhibit vaccine potency [96, 97]. More recently, the addition of IFN-α subtype 4 to the Ad vaccine increased CD4+ T cell activation and NAb titers to the point that CD8+ T cells can be depleted with no increases to plasma viremia or spleen size [98]. This recent study suggests that Type I IFN could be an effective adjuvant to augment NAb-based vaccine efficacy.
Live-“attenuated” vaccines
By far, the best vaccination outcome in the FV model involved the administration of live N-tropic F-MuLV (FV-N) in Fv1 b/b mice. Fv1 encodes an endogenous retroviral Gag gene that determines tropism by interfering with the proviral integration of certain FV strains [99]. FV-N viruses replicate in Fv1 n/n NIH Swiss mice while FV-B viruses replicate in Fv1 b/b BALB or B6 strains. However, when an FV-N virus is used to infect an Fv1 b/b mouse, the virus acts as a live-“attenuated” vaccine and induces NAbs, FV-specific CTL activity and FV-specific T cell proliferation prior to challenge [85]. These parameters correlate with transient levels of spleen ICs at 12 days post-challenge and complete protection from splenomegaly [76, 83, 85].
Consistent with the success of live-attenuated vaccines in the FV model, the best vaccination outcome observed in the SIV model involved SIV with a deletion in the accessory protein Nef [100]. SIVΔNef viruses replicate at low levels in the host, likely driving a potent CD8+ T cell response. These attenuated viruses should also somewhat preserve the CD4+ T cell response and thereby improve SIV antibody development. Interestingly, the SIV Vif protein, which antagonizes the respective simian APOBEC3 proteins, can be mutated to generate a live-attenuated virus [101]. Studies are ongoing to determine if SIVΔNef and SIV Vif-attenuated viruses can elicit potent neutralizing or blocking antibodies.
Virus-like particles
While highly potent, live-attenuated vaccines for HIV-1 may not be clinically feasible because of serious safety concerns. Our current working model (Fig. 2) on Apobec3-restricted noninfectious particles driving the B cell response suggests that VLP configurations may be a suitable alternative vaccine to elicit NAbs. These VLPs should ideally preserve the functional, fusion-competent Env trimer and contain nucleic acids that can prime the B cell response through TLR signaling. HIV-1 pseudovirions expressing Gag and Env have the potential to express epitopes of the native virion trimer [102]. VLP-based vaccine regimens offer advantages to other approaches since it could circumvent the need for adjuvants and will not be subject to concerns on prior immunity to the vaccine vector and potential reversion from live attenuation. We anticipate that the FV model will play an important role in dissecting the induction of humoral immunity using the VLP vaccine platform.
Concluding remarks
The FV infection model was initially used to understand basic mechanisms involved in oncogenesis. However, concepts derived from more than 5 decades of work in the FV model are now proving to be quite relevant to HIV-1 infection. This was most recently shown by the discovery of pathways that connect the innate and humoral arms of the immune response against retroviruses. These studies have set the stage for the modern use of the FV model in investigating basic mechanisms underlying the retroviral antibody response in the context of natural infection and vaccination.
References
Shedlock DJ, Silvestri G, Weiner DB. Monkeying around with HIV vaccines: using rhesus macaques to define ‘gatekeepers’ for clinical trials. Nat Rev Immunol. 2009;9(10):717–28.
Zhang JX, Diehl GE, Littman DR. Relief of preintegration inhibition and characterization of additional blocks for HIV replication in primary mouse T cells. PLoS ONE. 2008;3(4):e2035.
Denton PW, Garcia JV. Novel humanized murine models for HIV research. Curr HIV/AIDS Rep. 2009;6(1):13–9.
Chesebro B, Miyazawa M, Britt WJ. Host genetic control of spontaneous and induced immunity to Friend murine retrovirus infection. Annu Rev Immunol. 1990;8:477–99.
Hasenkrug KJ, Chesebro B. Immunity to retroviral infection: the Friend virus model. PNAS USA. 1997;94(15):7811–6.
Miyazawa M, Tsuji-Kawahara S, Kanari Y. Host genetic factors that control immune responses to retrovirus infections. Vaccine. 2008;26(24):2981–96.
Li JP, D’Andrea AD, Lodish HF, Baltimore D. Activation of cell growth by binding of friend spleen focus-forming virus gp55 glycoprotein to the erythropoietin receptor. Nature. 1990;343(6260):762–4.
Persons DA, Paulson RF, Loyd MR, Herley MT, Bodner SM, Bernstein A, et al. Fv2 encodes a truncated form of the Stk receptor tyrosine kinase. Nat Genet. 1999;23(2):159–65.
Nishigaki K, Hanson C, Jelacic T, Thompson D, Ruscetti S. Friend spleen focus-forming virus transforms rodent fibroblasts in cooperation with a short form of the receptor tyrosine kinase Stk. PNAS USA. 2005;102(43):15488–93.
Finkelstein LD, Ney PA, Liu QP, Paulson RF, Correll PH. Sf-Stk kinase activity and the Grb2 binding site are required for Epo-independent growth of primary erythroblasts infected with Friend virus. Oncogene. 2002;21(22):3562–70.
Moreau-Gachelin F, Tavitian A, Tambourin P. Spi-1 is a putative oncogene in virally induced murine erythroleukaemias. Nature. 1988;331(6153):277–80.
Munroe DG, Peacock JW, Benchimol S. Inactivation of the cellular p53 gene is a common feature of Friend virus-induced erythroleukemia: relationship of inactivation to dominant transforming alleles. Mol Cell Biol. 1990;10(7):3307–13.
Robertson SJ, Ammann CG, Messer RJ, Carmody AB, Myers L, Dittmer U, et al. Suppression of acute anti-friend virus CD8+ T-cell responses by coinfection with lactate dehydrogenase-elevating virus. J Virol. 2008;82(1):408–18.
Li K, Schuler T, Chen Z, Glass GE, Childs JE, Plagemann PG. Isolation of lactate dehydrogenase-elevating viruses from wild house mice and their biological and molecular characterization. Virus Res. 2000;67(2):153–62.
Ammann CG, Messer RJ, Peterson KE, Hasenkrug KJ. Lactate dehydrogenase-elevating virus induces systemic lymphocyte activation via TLR7-dependent IFNalpha responses by plasmacytoid dendritic cells. PLoS ONE. 2009;4(7):e6105.
Gerlach N, Schimmer S, Weiss S, Kalinke U, Dittmer U. Effects of type I interferons on Friend retrovirus infection. J Virol. 2006;80(7):3438–44.
Marques R, Antunes I, Eksmond U, Stoye J, Hasenkrug K, Kassiotis G. B lymphocyte activation by coinfection prevents immune control of friend virus infection. J Immunol. 2008;181(5):3432–40.
Bila C, Oberhauser V, Ammann CG, Ejaz A, Huber G, Schimmer S, et al. Complement opsonization enhances friend virus infection of B cells and thereby amplifies the virus-specific CD8+ T cell response. J Virol. 2011;85(2):1151–5.
Riley V. Enzymatic determination of transmissible replicating factors associated with mouse tumors. Ann N Y Acad Sci. 1963;100:762–90.
Ebert PS, Chirigos MA, Fields LA, Ellsworth PA. Plasma lactate dehydrogenase and spleen heme biosynthetic activity following Friend and Rauscher leukemia virus infections. Life Sci. 1967;6(18):1963–71.
Hasenkrug KJ, Dittmer U. The role of CD4 and CD8 T cells in recovery and protection from retroviral infection: lessons from the Friend virus model. Virology. 2000;272(2):244–9.
Hasenkrug KJ, Dittmer U. Immune control and prevention of chronic Friend retrovirus infection. Front Biosci. 2007;12:1544–51.
Forster F, Medalia O, Zauberman N, Baumeister W, Fass D. Retrovirus envelope protein complex structure in situ studied by cryo-electron tomography. PNAS USA. 2005;102(13):4729–34.
Zhu P, Liu J, Bess J Jr, Chertova E, Lifson JD, Grise H, et al. Distribution and three-dimensional structure of AIDS virus envelope spikes. Nature. 2006;441(7095):847–52.
Chesebro B, Wehrly K. Identification of a non-H-2 gene (Rfv-3) influencing recovery from viremia and leukemia induced by Friend virus complex. PNAS USA. 1979;76(1):425–9.
Santiago ML, Montano M, Benitez R, Messer RJ, Yonemoto W, Chesebro B, et al. Apobec3 encodes Rfv3, a gene influencing neutralizing antibody control of retrovirus infection. Science. 2008;321(5894):1343–6.
Friend C. Cell-free transmission in adult Swiss mice of a disease having the character of a leukemia. J Exp Med. 1957;105(4):307–18.
Lilly F. Susceptibility to two strains of Friend leukemia virus in mice. Science. 1967;155(761):461–2.
Lilly F. Fv-2: identification and location of a second gene governing the spleen focus response to Friend leukemia virus in mice. J Natl Cancer Inst. 1970;45(1):163–9.
Lilly F. The effect of histocompatibility-2 type on response to friend leukemia virus in mice. J Exp Med. 1968;127(3):465–73.
Chesebro B, Wehrly K, Stimpfling J. Host genetic control of recovery from Friend leukemia virus-induced splenomegaly: mapping of a gene within the major histocompatability complex. J Exp Med. 1974;140(6):1457–67.
Chesebro B, Wehrly K. Rfv-1 and Rfv-2, two H-2-associated genes that influence recovery from Friend leukemia virus-induced splenomegaly. J Immunol. 1978;120(4):1081–5.
Doig D, Chesebro B. Anti-Friend virus antibody is associated with recovery from viremia and loss of viral leukemia cell-surface antigens in leukemic mice. Identification of Rfv-3 as a gene locus influencing antibody production. J Exp Med. 1979;150(1):10–9.
Hasenkrug KJ, Valenzuela A, Letts VA, Nishio J, Chesebro B, Frankel WN. Chromosome mapping of Rfv3, a host resistance gene to Friend murine retrovirus. J Virol. 1995;69(4):2617–20.
Super HJ, Hasenkrug KJ, Simmons S, Brooks DM, Konzek R, Sarge KD, et al. Fine mapping of the friend retrovirus resistance gene, Rfv3, on mouse chromosome 15. J Virol. 1999;73(9):7848–52.
Kanari Y, Clerici M, Abe H, Kawabata H, Trabattoni D, Caputo SL, et al. Genotypes at chromosome 22q12-13 are associated with HIV-1-exposed but uninfected status in Italians. AIDS. 2005;19(10):1015–24.
Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, et al. Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;420(6915):520–62.
Sheehy AM, Gaddis NC, Choi JD, Malim MH. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature. 2002;418(6898):646–50.
Santiago ML, Smith DS, Barrett BS, Montano M, Benitez RL, Pelanda R, et al. Persistent Friend virus replication and disease in Apobec3-deficient mice expressing functional B-cell-activating factor receptor. J Virol. 2011;85(1):189–99.
Smith DS, Guo K, Barrett BS, Heilman KJ, Evans LH, Hasenkrug KJ, et al. Noninfectious retrovirus particles drive the Apobec3/Rfv3 dependent neutralizing antibody response. PLoS Pathog. 2011;7(10):e1002284.
Tsuji-Kawahara S, Chikaishi T, Takeda E, Kato M, Kinoshita S, Kajiwara E, et al. Persistence of viremia and production of neutralizing antibodies differentially regulated by polymorphic APOBEC3 and BAFF-R loci in Friend virus-infected mice. J Virol. 2010;84(12):6082–95.
Abudu A, Takaori-Kondo A, Izumi T, Shirakawa K, Kobayashi M, Sasada A, et al. Murine retrovirus escapes from murine APOBEC3 via two distinct novel mechanisms. Curr Biol. 2006;16(15):1565–70.
Takeda E, Tsuji-Kawahara S, Sakamoto M, Langlois MA, Neuberger MS, Rada C, et al. Mouse APOBEC3 restricts friend leukemia virus infection and pathogenesis in vivo. J Virol. 2008;82(22):10998–1008.
Browne EP, Littman DR. Species-specific restriction of Apobec3-mediated hypermutation. J Virol. 2008;82(3):1305–13.
Sanville B, Dolan MA, Wollenberg K, Yan Y, Martin C, Yeung ML, et al. Adaptive evolution of Mus Apobec3 includes retroviral insertion and positive selection at two clusters of residues flanking the substrate groove. PLoS Pathog. 2010;6:e1000974.
Li J, Hakata Y, Takeda E, Liu Q, Iwatani Y, Kozak CA, et al. Two genetic determinants acquired late in Mus evolution regulate the inclusion of exon 5, which alters mouse APOBEC3 translation efficiency. PLoS Pathog. 2012;8(1):e1002478.
Okeoma CM, Petersen J, Ross SR. Expression of murine APOBEC3 alleles in different mouse strains and their effect on mouse mammary tumor virus infection. J Virol. 2009;83(7):3029–38.
Langlois MA, Kemmerich K, Rada C, Neuberger MS. The AKV murine leukemia virus is restricted and hypermutated by mouse APOBEC3. J Virol. 2009;83(22):11550–9.
Kolokithas A, Rosenke K, Malik F, Hendrick D, Swanson L, Santiago ML, et al. The glycosylated Gag protein of a murine leukemia virus inhibits the antiretroviral function of APOBEC3. J Virol. 2010;84(20):10933–6.
Kozak CA, Chakraborti A. Single amino acid changes in the murine leukemia virus capsid protein gene define the target of Fv1 resistance. Virology. 1996;225(2):300–5.
Casey RE. Mouse strain-specific splicing of Apobec3. MSc Thesis, Worcester Polytechnic Institute. 2006: http://www.wpi.edu/Pubs/ETD/Available/etd-082206-113216/unrestricted/Casey.pdf.
Lassen KG, Wissing S, Lobritz MA, Santiago M, Greene WC. Identification of two APOBEC3F splice variants displaying HIV-1 antiviral activity and contrasting sensitivity to Vif. J Biol Chem. 2010;285(38):29326–35.
Biasin M, Piacentini L, Lo Caputo S, Kanari Y, Magri G, Trabattoni D et al. Apolipoprotein B mRNA-editing enzyme, catalytic polypeptide-like 3G: a possible role in the resistance to HIV of HIV-exposed seronegative individuals. J Inf Dis. 2007;195(7):960–4.
Sui Y, Zhu Q, Gagnon S, Dzutsev A, Terabe M, Vaccari M, et al. Innate and adaptive immune correlates of vaccine and adjuvant-induced control of mucosal transmission of SIV in macaques. PNAS. 2010;107(21):9843–8.
Santiago ML, Benitez RL, Montano M, Hasenkrug KJ, Greene WC. Innate retroviral restriction by Apobec3 promotes antibody affinity maturation in vivo. J Immunol. 2010;185(2):1114–23.
Conticello SG, Thomas CJ, Petersen-Mahrt SK, Neuberger MS. Evolution of the AID/APOBEC family of polynucleotide (deoxy)cytidine deaminases. Mol Biol Evol. 2005;22(2):367–77.
Muramatsu M, Kinoshita K, Fagarasan S, Yamada S, Shinkai Y, Honjo T. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell. 2000;102(5):553–63.
Okeoma CM, Lovsin N, Peterlin BM, Ross SR. APOBEC3 inhibits mouse mammary tumour virus replication in vivo. Nature. 2007;445(7130):927–30.
Low A, Okeoma CM, Lovsin N, de las Heras M, Taylor TH, Peterlin BM et al. Enhanced replication and pathogenesis of Moloney murine leukemia virus in mice defective in the murine APOBEC3 gene. Virology. 2009;385(2):455–63.
Dittmer U, Race B, Peterson KE, Stromnes IM, Messer RJ, Hasenkrug KJ. Essential roles for CD8+ T cells and gamma interferon in protection of mice against retrovirus-induced immunosuppression. J Virol. 2002;76(1):450–4.
Voltersvik P, Albrektsen G, Ulvestad E, Dyrhol-Riise AM, Sorensen B, Asjo B. Changes in immunoglobulin isotypes and immunoglobulin G (IgG) subclasses during highly active antiretroviral therapy: anti-p24 IgG1 closely parallels the biphasic decline in plasma viremia. J Acquir Immune Defic Syndr. 2003;34(4):358–67.
Morris L, Binley JM, Clas BA, Bonhoeffer S, Astill TP, Kost R, et al. HIV-1 antigen-specific and -nonspecific B cell responses are sensitive to combination antiretroviral therapy. J Exp Med. 1998;188(2):233–45.
Binley JM, Trkola A, Ketas T, Schiller D, Clas B, Little S, et al. The effect of highly active antiretroviral therapy on binding and neutralizing antibody responses to human immunodeficiency virus type 1 infection. J Inf Dis. 2000;182(3):945–9.
Mariani R, Chen D, Schrofelbauer B, Navarro F, Konig R, Bollman B, et al. Species-specific exclusion of APOBEC3G from HIV-1 virions by Vif. Cell. 2003;114(1):21–31.
Bachmann MF, Jennings GT. Vaccine delivery: a matter of size, geometry, kinetics and molecular patterns. Nat Rev Immunol. 2010;10(11):787–96.
Browne EP, Littman DR. Myd88 is required for an antibody response to retroviral infection. PLoS Pathog. 2009;5(2):e1000298.
Browne EP. Toll-like receptor 7 controls the anti-retroviral germinal center response. PLoS Pathog. 2011;7(10):e1002293.
Kane M, Case LK, Wang C, Yurkovetskiy L, Dikiy S, Golovkina TV. Innate immune sensing of retroviral infection via Toll-like receptor 7 occurs upon viral entry. Immunity. 2011;35(1):135–45.
Walker LM, Huber M, Doores KJ, Falkowska E, Pejchal R, Julien JP, et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature. 2011;477(7365):466–70.
Rerks-Ngarm S, Pitisuttithum P, Nitayaphan S, Kaewkungwal J, Chiu J, Paris R, et al. Vaccination with ALVAC and AIDSVAX to prevent HIV-1 infection in Thailand. NEJM. 2009;361(23):2209–20.
Ackerman ME, Dugast AS, Alter G. Emerging concepts on the role of innate immunity in the prevention and control of HIV infection. Ann Rev Med. 2012;63:113–30.
Barouch DH, Liu J, Li H, Maxfield LF, Abbink P, Lynch DM, et al. Vaccine protection against acquisition of neutralization-resistant SIV challenges in rhesus monkeys. Nature. 2012;482(7383):89–93.
Chesebro B, Wehrly K, Cloyd M, Britt W, Portis J, Collins J, et al. Characterization of mouse monoclonal antibodies specific for Friend murine leukemia virus-induced erythroleukemia cells: Friend-specific and FMR-specific antigens. Virology. 1981;112(1):131–44.
Britt WJ, Chesebro B. Use of monoclonal anti-gp70 antibodies to mimic the effects of the Rfv-3 gene in mice with Friend virus-induced leukemia. J Immunol. 1983;130(5):2363–7.
Hasenkrug KJ, Brooks DM, Chesebro B. Passive immunotherapy for retroviral disease: influence of major histocompatibility complex type and T-cell responsiveness. PNAS. 1995;92(23):10492–5.
Messer RJ, Dittmer U, Peterson KE, Hasenkrug KJ. Essential role for virus-neutralizing antibodies in sterilizing immunity against Friend retrovirus infection. PNAS USA. 2004;101(33):12260–5.
Sala A, Gresser I, Chassoux D, Maury C, Santodonato L, Eid P, et al. Inhibition of Friend leukemia cell visceral metastases by a new monoclonal antibody and role of the immune system of the host in its action. Cancer Res. 1992;52(10):2880–9.
Friend C. Immunological relationships of a filterable agent causing a leukemia in adult mice. I. The neutralization of infectivity by specific antiserum. J Exp Med. 1959;109(2):217–28.
Chesebro B, Wehrly K. Studies on the role of the host immune response in recovery from Friend virus leukemia. I. Antiviral and antileukemia cell antibodies. J Exp Med. 1976;143(1):73–84.
Sanders RW, Vesanen M, Schuelke N, Master A, Schiffner L, Kalyanaraman R, et al. Stabilization of the soluble, cleaved, trimeric form of the envelope glycoprotein complex of human immunodeficiency virus type 1. J Virol. 2002;76(17):8875–89.
Pantophlet R, Wang M, Aguilar-Sino RO, Burton DR. The human immunodeficiency virus type 1 envelope spike of primary viruses can suppress antibody access to variable regions. J Virol. 2009;83(4):1649–59.
Ishihara C, Miyazawa M, Nishio J, Chesebro B. Induction of protective immunity to Friend murine leukemia virus in genetic nonresponders to virus envelope protein. J Immunol. 1991;146(11):3958–63.
Dittmer U, Brooks DM, Hasenkrug KJ. Characterization of a live-attenuated retroviral vaccine demonstrates protection via immune mechanisms. J Virol. 1998;72(8):6554–8.
Delgado MF, Coviello S, Monsalvo AC, Melendi GA, Hernandez JZ, Batalle JP, et al. Lack of antibody affinity maturation due to poor Toll-like receptor stimulation leads to enhanced respiratory syncytial virus disease. Nat Med. 2009;15(1):34–41.
Earl PL, Moss B, Morrison RP, Wehrly K, Nishio J, Chesebro B. T-lymphocyte priming and protection against Friend leukemia by vaccinia-retrovirus env gene recombinant. Science. 1986;234(4777):728–31.
Ishihara C, Miyazawa M, Nishio J, Azuma I, Chesebro B. Use of low toxicity adjuvants and killed virus to induce protective immunity against the Friend murine leukaemia retrovirus-induced disease. Vaccine. 1992;10(5):353–6.
Schlecht-Louf G, Renard M, Mangeney M, Letzelter C, Richaud A, Ducos B, et al. Retroviral infection in vivo requires an immune escape virulence factor encrypted in the envelope protein of oncoretroviruses. PNAS USA. 2010;107(8):3782–7.
Hunsmann G, Moennig V, Schafer W. Properties of mouse leukemia viruses. IX. Active and passive immunization of mice against Friend leukemia with isolated viral GP71 glycoprotein and its corresponding antiserum. Virology. 1975;66(1):327–9.
Hunsmann G, Schneider J, Schulz A. Immunoprevention of Friend virus-induced erythroleukemia by vaccination with viral envelope glycoprotein complexes. Virology. 1981;113(2):602–12.
Morrison RP, Earl PL, Nishio J, Lodmell DL, Moss B, Chesebro B. Different H-2 subregions influence immunization against retrovirus and immunosuppression. Nature. 1987;329(6141):729–32.
Miyazawa M, Nishio J, Chesebro B. Genetic control of T cell responsiveness to the Friend murine leukemia virus envelope antigen. Identification of class II loci of the H-2 as immune response genes. J Exp Med. 1988;168(5):1587–605.
Hasenkrug KJ, Brooks DM, Nishio J, Chesebro B. Differing T-cell requirements for recombinant retrovirus vaccines. J Virol. 1996;70(1):368–72.
Miyazawa M, Nishio J, Chesebro B. Protection against Friend retrovirus-induced leukemia by recombinant vaccinia viruses expressing the gag gene. J Virol. 1992;66(7):4497–507.
McElrath MJ, Haynes BF. Induction of immunity to human immunodeficiency virus type-1 by vaccination. Immunity. 2010;33(4):542–54.
Bayer W, Schimmer S, Hoffmann D, Dittmer U, Wildner O. Evaluation of the Friend Virus model for the development of improved adenovirus-vectored anti-retroviral vaccination strategies. Vaccine. 2008;26(5):716–26.
Casimiro DR, Chen L, Fu TM, Evans RK, Caulfield MJ, Davies ME, et al. Comparative immunogenicity in rhesus monkeys of DNA plasmid, recombinant vaccinia virus, and replication-defective adenovirus vectors expressing a human immunodeficiency virus type 1 gag gene. J Virol. 2003;77(11):6305–13.
Roberts DM, Nanda A, Havenga MJ, Abbink P, Lynch DM, Ewald BA, et al. Hexon-chimaeric adenovirus serotype 5 vectors circumvent pre-existing anti-vector immunity. Nature. 2006;441(7090):239–43.
Bayer W, Lietz R, Ontikatze T, Johrden L, Tenbusch M, Nabi G, et al. Improved vaccine protection against retrovirus infection after co-administration of adenoviral vectors encoding viral antigens and type I interferon subtypes. Retrovirology. 2011;8:75.
Best S, Le Tissier P, Towers G, Stoye JP. Positional cloning of the mouse retrovirus restriction gene Fv1. Nature. 1996;382(6594):826–9.
Daniel MD, Kirchhoff F, Czajak SC, Sehgal PK, Desrosiers RC. Protective effects of a live attenuated SIV vaccine with a deletion in the nef gene. Science. 1992;258(5090):1938–41.
Schmitt K, Hill MS, Ruiz A, Culley N, Pinson DM, Wong SW, et al. Mutations in the highly conserved SLQYLA motif of Vif in a simian-human immunodeficiency virus result in a less pathogenic virus and are associated with G-to-A mutations in the viral genome. Virology. 2009;383(2):362–72.
Tong T, Crooks ET, Osawa K, Binley JM. HIV-1 virus-like particles bearing pure Env trimers expose neutralizing epitopes but occlude nonneutralizing epitopes. J Virol. 2012;86(7):3574–87.
Acknowledgments
This work was supported by the National Institutes of Health R01 AI090795 (MLS), University of Colorado (UCD) Department of Medicine Start-up Funds (MLS), the Tim Gill Foundation (MSH) and the UCD Outstanding Early Career Scholar Program (MLS). KH is a Robert D. Watkins fellow of the American Society for Microbiology.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Halemano, K., Harper, M.S., Guo, K. et al. Humoral immunity in the Friend retrovirus infection model. Immunol Res 55, 249–260 (2013). https://doi.org/10.1007/s12026-012-8370-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-012-8370-y