Abstract
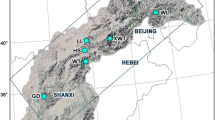
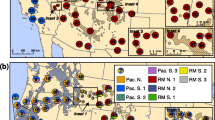
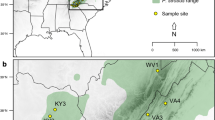
Shortleaf pine (n = 93) and loblolly pine (n = 112) trees representing 22 seed sources or 16 physiographic populations were sampled from Southwide Southern Pine Seed Source Study plantings located in Oklahoma, Arkansas, and Mississippi. The sampled trees were grown from shortleaf pine and loblolly pine seeds formed in 1951 and 1952, prior to the start of intensive forest management across their native ranges. Amplification fragment length polymorphism (AFLP) markers were developed and used to study genetic diversity and its structure in these pine species. After screening 48 primer pairs, 17 and 21 pairs were selected that produced 794 and 647 AFLPs in shortleaf pine and loblolly pine, respectively. High-AFLP-based genetic diversity exists within shortleaf pine and loblolly pine, and most (84.73% in shortleaf pine; 87.69% in loblolly pine) of this diversity is maintained within physiographic populations. The high value of unbiased measures of genetic identity and low value of genetic distance for all pairwise comparisons indicates that the populations have similar genetic structures. For shortleaf pine, there was no significant correlation between geographic distance and genetic distance (r = 0.28), while for loblolly pine there was a weak but significant correlation (r = 0.51).




Similar content being viewed by others
References
Al-Rabab’ah MA, Williams CG (2002) Population dynamics of Pinus taeda L. based on nuclear microsatellites. For Ecol Manag 163:263–271
Chen JW, Tauer CG, Bai G, Huang Y, Payton ME, Holley AG (2004) Bidirectional introgression between Pinus taeda and Pinus echinata: evidence from morphological and molecular data. Can J For Res 34:2508–2516
Doyle JJ, Doyle J (1988) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Edwards MA, Hamrick JL (1995) Genetic variation in shortleaf pine, Pinus echinata Mill. (Pinaceae). For Genet 2:21–28
Florence Z, Rink G (1979) Geographic patterns of allozymic variation in loblolly pine. In: Proceedings of the 15th Southern Forest Tree Improvement Conference, Starkville, MS, 19–21 June 1979. Mississippi State University, MS, pp 33–41
Hartl DL, Clark AG (1989) Principles of population genetics, 2nd edn. Sinauer, Sunderland
Heun M, Murphy JP, Phillips TD (1994) A comparison of RAPD and isozyme analyses for determining the genetic relationships among Avena sterilis L. accessions. Theor Appl Genet 87:689–696
Huneycutt M, Askew GR (1989) Electrophoretic identification of loblolly pine shortleaf pine hybrids. Silvae Genet 38:95–96
Lanner-Herrera C, Gustafsson M, Fält AS, Bryngelsson T (1996) Diversity in natural populations of wild Brassica oleracea as estimated by isozyme and RAPD analysis. Genet Resour Crop Evol 43:13–23
Manly BFJ (1985) The statistics of natural selection. Chapman and Hall, London, pp 272–282
Messmer MM, Melchinger AE, Woodman WL, Lee EA, Lamkey KR (1991) Genetic diversity among progenitors and elite lines from the Iowa Stiff Stalk Synthetic (BSSS) maize population: comparison of allozyme and RFLP data. Theor Appl Genet 83:97–107
Moser H, Lee M (1994) RFLP variation and genealogical distance, multivariate distance, heterosis, and genetic variation in oats. Theor Appl Genet 87:947–956
Muluvi GM, Sprent JI, Soranzo N, Provan J, Odee D, Folkard G, McNicol JW, Powell W (1999) Amplified fragment length polymorphism (AFLP) analysis of genetic variation in Moringa oleifera Lam. Mol Ecol 8:463–470
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York, pp 187–192
Raja RG, Tauer CG, Wittwer RF, Huang YH (1997) Isoenzyme variation and genetic structure in natural populations of shortleaf pine (Pinus echinata). Can J For Res 27:740–749
Raja RG, Tauer CG, Wittwer RF, Huang Y (1998) Regeneration methods affect genetic variation and structure in shortleaf pine (Pinus echinata Mill.). For Genet 5:171–178
Remington DL, O’Malley DM (2000) Whole-genome characterization of embryonic stage inbreeding depression in a selfed loblolly pine family. Genetics 155:337–348
Remington DL, Whetten RW, Liu BH, O’Malley DM (1999) Construction of an AFLP genetic map with nearly complete genome coverage in Pinus taeda. Theor Appl Genet 98:1279–1292
Russell JR, Fuller JD, Macaulay M, Hatz BG, Jahoor A, Powell W, Waugh R (1997) Direct comparison of levels of genetic variation among barley accessions detected by RFLPs, AFLPs, SSRs and RAPDs. Theor Appl Genet 95:714–722
Schmidtling RC, Carroll E, LaFarge T (1999) Allozyme diversity of selected and natural loblolly pine populations. Silvae Genet 48:35–45
Schultz RP (1997) Loblolly pine: the ecology and culture of loblolly pine (Pinus taeda L). US Dept. of Agric., Ag. Handb. 713
Smith OS, Smith JSC, Bowen SL, Tenborg RA (1992) Numbers of RFLP probes necessary to show associations between lines. Maize Genet Coop News Lett 66:66
South DB, Buckner ER (2003) The decline of southern yellow pine timberland. J For 101:30–35
Sun GL, Díaz O, Salomon B, Bothmer R (1999) Genetic diversity in Elymus caninus as revealed by isozyme, RAPD, and microsatellite markers. Genome 42:420–431
Tauer CG (1980) Twenty-year results of a shortleaf pine seed source study in Oklahoma. OSU Ag Exp Sta Bull B-752, pp 12
Wells OO (1973) Variation among shortleaf pines in a Mississippi seed source planting. USDA For Serv Res Note 3-SO-162, pp 8
Wells OO, Lambeth CC (1983) Loblolly pine province test in southern Arkansas 25th year results. South J Appl For 2:71–75
Wells OO, Wakley PC (1970) Variation in shortleaf pine from several geographic sources. For Sci 16:415–423
Wells OO, Switzer GL, Schmidtling RC (1991) Geographic variation in Mississippi loblolly pine and sweet gum. Silvae Genet 40:105–119
Xu S, Tauer CG, Nelson CD (2008) Natural hybridization within seed sources of shortleaf pine (Pinus echinata Mill.) and loblolly pine (Pinus taeda L.). Tree Genetics and Genomes (in press)
Yeh FC, Boyle TJB (1997) Population genetic analysis of co-dominant and dominant markers and quantitative traits. Belg J Bot 129:157
Acknowledgements
This study is supported by the USDA Forest Service, Southern Research Station, Cooperative Agreement SRS 05-CA-11330126-168 and by the Oklahoma State University Agricultural Experiment Station. We thank Larry Lott (Southern Institute of Forest Genetics) and personnel of the Oklahoma State University Kiamichi Forestry Research Station for assistance in locating and collecting needle samples.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Sederoff
Rights and permissions
About this article
Cite this article
Xu, S., Tauer, C.G. & Nelson, C.D. Genetic diversity within and among populations of shortleaf pine (Pinus echinata Mill.) and loblolly pine (Pinus taeda L.). Tree Genetics & Genomes 4, 859–868 (2008). https://doi.org/10.1007/s11295-008-0158-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-008-0158-9