Abstract
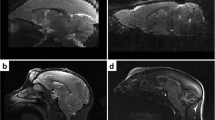
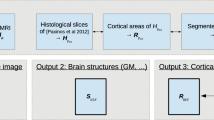
Currently available non-human primate templates typically require input of a skull-stripped brain for structural processing. This can be a manually intensive procedure, and considerably limits their utility. The purpose of this study was to create a vervet MRI population template, associated tissue probability maps (TPM), and a label atlas to facilitate true fully automated Magnetic Resonance Imaging (MRI) structural analyses for morphometric analyses. Structural MRI scans of ten vervet monkeys (Chlorocebus aethiops) scanned at three time points were used in this study. An unbiased population average template was created using a symmetric diffeomorphic registration (SyN) procedure. Skull stripping, segmentation, and label map generation were performed using the publically available rhesus INIA19 MRI template and NeuroMap label atlas. A six-class TPM and a six-layer two-class normalization template was created from the vervet segmentation for use within the Statistical Parametric Mapping (SPM) framework. Fully automated morphologic processing of all of the vervet MRI scans was then performed using the vervet TPM and vervet normalization template including skull-stripping, segmentation and normalization. The vervet template creation procedure resulted in excellent skull stripping, segmentation, and NeuroMap atlas labeling with 720 structures successfully registered. Fully automated processing was accomplished for all vervet scans, demonstrating excellent skull-stripping, segmentation, and normalization performance. We describe creation of an unbiased vervet structural MRI population template and atlas. The template includes an associated six-class TPM and DARTEL six-layer two-class normalization template for true fully automated skull-stripping, segmentation, and normalization of vervet structural T1-weighted MRI scans. We provide the most detailed vervet label atlas currently available based on the NeuroMaps atlas with 720 labels successfully registered. We additionally describe a novel method for atlas label generation that capitalizes on previous work in this area using high-dimensional highly accurate image matching procedures for inter-species morphologic normalization.





Similar content being viewed by others
References
Ashburner, J., & Friston, K. J. (2000). Voxel-based morphometry–the methods. NeuroImage, 11(6 Pt 1), 805–821.
Avants, B., & Gee, J. C. (2004). Geodesic estimation for large deformation anatomical shape averaging and interpolation. NeuroImage, 23(Suppl 1), S139–150.
Avants, B. B., Epstein, C. L., Grossman, M., & Gee, J. C. (2008). Symmetric diffeomorphic image registration with cross-correlation: evaluating automated labeling of elderly and neurodegenerative brain. Medical Image Analysis, 12(1), 26–41.
Bowden, D. M., & Dubach, M. F. (2000). Applicability of the template atlas to various primate species. In D. M. Bowden & R. F. Martin (Eds.), Primate brain maps: structure of the macaque brain (pp. 38–47). Amsterdam: Elsevier Science.
Deogaonkar, M., Heers, M., Mahajan, S., Brummer, M., & Subramanian, T. (2005). Method of construction of a MRI-based tabular database of 3D stereotaxic co-ordinates for individual structures in the basal ganglia of Macaca mulatta. Journal of Neuroscience Methods, 149(2), 154–163. doi:10.1016/j.jneumeth.2005.05.016.
Fedorov, A., Li, X., Pohl, K. M., Bouix, S., Styner, M., Addicott, M., et al. (2011). Atlas-guided segmentation of vervet monkey brain MRI. Open NeuroImage Journal, 5, 186–197. doi:10.2174/1874440001105010186.
Frey, S., Pandya, D. N., Chakravarty, M. M., Bailey, L., Petrides, M., & Collins, D. L. (2011). An MRI based average macaque monkey stereotaxic atlas and space (MNI monkey space). NeuroImage, 55(4), 1435–1442. doi:10.1016/j.neuroimage.2011.01.040.
Jenkinson, M., Beckmann, C. F., Behrens, T. E., Woolrich, M. W., & Smith, S. M. (2012). Fsl. [Historical article review]. NeuroImage, 62(2), 782–790. doi:10.1016/j.neuroimage.2011.09.015.
Klein, A., Andersson, J., Ardekani, B. A., Ashburner, J., Avants, B., Chiang, M. C., et al. (2009). Evaluation of 14 nonlinear deformation algorithms applied to human brain MRI registration. NeuroImage, 46(3), 786–802.
Maldjian, J. A., Laurienti, P. J., Kraft, R. A., & Burdette, J. H. (2003). An automated method for neuroanatomic and cytoarchitectonic atlas-based interrogation of fMRI data sets. NeuroImage, 19(3), 1233–1239.
McLaren, D. G., Kosmatka, K. J., Oakes, T. R., Kroenke, C. D., Kohama, S. G., Matochik, J. A., et al. (2009). A population-average MRI-based atlas collection of the rhesus macaque. [Research support, N.I.H., Extramural research support, N.I.H., intramural]. NeuroImage, 45(1), 52–59. doi:10.1016/j.neuroimage.2008.10.058.
Rohlfing, T., Kroenke, C. D., Sullivan, E. V., Dubach, M. F., Bowden, D. M., Grant, K. A., et al. (2012). The INIA19 template and neuromaps atlas for primate brain image parcellation and spatial normalization. Front Neuroinformation, 6, 27. doi:10.3389/fninf.2012.00027.
Steiper, M. E., & Young, N. M. (2006). Primate molecular divergence dates. [Research support, N.I.H., Extramural research support, non-U.S. Gov’t]. Molecular Phylogenetics and Evolution, 41(2), 384–394. doi:10.1016/j.ympev.2006.05.021.
Sultan, F., Hamodeh, S., Murayama, Y., Saleem, K. S., & Logothetis, N. (2010). Flat map areal topography in Macaca mulatta based on combined MRI and histology. Magnetic Resonance Imaging, 28(8), 1159–1164. doi:10.1016/j.mri.2010.03.023.
Wisco, J. J., Rosene, D. L., Killiany, R. J., Moss, M. B., Warfield, S. K., Egorova, S., et al. (2008). A rhesus monkey reference label atlas for template driven segmentation. [Research Support, N.I.H., Extramural Research Support, Non-U.S. Gov’t Validation Studies]. Journal of Medical Primatology, 37(5), 250–260. doi:10.1111/j.1600-0684.2008.00288.x.
Woods, R. P., Fears, S. C., Jorgensen, M. J., Fairbanks, L. A., Toga, A. W., & Freimer, N. B. (2011). A web-based brain atlas of the vervet monkey, Chlorocebus aethiops. [Research support, N.I.H., Extramural research support, Non-U.S. Gov’t research support, U.S. Gov’t, Non-P.H.S.]. NeuroImage, 54(3), 1872–1880. doi:10.1016/j.neuroimage.2010.09.070.
Acknowledgments
This study was supported in part by National Institute on Alcohol Abuse and Alcoholism: AA019431, AA017710 and AA016748 (JBD), and AA014106 (PI: David Friedman). Support for this research was also provided by NIH grants R019963/OD010965 (PI: Jay R. Kaplan), NS058700 (PI: Don Bowden) and NS0075107 (JAM). The authors would also like to thank Ben Wagner for programming assistance and the Center for Biomolecular Imaging along with Bob Kraft for vervet MRI scanning.
Conflict of Interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maldjian, J.A., Daunais, J.B., Friedman, D.P. et al. Vervet MRI Atlas and Label Map for Fully Automated Morphometric Analyses. Neuroinform 12, 543–550 (2014). https://doi.org/10.1007/s12021-014-9231-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12021-014-9231-8