Abstract
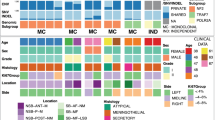
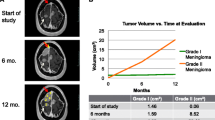
Although meningioma is the most common primary tumor of the central nervous system, the mechanism of progression from benign to atypical or anaplastic grade remains elusive. The present case reports the genomic evaluation of two synchronous meningiomas with different histological grades (benign and atypical) in the same patient. Under the assumption that the atypical tumor may have progressed from the benign tumor, the clonal origin of the lesions was investigated to identify genomic events responsible for the oncogenic process of evolution to higher grades in meningioma. A 59 year-old female patient was diagnosed with two synchronous meningiomas with different histological grades, benign and atypical. Whole-exome sequencing (WES) and RNA sequencing (RNA-seq) analysis of both tumors were done. WES analysis showed that each meningioma harbored distinct mutation profiles, and RNA-seq analysis revealed distinct gene expression profiles between the two tumors. The only apparent common genetic abnormality found in both tumors was the loss of heterozygosity of chromosome 22, raising the possibility that this event is the initial step in tumor formation, after which distinct subsequent mutations lead to the evolvement of two separate tumors of different grades. The result provides additional evidence on previous reports suggesting separate, independent mechanism of progression into higher grades in meningioma.



Similar content being viewed by others
References
Dolecek TA, Propp JM, Stroup NE, Kruchko C (2012) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2005–2009. Neuro-Oncol 14(Suppl 5):v1–v49. https://doi.org/10.1093/neuonc/nos218
Louis DN, Perry A, Reifenberger G, Deimling AV, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Zang KD (2001) Meningioma: a cytogenetic model of a complex benign human tumor, including data on 394 karyotyped cases. Cytogenet Cell Genet 93(3–4):207–220
Brastianos PK, Horowitz PM, Santagata S, Jones RT, McKenna A, Getz G, Ligon KL, Palescandolo E, Van Hummelen P, Ducar MD, Raza A, Sunkavalli A, MacConaill LE, Stemmer-Rachamimov AO, Louis DN, Hahn WC, Dunn IF, Beroukhim R (2013) Genomic sequencing of meningiomas identifies oncogenic SMO and AKT1 mutations. Nat Genet 45(3):285–289. https://doi.org/10.1038/ng.2526
Clark VE, Erson-Omay EZ, Serin A, Yin J, Cotney J, Ozduman K, Avşar T, Li J, Murray PB, Henegariu O, Yilmaz S, Günel JM, Carrión-Grant G, Yilmaz B, Grady C, Tanrikulu B, Bakircioğlu M, Kaymakçalan H, Caglayan AO, Sencar L, Ceyhun E, Atik AF, Bayri Y, Bai H, Kolb LE, Hebert RM, Omay SB, Mishra-Gorur K, Choi M, Overton JD, Holland EC, Mane S, State MW, Bilgüvar K, Baehring JM, Gutin PH, Piepmeier JM, Vortmeyer A, Brennan CW, Pamir MN, Kiliç T, Lifton RP, Noonan JP, Yasuno K, Günel M (2013) Genomic analysis of non-NF2 meningiomas reveals mutations in TRAF7, KLF4, AKT1, and SMO. Science 339(6123):1077–1080. https://doi.org/10.1126/science.1233009
Sandberg AA, Stone JF (2008) The genetics and molecular biology of neural tumors. Springer, New York
Jacoby LB, Pulaski K, Rouleau GA, Martuza RL (1990) Clonal analysis of human meningiomas and schwannomas. Cancer Res 50(21):6783–6786
Zhu J, Frosch MP, Busque L, Beggs AH, Dashner K, Gilliland DG, Black PM (1995) Analysis of meningiomas by methylation- and transcription-based clonality assays. Cancer Res 55(17):3865–3872
van den Munckhof P, Christiaans I, Kenter SB, Baas F, Hulsebos TJ (2012) Germline SMARCB1 mutation predisposes to multiple meningiomas and schwannomas with preferential location of cranial meningiomas at the falx cerebri. Neurogenetics 13(1):1–7. https://doi.org/10.1007/s10048-011-0300-y
Collins VP, Nordenskjold M, Dumanski JP (1990) Molecular genetics of meningioma. Brain Pathol 1(1):19–24
Simon M, von Deimling A, Larson JJ, Wellenreuther R, Kaskel P, Waha A, Warnick RE, Tew JM, Menon AG (1995) Allelic losses on chromosomes 14, 10, and 1 in atypical and malignant meningiomas: a genetic model of meningioma progression. Cancer Res 55(20):4696
Choi M, Scholl UI, Ji W, Liu T, Tikhonova IR, Zumbo P, Nayir A, Bakkaloglu A, Ozen S, Sanjad S, Nelson-Williams C, Farhi A, Mane S, Lifton RP (2009) Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci USA 106(45):19096–19101. https://doi.org/10.1073/pnas.0910672106
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform 12:323. https://doi.org/10.1186/1471-2105-12-323
Warde-Farley D, Donaldson SL, Comes O, Zuberi K, Badrawi R, Chao P, Franz M, Grouios C, Kazi F, Lopes CT, Maitland A, Mostafavi S, Montojo J, Shao Q, Wright G, Bader GD, Morris Q (2010) The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res 38(suppl_2):W214–W220. https://doi.org/10.1093/nar/gkq537
Chen J, Bardes EE, Aronow BJ, Jegga AG (2009) ToppGene suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res 37(suppl_2):W305–W311. https://doi.org/10.1093/nar/gkp427
Mi H, Muruganujan A, Thomas PD (2013) PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res 41(D1):D377–D386. https://doi.org/10.1093/nar/gks1118
Koh YC, Yoo H, Whang GC, Kwon OK, Park HI (2001) Multiple meningiomas of different pathological features: case report. J Clin Neurosci 8(Suppl 1):40–43. https://doi.org/10.1054/jocn.2001.0875
Mocker K, Holland H, Ahnert P, Schober R, Bauer M, Kirsten H, Koschny R, Meixensberger J, Krupp W (2011) Multiple meningioma with different grades of malignancy: case report with genetic analysis applying single-nucleotide polymorphism array and classical cytogenetics. Pathol Res Pract 207(1):67–72. https://doi.org/10.1016/j.prp.2010.09.001
Tomita T, Kurimoto M, Yamatani K, Nagai S, Kuwayama N, Hirashima Y, Endo S (2003) Multiple meningiomas consisting of fibrous meningioma and anaplastic meningioma. J Clin Neurosci 10(5):622–624. https://doi.org/10.1016/s0967-5868(03)00161-9
Yang G, Chen X, Li CG, Zhang DM, Zhang WG, Shi CB, Zhao SG (2013) Multiple meningiomas characterized by benign and malignant tumor entities. CNS Neurosci Ther 19(12):984–986. https://doi.org/10.1111/cns.12179
Yudoyono F, Sidabutar R, Arifin MZ, Faried A (2015) Multiple meningiomas consisting of fibrous meningioma, transitional meningioma, and meningotheliomatous meningioma in one adult patient. Asian J Neurosurg 10(4):348–349. https://doi.org/10.4103/1793-5482.162729
Dewan R, Pemov A, Dutra AS, Pak ED, Edwards NA, Ray-Chaudhury A, Hansen NF, Chandrasekharappa SC, Mullikin JC, Asthagiri AR, Program NCS, Heiss JD, Stewart DR, Germanwala AV (2017) First insight into the somatic mutation burden of neurofibromatosis type 2-associated grade I and grade II meningiomas: a case report comprehensive genomic study of two cranial meningiomas with vastly different clinical presentation. BMC Cancer 17:127. https://doi.org/10.1186/s12885-017-3127-6
Liu Y, Zhang J, Li L, Yin G, Zhang J, Zheng S, Cheung H, Wu N, Lu N, Mao X, Yang L, Zhang J, Zhang L, Seth S, Chen H, Song X, Liu K, Xie Y, Zhou L, Zhao C, Han N, Chen W, Zhang S, Chen L, Cai W, Li L, Shen M, Xu N, Cheng S, Yang H, Lee JJ, Correa A, Fujimoto J, Behrens C, Chow CW, William WN, Heymach JV, Hong WK, Swisher S, Wistuba II, Wang J, Lin D, Liu X, Futreal PA, Gao Y (2016) Genomic heterogeneity of multiple synchronous lung cancer. Nature Commun 7:13200. https://doi.org/10.1038/ncomms13200
Wheeler SR, Shi C, Holt JA, Vnencak-Jones CL (2016) Mutation profiles of synchronous colorectal cancers from a patient with Lynch syndrome suggest distinct oncogenic pathways. J Gastrointest Oncol 7(3):E64-71. https://doi.org/10.21037/jgo.2016.01.07
Akagi K, Kurahashi H, Arita N, Hayakawa T, Monden M, Mori T, Takai S, Nishisho I (1995) Deletion mapping of the long arm of chromosome 22 in human meningiomas. Int J Cancer 60(2):178–182. https://doi.org/10.1002/ijc.2910600208
Chang IB, Cho BM, Moon SM, Park SH, Oh SM, Cho SJ (2010) Loss of heterozygosity at 1p, 7q, 17p, and 22q in meningiomas. J Korean Neurosurg Soci 48(1):14. https://doi.org/10.3340/jkns.2010.48.1.14
Ruttledge MH, Xie YG, Han FY, Peyrard M, Collins VP, Nordenskjöld M, Dumanski JP (1994) Deletions on chromosome 22 in sporadic meningioma. Genes Chromosom Cancer 10(2):122–130
Seizinger BR, de la Monte S, Atkins L, Gusella JF, Martuza RL (1987) Molecular genetic approach to human meningioma: loss of genes on chromosome 22. Proc Natl Acad Sci USA 84(15):5419–5423
Meese E, Blin N, Zang KD (1987) Loss of heterozygosity and the origin of meningioma. Hum Genet 77(4):349–351
Mihaila D, Gutiérrez JA, Rosenblum ML, Newsham IF, Bögler O, Rempel SA, Consortium NC (2003) Meningiomas: analysis of loss of heterozygosity on chromosome 10 in tumor progression and the delineation of four regions of chromosomal deletion in common with other cancers. Clin Cancer Res 9(12):4435–4442
Chang S-C, Lin J-K, Lin T-C, Liang W-Y (2005) Loss of heterozygosity: an independent prognostic factor of colorectal cancer. World J Gastroenterol 11(6):778
Bale AE (2002) Hedgehog signaling and human disease. Ann Rev Genomics Hum Genet 3:47–65. https://doi.org/10.1146/annurev.genom.3.022502.103031
Laurendeau I, Ferrer M, Garrido D, D’Haene N, Ciavarelli P, Basso A, Vidaud M, Bieche I, Salmon I, Szijan I (2010) Gene expression profiling of the hedgehog signaling pathway in human meningiomas. Mol Med 16(7–8):262–270. https://doi.org/10.2119/molmed.2010.00005
Mawrin C, Perry A (2010) Pathological classification and molecular genetics of meningiomas. J Neurooncol. https://doi.org/10.1007/s11060-010-0342-2
Pecina-Slaus N, Kafka A, Lechpammer M (2016) Molecular genetics of intracranial meningiomas with emphasis on canonical Wnt signalling. Cancers 8(7):67. https://doi.org/10.3390/cancers8070067
Ragel BT, Jensen RL (2010) Aberrant signaling pathways in meningiomas. J Neurooncol 99(3):315–324. https://doi.org/10.1007/s11060-010-0381-8
Wrobel G, Roerig P, Kokocinski F, Neben K, Hahn M, Reifenberger G, Lichter P (2005) Microarray-based gene expression profiling of benign, atypical and anaplastic meningiomas identifies novel genes associated with meningioma progression. Int J Cancer 114(2):249–256. https://doi.org/10.1002/ijc.20733
Choy W, Kim W, Nagasawa D, Stramotas S, Yew A, Gopen Q, Parsa AT, Yang I (2011) The molecular genetics and tumor pathogenesis of meningiomas and the future directions of meningioma treatments. Neurosurg Focus 30(5):E6. https://doi.org/10.3171/2011.2.FOCUS1116
Song L, Li ZY, Liu WP, Zhao MR (2015) Crosstalk between Wnt/beta-catenin and Hedgehog/Gli signaling pathways in colon cancer and implications for therapy. Cancer Biol Ther 16(1):1–7. https://doi.org/10.4161/15384047.2014.972215
Taipale J, Beachy PA (2001) The hedgehog and Wnt signalling pathways in cancer. Nature 411
Huard CC, Tremblay CeS, Helsper K, Delisle M-C, Schindler D, L ´evesque G, Carreau M (2012) Fanconi anemia proteins interact with CtBP1 and modulate the expression of the Wnt antagonist Dickkopf-1. Blood. https://doi.org/10.1182/blood-201202-408997
Huard CC, Tremblay CS, Magron A, Levesque G, Carreau M (2014) The Fanconi anemia pathway has a dual function in Dickkopf-1 transcriptional repression. Proc Natl Acad Sci USA 111(6):2152–2157. https://doi.org/10.1073/pnas.1314226111
Acknowledgements
This work was supported by Creative-Pioneering Researchers Program through Seoul National University (SNU No. 800-20160282).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
This study was approved by the Seoul National University Hospital institutional review board.
Research involving human and animal rights
This article does not contain any studies with animals performed by any of the authors.
Informed consent
The written consent was taken from the patient according to the Institutional Review Board guidelines before tumor removal.
Additional information
Data availability
The datasets analyzed during the current study are available in the NCBI sequence read archive under BioSample accessions SAMN7566682, SAMN7566683, SAMN7566684.
Tamrin Chowdhury and Yongjin Yoo are co-first authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chowdhury, T., Yoo, Y., Seo, Y. et al. Genomic analysis of synchronous intracranial meningiomas with different histological grades. J Neurooncol 138, 41–48 (2018). https://doi.org/10.1007/s11060-018-2772-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-018-2772-1