Abstract
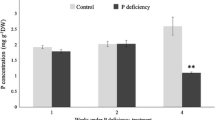
Phosphorus (P) is often a limiting macronutrient because of its low availability in soils. White lupin (Lupinus albus L.) plants are well adapted to growth under low-P conditions. White lupin acclimation to low-P conditions includes changes in root architecture and enhanced expression of numerous genes encoding for secreted acid phosphatases and phosphate transporters. However, information about transcription factors and signaling proteins that coordinate the P-starvation responses is limited in white lupin. In this study, cDNAs and ESTs encoding for transcription factors and signaling proteins were isolated and their transcription profiles were clarified to facilitate the identification of key signal transduction genes necessary to improve P acquisition, allocation, and use. 34 cDNA fragments of MYB-coiled coil (MYB-CC) and R2R3-MYB, and 26 ESTs encoding for transcription factors and signaling proteins were isolated. Four MYB-CC cDNAs showed high similarity to the transcription factor Phosphate starvation response 1 in Arabidopsis, which has been implicated in regulation of many P-starvation response genes. In addition, deduced amino acid sequences of 29 R2R3-MYB cDNAs showed similarities to Arabidopsis R2R3-MYB proteins. Transcription of the 60 genes, as measured by real-time reverse transcription-PCR, in normal roots, cluster roots, leaves, and shoot tips under P sufficient and low-P conditions revealed that six (10%) and two (3.3%) sequences were either induced or suppressed, respectively, by low-P condition. In addition, 36 genes (60%) showed an organ specific expression.




Similar content being viewed by others
Abbreviations
- 3′RACE:
-
Rapid amplification of cDNA 3′-end
- CC:
-
coiled coil
- DAE:
-
days after emergence
- EST:
-
expressed sequence tag
- LaMATE:
-
Lupinus albus multidrug and toxin efflux protein
- LaPT1:
-
Lupinus albus phosphate transporter 1
- LaSAP:
-
Lupinus albus secreted acid phosphatase
- P:
-
Phosphorus
- PHR:
-
phosphate starvation response
- RT-PCR:
-
reverse transcription-PCR
- TF:
-
transcription factor
References
Agarwal M, Hao Y, Kapoor A, Dong C-H, Fujii H, Zheng X, Zhu J-K (2006) A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J Biol Chem 281:37636–37645
Bogs J, Jaffe FW, Takos AM, Walker AR, Robinson SP (2007) The grapevine transcription factor VvMYBPA1 regulates proanthocyanidin synthesis during fruit development. Plant Physiol 143:1347–1361
Cesco S, Neumann G, Tomasi N, Pinton R, Weisskopf L (2010) Release of plant-borne flavonoids into the rhizosphere and their role in plant nutrition. Plant Soil 329:1–25
Chen YF, Li LQ, Xu Q, Kong YH, Wang H, Wu WH (2009) The WRKY6 transcription factor modulates PHOSPHATE1 expression in response to low Pi stress in Arabidopsis. Plant Cell 21:3554–3566
Cominelli E, Galbiati M, Vavasseur A, Conti L, Sala T, Vuylsteke M, Leonhardt N, Dellaporta SL, Tonelli C (2005) A guard-cell-specific MYB transcription factor regulates stomatal movements and plant drought tolerance. Curr Biol 15:1196–1200
Czechowski T, Bari RP, Stitt M, Scheible WR, Udvardi MK (2004) Real-time RT-PCR profiling of over 1400 Arabidopsis transcription factors: unprecedented sensitivity reveals novel root- and shoot-specific genes. Plant J 38:366–379
Devaiah BN, Karthikeyan AS, Raghothama KG (2007) WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol 143:1789–1801
Devaiah BN, Madhuvanthi R, Karthikeyan AS, Raghothama KG (2009) Phosphate starvation responses and gibberellic acid biosynthesis are regulated by the MYB62 transcription factor in Arabidopsis. Mol Plant 2:43–58
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trend Plant Sci 5:199–206
Hernández G, Ramírez M, Valdés-López O, Tesfaye M, Graham MA, Czechowski T, Schlereth A, Wandrey M, Erban A, Cheung F, Wu HC, Lara M, Town CD, Kopka J, Udvardi MK, Vance CP (2007) Phosphorus stress in common bean: root transcript and metabolic responses. Plant Physiol 144:752–767
Johnson JF, Allan DL, Vance CP (1994) Phosphorus stress-induced proteoid roots show altered metabolism in Lupinus albus. Plant Physiol 104:657–665
Johnson JF, Allan DL, Vance CP (1996a) Phosphorus deficiency in Lupinus albus. Altered lateral root development and enhanced expression of phosphoenolpyruvate carboxylase. Plant Physiol 112:31–41
Johnson JF, Allan DL, Vance CP, Weiblen G (1996b) Root carbon dioxide fixation by phosphorus-deficient Lupinus albus. Contribution to organic acid exudation by proteoid roots. Plant Physiol 112:19–30
Keerthisinghe G, Hocking PJ, Ryan PR, Delhaize E (1998) Effect of phosphorus supply on the formation and function of proteoid roots of white lupin (Lupinus albus L.). Plant Cell Environ 21:467–478
Kim MC, Chung WS, Yun D-J, Cho MJ (2009) Calcium and calmodulin-mediated regulation of gene expression in plants. Mol Plant 2:13–21
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kirik V, Kölle K, Wohlfarth T, Miséra S, Bäumlein H (1998) Ectopic expression of a novel MYB gene modifies the architecture of the Arabidopsis inflorescence. Plant J 13:729–742
Kranz HD, Denekamp M, Greco R, Jin H, Leyva A, Meissner RC, Petroni K, Urzainqui A, Bevan M, Martin C, Smeekens S, Tonelli C, Paz-Ares J, Weisshaar B (1998) Towards functional characterization of the members of the R2R3-MYB gene family from Arabidopsis thaliana. Plant J 16:263–276
Lin W-Y, Lin S-I, Chiou T-J (2009) Molecular regulators of phosphate homeostasis in plants. J Exp Bot 60:1427–1438
Liu J, Uhde-Stone C, Li A, Vance C, Allan D (2001) A phosphate transporter with enhanced expression in proteoid roots of white lupin (Lupinus albus L.). Plant Soil 237:257–266
Liu J, Samac DA, Bucciarelli B, Allan DL, Vance CP (2005) Signaling of phosphorus deficiency-induced gene expression in white lupin requires sugar and phloem transport. Plant J 41:257–268
Miller SS, Liu J, Allan DL, Menzhuber CJ, Fedorova M, Vance CP (2001) Molecular control of acid phosphatase secretion into the rhizosphere of proteoid roots from phosphorus-stressed white lupin. Plant Physiol 127:594–606
Morcuende R, Bari R, Gibon Y, Zheng W, Pant BD, Bläsing O, Usadel B, Czechowski T, Udvardi MK, Sttitt M, Scheible W-R (2007) Genome-wide reprogramming of metabolism and regulatory networks of Arabidopsis in response to phosphorus. Plant Cell Environ 30:85–112
Müller R, Morant M, Jarmer H, Nilsson L, Nielsen TH (2007) Genome-wide analysis of the Arabidopsis leaf transcriptome reveals interaction of phosphate and sugar metabolism. Plant Physiol 143:156–171
Nesi N, Jond C, Debeaujon I, Caboche M, Lepiniec L (2001) The Arabidopsis TT2 gene encodes an R2R3 MYB domain protein that acts as a key determinant for proanthocyanidin accumulation in developing seed. Plant Cell 13:2099–2114
Neumann G, Massonneau A, Martinoia E, Römheld V (1999) Physiological adaptations to phosphorus deficiency during proteoid root development in white lupin. Planta 208:373–382
Nilsson L, Müller R, Nielsen TH (2007) Increased expression of the MYB-related transcription factor, PHR1, leads to enhanced phosphate uptake in Arabidopsis thaliana. Plant Cell Environ 30:1499–1512
Peñaloza E, Muñoz G, Salvo-Garrido H, Silva H, Corcuera LJ (2005) Phosphate deficiency regulates phosphoenolpyruvate carboxylase expression in proteoid root clusters of white lupin. J Exp Bot 56:145–153
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Rabinowicz PD, Braun EL, Wolfe AD, Bowen B, Grotewold E (1999) Maize R2R3 Myb genes: sequence analysis reveals amplification in the higher plants. Genetics 153:427–444
Ramakers C, Ruijter JM, Lekanne Deprez RH, Moorman FM (2003) Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339:62–66
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C-Z, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G-L (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110
Rubio V, Linhares F, Solano R, Martin AC, Iglesias J, Leyva A, Paz-Ares J (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15:2122–2133
Saitou N, Nei N (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sbabou L, Bucciarelli B, Miller S, Liu J, Berhada F, Filali-Maltouf A, Allan D, Vance C (2010) Molecular analysis of SCARECROW genes expressed in white lupin cluster roots. J Exp Bot 61:1351–1363
Shane MW, De Vos M, De Roock S, Lambers H (2003) Shoot P status regulates cluster-root growth and citrate exudation in Lupinus albus grown with a divided root system. Plant Cell Environ 26:265–273
Shen J, Li H, Neumann G, Zhang F (2005) Nutrient uptake, cluster root formation and exudation of protons and citrates in Lupinus albus as affected by localized supply of phosphorus in a split-root system. Plant Sci 168:837–845
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4:447–456
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B (2007) Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J 50:660–677
Tesfaye M, Liu J, Allan DL, Vance CP (2007) Genomic and genetic control of phosphate stress in legumes. Plant Physiol 144:594–603
Uhde-Stone C, Gilbert G, Johnson JMF, Litjens R, Zinn KE, Temple SJ, Vance CP, Allan DL (2003a) Acclimation of white lupin to phosphorus deficiency involves enhanced expression of genes related to organic acid metabolism. Plant Soil 248:99–116
Uhde-Stone C, Zinn KE, Ramírez M, Li A, Vance CP, Allan DL (2003b) Nylon filter arrays reveal differential gene expression in proteoid roots of white lupin in response to phosphorus deficiency. Plant Physiol 131:1064–1079
Uhde-Stone C, Liu J, Zinn KE, Allan DL, Vance CP (2005) Transgenic proteoid roots of white lupin: a vehicle for characterizing and silencing root genes involved in adaptation to P stress. Plant J 44:840–853
Vailleau F, Daniel X, Tronchet M, Montillet JL, Triantaphylidès C, Roby D (2002) A R2R3-MYB gene, AtMYB30, acts as a positive regulator of the hypersensitive cell death program in plants in response to pathogen attack. Proc Natl Acad Sci USA 99:10179–10184
Vance CP (2010) Quantitative trait loci, epigenetics, sugars, and microRNAs: quaternaries in phosphate acquisition and use. Plant Physiol 154:582–588
Vance CP, Uhde-Stone C, Allan DL (2003) Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytol 157:423–447
Wasaki J, Omura M, Osaki M, Ito H, Matsui H, Shinano T, Tadano T (1999) Structure of a cDNA for an acid phosphatase from phosphate-deficient lupin (Lupinus albus L.) roots. Soil Sci Plant Nutr 45:439–449
Watt M, Evans JR (1999a) Linking development and determinacy with organic acid efflux from proteoid roots of white lupin grown with low phosphorus and ambient or elevated atmospheric CO2 concentration. Plant Physiol 120:705–716
Watt M, Evans JR (1999b) Proteoid roots. Physiology and development. Plant Physiol 121:317–323
Weisskopf L, Abou-Mansour E, Fromin N, Tomasi N, Santelia D, Edelkott I, Neumann G, Aragno M, Tabacchi R, Martinoia E (2006a) White lupin has developed a complex strategy to limit microbial degradation of secreted citrate required for phosphate acquisition. Plant Cell Environ 29:919–927
Weisskopf L, Tomasi N, Santelia D, Martinoia E, Langlade NB, Tabacchi R, Abou-Mansour E (2006b) Isoflavonoid exudation from white lupin roots is influenced by phosphate supply, root type to cluster-root stage. New Phytol 171:657–668
Yamagishi M, Shimoyamada Y, Nakatsuka T, Masuda K (2010) Two R2R3-MYB genes, homologues of petunia AN2, regulate anthocyanin biosyntheses in flower tepals, tepal spots and leaves of Asiatic hybrid lily. Plant Cell Physiol 51:463–474
Yan F, Zhu Y, Muller C, Zorb C, Schubert S (2002) Adaptation of H+-pumping and plasma membrane H+ ATPase activity in proteoid roots of white lupin under phosphate deficiency. Plant Physiol 129:50–63
Yang XJ, Finnegan PM (2010) Regulation of phosphate starvation responses in higher plants. Ann Bot 105:513–526
Yang Y, Klessig DF (1996) Isolation and characterization of a tobacco mosaic virus-inducible myb oncogene homolog from tobacco. Proc Natl Acad Sci USA 93:14972–14977
Yanhui C, Yang X, He K, Liu M, Li J, Gao Z, Lin Z, Zhang Y, Wang X, Qiu X, Shen Y, Zhang L, Deng X, Luo J, Deng XW, Chen Z, Gu QuLJ (2006) The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60:107–124
Yoo JH, Park CY, Kim JC, Heo WD, Cheong MS, Park HC, Kim MC, Moon BC, Choi MS, Kang YH, Lee JH, Kim HS, Lee SM, Yoon HW, Lim CO, Yun DJ, Lee SY, Chung WS, Cho MJ (2005) Direct interaction of a divergent CaM isoform and the transcription factor, MYB2, enhance salt tolerance in Arabidopsis. J Biol Chem 280:3697–3706
Zhou J, Jiao F, Wu Z, Li Y, Wang X, He X, Zhong W, Wu P (2008a) OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol 146:1673–1686
Zhou K, Yamagishi M, Osaki M, Masuda K (2008b) Sugar signaling mediates cluster root formation and phosphorus starvation-induced gene expression in white lupin. J Exp Bot 59:2749–2756
Zimmermann IM, Heim MA, Weisshaar B, Uhrig JF (2004) Comprehensive identification of Arabidopsis thaliana MYB transcription factors interacting with R/B-like BHLH proteins. Plant J 40:22–34
Zinn KE, Liu J, Allan DL, Vance CP (2009) White lupin (Lupinus albus) response to phosphorus stress: evidence for complex regulation of LaSAP1. Plant Soil 322:1–15
Financial source
This research was supported by a Grants-In-Aid for Scientific Research (No. 17658032), the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Hans Lambers.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
The primary lateral root of white lupin at 6 DAE. A bleached root was stained by methylene blue The primary lateral root including rootlet primordia (very young proteoid root) was used for cDNA library construction. (GIF 149 kb)
Supplementary Fig. 2
(XLS 71 kb)
Supplementary Fig. 3-1
(GIF 258 kb)
Supplementary Fig. 3-2
(GIF 255 kb)
Supplementary Fig. 3-3
(GIF 279 kb)
Supplementary Fig. 3-4
Quantification of transcript abundance of genes encoding for TFs and signaling proteins in normal roots (NR), cluster roots (CR), leaves (Leaf), and shoot tips including shoot apical meristems (ST) of white lupin plants grown under P sufficient (white bars) and low-P (dark gray bars) growth conditions. Vertical bars show standard errors of the biological replicates. Within each figure, different letters above the bars represent significant differences at the 5% level. LaUbiquitin was used to normalize the expression. (GIF 230 kb)
Supplementary Table 1
List of cDNAs and ESTs analyzed in this study and primer sequences (XLS 35 kb)
Rights and permissions
About this article
Cite this article
Yamagishi, M., Zhou, K., Osaki, M. et al. Real-time RT-PCR profiling of transcription factors including 34 MYBs and signaling components in white lupin reveals their P status dependent and organ-specific expression. Plant Soil 342, 481–493 (2011). https://doi.org/10.1007/s11104-010-0711-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-010-0711-9