Abstract
The actual conformation switching of proteins in the crowded cellular environment is completely different from that in vitro. Proteins in cytoplasm are continually subject to confinement and/or attraction to other molecules in their surroundings due to the existence of various biological species. To gain insight into the nature of crowded environments, we investigated the effects of confinement and affinity on the conformation switching of adenylate kinase (ADK) in a spherical cavity. It was found that even a small degree of confinement reduces the entropy of the open state and stabilizes the closed state, which leads to increased energy barriers for transition. Furthermore, the analysis of transition temperatures and mean first passage times indicates that the proper affinity can promote the transition of ADK from closed state to open state. This study reveals that the crowded cellular environment plays an important role in the thermodynamics and kinetics of proteins in vivo.

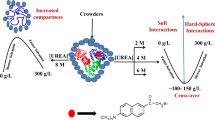
Cartoon representation of adenylate kinase in a spherical cavity. The LID, NMPand Core domains are highlighted in yellow, blue and magenta, respectively








Similar content being viewed by others
References
Ellis R (2001) J Curr Opin Struct Biol 11:500
Minton AP (2005) J Pharm Sci 94:1668
Ellis RJ, Minton AP (2003) Nature 425:27
Minton AP (2001) J Biol Chem 276:10577
Minton AP (1983) Mol Cell Biochem 55:119
Minton AP (1998) Methods Enzymol 295:127
Zimmerman SB, Minton AP (1993) Annu Rev Biophys Biomol Struct 22:27
Zimmerman SB, Trach SO (1991) J Mol Biol 222:599
Friedel M, Sheeler DJ, Shea JE (2003) J Chem Phys 118:8106
Eggers DK, Valentine JS (2001) Protein Sci 10:250
Eggers DK, Valentine JS (2001) J Mol Biol 314:911
Lei C, Shin Y, Liu J, Ackerman EJ (2002) J Am Chem Soc 124:11242
Wang YQ, Sarkar M, Smith AE, Krois AS, Pielak GJ (2012) J Am Chem Soc 134:16614
Arkin H, Janke W (2012) J Phys Chem B 116:10379
Chen E, Christiansen A, Wang Q, Cheung MS, Kliger DS, Wittung-Stafshede P (2012) Biochemistry 51:9836
Klimov DK, Newfield D, Thirumalai D (2002) Proc Natl Acad Sci USA 99:8019
Kurniawan NA, Enemark S, Rajagopalan R (2012) J Am Chem Soc 134:10200
Lucent D, Vishal V, Pande VS (2007) Proc Natl Acad Sci USA 104:10430
Malik A, Kundu J, Mukherjee SK, Chowdhury PK (2012) J Phys Chem B 116:12895
Marino KA, Bolhuis PG (2012) J Phys Chem B 116:11872
Martin J (2004) J Mol Recognit 17:465
Mittal J, Best RB (2008) Proc Natl Acad Sci USA 105:20233
Predeus AV, Gul S, Gopal SM, Feig M (2012) J Phys Chem B 116:8610
Rao JS, Cruz L (2013) J Phys Chem B 117:3707
Rathore N, Knotts TA, de Pablo J (2006) J Biophys J 90:1767
Wojciehowski M, Cieplak M (2008) Biosystems 94:248
Xu WX, Wang J, Wang W (2005) Proteins 61:777
Wang W, Xu WX, Levy Y, Trizac E, Wolynes PG (2009) Proc Natl Acad Sci USA 106:5517
Benton LA, Smith AE, Young GB, Pielak G (2012) J Biochem 51:9773
Zhou HX (2004) J Mol Recognit 17:368
Zhou HX, Dill KA (2001) Biochemistry 40:11289
Nakamura HK, Sasai M, Takano M (2004) Chem Phys 307:259
Cieplak M, Hoang TX, Robbins MO (2002) Proteins 49:114
Hayward S, Go N (1995) Annu Rev Phys Chem 46:223
Kim J, Keyes T (2008) J Phys Chem B 112:954
Ueeda Y, Taketomi H, Go N (1978) Bioplymers 17:1531
Hills RD, Brooks CL (2009) Int J Mol Sci 10:889
Lu Q, Wang J (2008) J Am Chem Soc 130:4772
Whitford PC, Miyashita O, Levy Y, Onuchic JN (2007) J Mol Biol 366:1661
Daily MD, Phillips GN, Cui QA (2010) J Mol Biol 400:618
Beckstein O, Denning EJ, Perilla JR, Woolf TB (2009) J Mol Biol 394:160
Lai ZZ, Lu Q, Wang J (2011) J Phys Chem B 115:4147
Chu JW, Voth GA (2007) Biophys J 93:3860
Levy Y, Wolynes PG, Onuchic JN (2004) Proc Natl Acad Sci USA 101:511
Lu Q, Wang J (2009) J Phys Chem B 113:1517
Cellmer T, Bratko D, Blanch H (2003) Biophys J 84:41a
Dukovski I, Muthukumar M (2003) J Chem Phys 118:6648
Liu C, Muthukumar M (1998) J Chem Phys 109:2536
Clementi C, Nymeyer H, Onuchic JN (2000) J Mol Biol 298:937
Bryngelson JD, Onuchic JN, Socci ND, Wolynes PG (1995) Proteins 21:167
Dill KA, Chan HS (1997) Nat Struct Biol 4:10
Onuchic JN, LutheySchulten Z, Wolynes PG (1997) Annu Rev Phys Chem 48:545
Wolynes PG (2005) Philos T R Soc A 363:453
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Grants No. 21433004 and 21473056), Shanghai Pu Jiang Program (12PJ1403000), Shanghai Natural Science Foundation (14ZR1411800) and a start-up grant of ECNU (41500-515430-14100/001/136). We also thank the supercomputer center of ECNU for computer time.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Sampled trajectories obtained at temperatures T = 0.5 (a), 0.65 (b), 0.75 (c) and 0.95 (d). The black and red curves denote the calculated RMSD of closed and open states with respect to the number of steps N, respectively. (GIF 80 kb)
Fig. S2
The 2D free energy profiles in bulk at temperatures T = 0.5 (a), 0.65 (b), 0.75 (c) and 0.95 (d) as a function of rmsd_close and rmsd_open in Ångstroms. The values of free energies range from 0.0 to 6.0 in units of K BTf 0. (GIF 136 kb)
Fig. S3
The 2D free-energy profiles are obtained at the radii R = 5.0,(a), 9.5 (b) and 12 Å (c), where the different temperatures T = 0.50, 0.65 and 0.95 correspond to the plots from left to right, respectively, as functions of rmsd_close and rmsd_open in Ångstroms. The values of free energies range from 0.0 to 6.0 in units of KBTf 0. (GIF 148 kb)
Fig. S4
The values of (Tf−Tf 0)/Tf 0 denoted by blue squares are approximately proportional to the function of R-3.75, which is indicated by the fitted red curve. (GIF 7 kb)
Rights and permissions
About this article
Cite this article
Li, M., Xu, W., Zhang, J.Z.H. et al. Combined effect of confinement and affinity of crowded environment on conformation switching of adenylate kinase. J Mol Model 20, 2530 (2014). https://doi.org/10.1007/s00894-014-2530-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-014-2530-z