Abstract
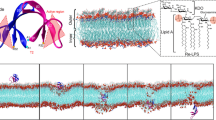
Emergence of antibiotic-resistant pathogens has paved way for development of newer class of drugs that would not be susceptible to resistance. Antimicrobial peptides such as defensins that target the microbial membrane are promising candidates. ROAD–1 is an alpha-defensin present in the oral cavity of rhesus macaque and shares very high sequence similarity to human enteric defensin 5. In this study we have performed microsecond long all atom molecular dynamic simulations to understand the mechanism of action of ROAD–1. We find that ROAD–1 is able to adopt an energetically stable conformation predominantly stabilized by electrostatic interactions only in presence of bacterial membranes. In mammalian membrane even though it gets absorbed onto the bilayer, it is unable to adopt an equilibrium conformation. Binding of ROAD–1 to bilayer induces clustering of POPG molecules up to 15 Å around the peptide. POPG molecules show higher order parameters than the neighboring POPE implying coexistence of different phases. Analysis of binding free energy of ROAD–1–membrane complex indicates Arg1, Arg2, Arg7, and Arg25 to play key role in its antimicrobial activity. Unlike its homolog HD5, ROAD–1 is not observed to form a dimer. Our study gives insight into the membrane-bound conformation of ROAD–1 and its mechanism of action that can aid in designing defensin-based therapeutics.

Antimicrobial peptide ROAD–1 adopts a different membrane-bound conformation as compared with HD5 even though they belong to the same family implying a different mechanism of action.








Similar content being viewed by others
References
Ganz T (2003) Defensins: antimicrobial peptides of innate immunity. Nat. Rev. Immunol. 3(9):710
Stotz HU, Thomson J, Wang Y (2009) Plant defensins: defense, development and application. Plant Signal. Behav. 4(11):1010–1012
Jarczak J, Kościuczuk EM, Lisowski P et al (2013) Defensins: natural component of human innate immunity. Hum. Immunol. 74(9):1069–1079
Selsted ME, Ouellette AJ (1995) Defensins in granules of phagocytic and non-phagocytic cells. Trends Cell Biol. 5(3):114–119
Lehrer RI (2004) Primate defensins. Nat. Rev. Microbiol. 2(9):727
Selsted ME, Ouellette AJ (2005) Mammalian defensins in the antimicrobial immune response. Nat. Immunol. 6(6):551
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature. 415(6870):389
Wimley WC (2010) Describing the mechanism of antimicrobial peptide action with the interfacial activity model. ACS Chem. Biol. 5(10):905–917
Wu Z, Hoover DM, Yang D et al (2003) Engineering disulfide bridges to dissect antimicrobial and chemotactic activities of human β-defensin 3. Proc. Natl. Acad. Sci. 100(15):8880–8885
Nguyen LT, Haney EF, Vogel HJ (2011) The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol. 29(9):464–472
Hristova K, Selsted ME, White SH (1996) Interactions of monomeric rabbit neutrophil defensins with bilayers: comparison with dimeric human defensin HNP-2. Biochemistry. 35(36):11888–11894
Wimley WC, Selsted ME, White SH (1994) Interactions between human defensins and lipid bilayers: evidence for formation of multimeric pores. Protein Sci. 3(9):1362–1373
Sagaram US, El-Mounadi K, Buchko GW et al (2013) Structural and functional studies of a phosphatidic acid-binding antifungal plant defensin MtDef4: identification of an RGFRRR motif governing fungal cell entry. PLoS One 8(12):e82485
Järvå M, Lay FT, Phan TK et al (2018) X-ray structure of a carpet-like antimicrobial defensin–phospholipid membrane disruption complex. Nat. Commun. 9(1):1962
Cools TL, Vriens K, Struyfs C et al (2017) The antifungal plant defensin HsAFP1 is a phosphatidic acid-interacting peptide inducing membrane permeabilization. Front. Microbiol. 8:2295
Payne JAE, Bleackley MR, Lee T-H et al (2016) The plant defensin NaD1 introduces membrane disorder through a specific interaction with the lipid, phosphatidylinositol 4, 5 bisphosphate. Biochim Biophys Acta 1858(6):1099–1109
Baxter AA, Poon IKH, Hulett MD (2017) The lure of the lipids: how defensins exploit membrane phospholipids to induce cytolysis in target cells. Nature Publishing Group, London
Phan TK, Lay FT, Poon IKH, Hinds MG, Kvansakul M, Hulett MD (2016) Human β-defensin 3 contains an oncolytic motif that binds PI (4, 5) P2 to mediate tumour cell permeabilisation. Oncotarget. 7(2):2054
Seo ES, Blaum BS, Vargues T et al (2010) Interaction of human β-defensin 2 (HBD2) with glycosaminoglycans. Biochemistry. 49(49):10486–10495
Schmitt P, Wilmes M, Pugnière M et al (2010) Insight into invertebrate defensin mechanism of action oyster defensins inhibit peptidoglycan biosynthesis by binding to lipid II. J. Biol. Chem. 285(38):29208–29216
Schneider T, Kruse T, Wimmer R et al (2010) Plectasin, a fungal defensin, targets the bacterial cell wall precursor lipid II. Science. 328(5982):1168–1172
Chileveru HR, Lim SA, Chairatana P, Wommack AJ, Chiang IL, Nolan EM (2015) Visualizing attack of Escherichia coli by the antimicrobial peptide human defensin 5. Biochemistry. 54(9):1767–1777
Chu H, Pazgier M, Jung G et al (2012) Human α-defensin 6 promotes mucosal innate immunity through self-assembled peptide nanonets. Science. 337(6093):477–481
Islam KT, Velivelli SLS, Berg RH, Oakley B, Shah DM (2017) A novel bi-domain plant defensin MtDef5 with potent broad-spectrum antifungal activity binds to multiple phospholipids and forms oligomers. Sci. Rep. 7(1):16157
Järvå M, Lay FT, Hulett MD, Kvansakul M (2017) Structure of the defensin NsD7 in complex with PIP2 reveals that defensin: lipid oligomer topologies are dependent on lipid type. FEBS Lett. 591(16):2482–2490
Baxter AA, Richter V, Lay FT et al (2015) The tomato defensin TPP3 binds phosphatidylinositol (4, 5)-bisphosphate via a conserved dimeric cationic grip conformation to mediate cell lysis. Mol Cell Biol 35(11):1964–1978
Zhang Y, Doherty T, Li J et al (2010) Resonance assignment and three-dimensional structure determination of a human α-defensin, HNP-1, by solid-state NMR. J. Mol. Biol. 397(2):408–422
Hong M, Su Y (2011) Structure and dynamics of cationic membrane peptides and proteins: insights from solid-state NMR. Protein Sci. 20(4):641–655
Zou G, de Leeuw E, Li C et al (2007) Toward understanding the cationicity of defensins ARG and LYS versus their noncoded analogs. J. Biol. Chem. 282(27):19653–19665
de Leeuw E, Rajabi M, Zou G, Pazgier M, Lu W (2009) Selective arginines are important for the antibacterial activity and host cell interaction of human α-defensin 5. FEBS Lett. 583(15):2507–2512
Tanabe H, Qu X, Weeks CS et al (2004) Structure-activity determinants in paneth cell α-defensins loss-of-function in mouse cryptdin-4 by charge-reversal at arginine residue positions. J. Biol. Chem. 279(12):11976–11983
Llenado RA, Weeks CS, Cocco MJ, Ouellette AJ (2009) Electropositive charge in α-defensin bactericidal activity: functional effects of Lys-for-Arg substitutions vary with the peptide primary structure. Infect. Immun. 77(11):5035–5043
Jung SW, Lee J, Cho AE (2017) Elucidating the bacterial membrane disruption mechanism of human α-Defensin 5: a theoretical study. J. Phys. Chem. B 121(4):741–748
Wang C, Shen M, Gohain N et al (2015) Design of a potent antibiotic peptide based on the active region of human defensin 5. J. Med. Chem. 58(7):3083–3093
Vasudevan S, Yuan J, Ösapay G et al (2008) Synthesis, structure, and activities of an oral mucosal α-defensin from rhesus macaque. J. Biol. Chem. 283(51):35869–35877
Tang Y-Q, Yuan J, Miller CJ, Selsted ME (1999) Isolation, characterization, cDNA cloning, and antimicrobial properties of two distinct subfamilies of α-defensins from rhesus macaque leukocytes. Infect. Immun. 67(11):6139–6144
Li J, Liu S, Lakshminarayanan R et al (2013) Molecular simulations suggest how a branched antimicrobial peptide perturbs a bacterial membrane and enhances permeability. Biochim. Biophys. Acta Biomembr. 1828(3):1112–1121
Sitaram N, Nagaraj R (1999) Interaction of antimicrobial peptides with biological and model membranes: structural and charge requirements for activity. Biochim Biophys Acta 1462(1):29–54
Murzyn K, Róg T, Pasenkiewicz-Gierula M (2005) Phosphatidylethanolamine-phosphatidylglycerol bilayer as a model of the inner bacterial membrane. Biophys. J. 88(2):1091–1103
Pastor RW, Feller SE (1996) Time scales of lipid dynamics and molecular dynamics. Biological Membranes. Springer, Berlin, pp 3–29
Hong C, Tieleman DP, Wang Y (2014) Microsecond molecular dynamics simulations of lipid mixing. Langmuir. 30(40):11993–12001
Domański J, Stansfeld PJ, Sansom MSP, Beckstein O (2010) Lipidbook: a public repository for force-field parameters used in membrane simulations. J. Membr. Biol. 236(3):255–258
Pronk S, Páll S, Schulz R et al (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics. 29(7):845–854
Vanommeslaeghe K, Hatcher E, Acharya C et al (2010) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31(4):671–690
Best RB, Zhu X, Shim J et al (2012) Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone ϕ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8(9):3257–3273
MacKerell Jr AD, Bashford D, Bellott M et al (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102(18):3586–3616
Klauda JB, Venable RM, Freites JA et al (2010) Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114(23):7830–7843
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79(2):926–935
Cuendet MA, van Gunsteren WF (2007) On the calculation of velocity-dependent properties in molecular dynamics simulations using the leapfrog integration algorithm. J Chem Phys 127(18):184102
Hess B (2008) P-LINCS: a parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4(1):116–122
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N· log (N) method for Ewald sums in large systems. J. Chem. Phys. 98(12):10089–10092
Nosé S (1984) A unified formulation of the constant temperature molecular dynamics methods. J. Chem. Phys. 81(1):511–519
Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A 31(3):1695
Berendsen HJC, Postma JPM, van Gunsteren WF, Di Nola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81(8):3684–3690
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52(12):7182–7190
Buchoux S (2016) FATSLiM: a fast and robust software to analyze MD simulations of membranes. Bioinformatics. 33(1):133–134
Carr M, MacPhee CE (2015) Membrainy: a ‘smart’, unified membrane analysis tool. Source Code Biol Med 10(1):3
Kumari R, Kumar R, Open source drug discovery consortium, Lynn A (2014) g_mmpbsa-- a GROMACS tool for high-throughput MM-PBSA calculations. J. Chem. Inf. Model. 54(7):1951–1962
Homeyer N, Gohlke H (2012) Free energy calculations by the molecular mechanics Poisson− Boltzmann surface area method. Mol Inform 31(2):114–122
Kollman PA, Massova I, Reyes C et al (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc. Chem. Res. 33(12):889–897
Kollman P (1993) Free-energy calculations—applications to chemical and biochemical phenomena. Chem. Rev. 93:2395–2417
Beveridge DL, Di Capua FM (1989) Free-energy via molecular simulation—applications to chemical and biomolecular systems. Annu. Rev. Biophys. Biophys. Chem. 18:431–492
Straatsma TP, McCammon JA (1991) Multiconfiguration thermodynamic integration. J. Chem. Phys. 95:1175
Lee J, Jung SW, Cho AE (2016) Molecular insights into the adsorption mechanism of human β-defensin-3 on bacterial membranes. Langmuir. 32(7):1782–1790
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25(13):1605–1612
Leekumjorn S, Sum AK (2007) Molecular characterization of gel and liquid-crystalline structures of fully hydrated POPC and POPE bilayers. J. Phys. Chem. B 111(21):6026–6033
Hall K, Lee TH, Aguilar MI (2011) The role of electrostatic interactions in the membrane binding of melittin. J. Mol. Recognit. 24(1):108–118
Jean-François F, Elezgaray J, Berson P, Vacher P, Dufourc EJ (2008) Pore formation induced by an antimicrobial peptide: electrostatic effects. Biophys. J. 95(12):5748–5756
Marrink SJ, De Vries AH, Tieleman DP (2009) Lipids on the move: simulations of membrane pores, domains, stalks and curves. Biochim. Biophys. Acta Biomembr. 1788(1):149–168
Matyus E, Kandt C, Tieleman DP (2007) Computer simulation of antimicrobial peptides. Curr. Med. Chem. 14(26):2789–2798
Kvansakul M, Lay FT, Adda CG et al (2016) Binding of phosphatidic acid by NsD7 mediates the formation of helical defensin–lipid oligomeric assemblies and membrane permeabilization. Proc. Natl. Acad. Sci. 113(40):11202–11207
Zhang Y, Lu W, Hong M (2010) The membrane-bound structure and topology of a human α-defensin indicate a dimer pore mechanism for membrane disruption. Biochemistry. 49(45):9770–9782
Rajabi M, Ericksen B, Wu X et al (2012) Functional determinants of human enteric α-defensin HD5: crucial role for hydrophobicity at the dimer interface. J. Biol. Chem. 287(26):21615–21627
Raschig J, Mailänder-Sánchez D, Berscheid A et al (2017) Ubiquitously expressed human beta defensin 1 (hBD1) forms bacteria-entrapping nets in a redox dependent mode of action. PLoS Pathog. 13(3):e1006261
Li J, Garg M, Shah D, Rajagopalan R (2010) Solubilization of aromatic and hydrophobic moieties by arginine in aqueous solutions. J. Chem. Phys. 133(5):054902
Su Y, Waring AJ, Ruchala P, Hong M (2010) Membrane-bound dynamic structure of an arginine-rich cell-penetrating peptide, the protein transduction domain of HIV TAT, from solid-state NMR. Biochemistry. 49(29):6009–6020
de Jong DH, Lopez CA, Marrink SJ (2013) Molecular view on protein sorting into liquid-ordered membrane domains mediated by gangliosides and lipid anchors. Faraday Discuss. 161:347–363
Lingwood D, Simons K (2010) Lipid rafts as a membrane-organizing principle. Science. 327(5961):46–50
Carruthers A, Melchior DL (1988) Effects of lipid environment on membrane transport: the human erythrocyte sugar transport protein/lipid bilayer system. Annu. Rev. Physiol. 50(1):257–271
Slater SJ, Kelly MB, Taddeo FJ, Ho C, Rubin E, Stubbs CD (1994) The modulation of protein kinase C activity by membrane lipid bilayer structure. J. Biol. Chem. 269(7):4866–4871
Zhu W, Xiong L, Peng J, Deng X, Gao J, Li C-M (2016) Molecular insight into affinities of gallated and nongallated proanthocyanidins dimers to lipid bilayers. Sci Rep 6:37680
Jang H, Ma B, Woolf TB, Nussinov R (2006) Interaction of protegrin-1 with lipid bilayers: membrane thinning effect. Biophys. J. 91(8):2848–2859
Chen F-Y, Lee M-T, Huang HW (2003) Evidence for membrane thinning effect as the mechanism for peptide-induced pore formation. Biophys. J. 84(6):3751–3758
Jefferies D, Hsu P-C, Khalid S (2017) Through the lipopolysaccharide glass: a potent antimicrobial peptide induces phase changes in membranes. Biochemistry. 56(11):1672–1679
Waheed Q, Tjörnhammar R, Edholm O (2012) Phase transitions in coarse-grained lipid bilayers containing cholesterol by molecular dynamics simulations. Biophys. J. 103(10):2125–2133
Tristram-Nagle S, Nagle JF (2004) Lipid bilayers: thermodynamics, structure, fluctuations, and interactions. Chem. Phys. Lipids 127(1):3–14
Hancock REW (1997) Peptide antibiotics. Lancet 349(9049):418–422
Bennett WFD, Hong CK, Wang Y, Tieleman DP (2016) Antimicrobial peptide simulations and the influence of force field on the free energy for pore formation in lipid bilayers. J. Chem. Theory Comput. 12(9):4524–4533
Tanizaki S, Feig M (2005) A generalized born formalism for heterogeneous dielectric environments: application to the implicit modeling of biological membranes. J. Chem. Phys. 122(12):124706
de Leeuw E, Burks SR, Li X, Kao JPY, Lu W (2007) Structure-dependent functional properties of human defensin 5. FEBS Lett. 581(3):515–520
Naafs MA (2018) The antimicrobial peptides: ready for clinical trials? Biomed J Sci Tech Res 7(4):6038–6042
Acknowledgments
SV would like to thank Department of Science and Technology, Govt. of India, for financial assistance (grant number SR/WOS-A/LS-566/2013) and C-DAC national supercomputing facilities for computational support. We would like to acknowledge Prof. P. V. Balaji, IIT Bombay for his valuable comments and computational support. SV would like to thank Nitin Kachariya and Rajalakshmi Panigrahi for assistance in making figures. SV would like to thank Prof. Micheal Selsted and Prof. Melaine Cocco for initiation into the world of antimicrobial peptides.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they no conflict of interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 42879 kb)
Rights and permissions
About this article
Cite this article
Vasudevan, S.V., Kumar, A. Antimicrobial peptide ROAD–1 triggers phase change in local membrane environment to execute its activity. J Mol Model 25, 281 (2019). https://doi.org/10.1007/s00894-019-4163-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-019-4163-8